Potri.001G427210 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
No hit found |
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G427210.1 pacid=42792678 polypeptide=Potri.001G427210.1.p locus=Potri.001G427210 ID=Potri.001G427210.1.v4.1 annot-version=v4.1
ATGTCGTGCTTATCCATTGAAGAATTACCTGATGAAATAGGAGAGCTCAAGGAGTTAAGGTTGTTGGATGTGACAGGTTGTCAAAGGCTAAGAAGGATCC
CTGTGAATTTGATTGGAAGGTTGAAGAAGTTAGAAGAGCTGTTGATCGGACATCTCAGCTTTAAGGGATGGGATGTTGTTGGATGTGACAGCACAGGAGG
AATGAATGCAAGCCTAACAGAACTAAATTCGTTGTCTCAGTTTGCCGTATTATCATTGAGGATACCGAAGGTTGAATGCATTCCCAGAGATTTTGTTTTT
CCAGTTAGCTTGCGCAAATATGATATAATATTAGGGAATGCTTTTGGCTATGGGTATTATTTATCAACCTCGACAAGATTATATTGGGCTGTTACATCCT
TAAATGCGAAGACATTTGAGCAATTGTTTTTACATAAATTAGAATCTGTAGTAGTGAGGGGTTGTGGGGATGTTTTCACTCTGTTTCCAGCAAGATTGCG
GCAAGGTTTAAAAAATCTGAAGGAGGTGGTTATTGACAACTGTGAATCATTGGAAGAGGTATTTGAATTGGGTGAGGAGAAGGAGCTGCCGCTGCTGTCA
TCTTTAACCAAGTTAGAGCTGTCACGTTTACCTGAGCTCAAATGCATATGGAAGGGGCCCACCAGTCATGTCAGCCTCCAAAGTCTTAATTTTCTGGAGT
TGGGGTATCTGGACAAACTGGCATTTATCTTCACACCGTCCCTCGCTCAAAGTCTACCACAACTGCAAACACTTGACATAACAGCATGCGGTGAATTGAA
GCATATTATAAGAGAAGAGGATGGTGAAAGGGAAATAATTCCAGAGTCTCCTTGCTTCCCACAATTAAAAAATATCTTCATACAACGGTGTGGTAAACTG
GAATATGTCTTCCCTGTCTCCATGTCTCCAAGCCTTCCGAACCTGGAACAGATGACGATTTATTATGCTGACAATTTAAAGCAAATATTTTACAGTGGAG
AAGGAGATGCACTCACCACAGATGGCATCATCAAGTTCCCTCAACTAAGAGAATTGTCTCTTGGGCTCCGATCAAATTACGGCTTTATAGGTCCAAGGAA
TTTTGATTTTCAATTGCCTTCTTTGCAGAATCTAAACATTAAAGGCCACGAAGAAGTGGGTAATTGGTTGGCACAGCTACAACAAAACGGCTTCTTACAA
AGATTAAAATCTGTAGCAGTGGACGATTGTGGGGATGTTCGCACTCCGTTTCCAGCAAAATTGCTGCGAGCTGTGAAAAATCTAAGCAGTGTGAACATTA
ACGGCTGCAAATCATTGGAAGAGGTATTTGAATTAGGTGAGCCTGATGAAGGAAGTAGGGAGGAGAAGGAGCTGCCGCTTCTGTCATCTTTAACAGGGTT
ACGGCTGTCACGTTTACCTGAGCTCAAATGCATTTGGAAGGGGCCCACCAGACATATCAGCCTCCAAAGTCTTGCTCATCTGTATTTGAATTCTCTCGAC
AAACTGATATTTATCCTCACACCGTCTCTCGCTCGAAGTCTGCCAAAGCTAGAAATACTTGAGATAAGTGAATGCGGTGAATTGAAGCATATTATCAGAG
AAGAGGATGGTGAAAGGGAAATAATTCCAGAGTCTCCTTGCTTCCCACAATTAAAAAATATCTTCATAGAACGGTGTGGTAAACTGGAATATGTCTTCCC
TGTCTCCATGTCTCCAAGCCTTCCGAACCTGGAACAGATGACGATTTATTATGCTGACAATTTAAAGCAAATATTTTACAGTGGAGAAGGAGACGCACTC
ACCACAGATGGCATCATCAAGTTCCCTCGATTAAGTGACTTGGTTCTTTCTTCCATATCAAATTACAGCTTTTTTGGTCCAACGAATCTTGCTGCCCAAT
TGCCTTCTTTGCGATTTCTAAAAATCAATGGCCACAAAGAATTGGGAAATTTGTTTGCACAGCTCCAATGCACAAAATTTCTTTTGATGCAGGGGTTCAC
AAATTTGAAAGAATTAAGCTTGGAATCCGTGCCTGACTTGAGGGGTCTTCTGCTGAGCAAATTGACTACTTTGGAGATGGCCGCCCATGGGGAGCAAAAC
GGCTCCTTACATAGATTAGAACGTGTACGAGTGGACGATTGTGGGGATGTTCGCGCTCCGTTTCCAGCAAAATTGCTGCGAGCTTTGAAAAATCTAAGCA
GTGTGAACATTAACGGCTGCAAATCATTGGAAGAGGTATTTGAATTAGGTGAGCCTGATGAAGGAAGTAGGGAGGAGAAGGAGCTGCCGCTTCTGTCATC
TTTAACAGGGTTACGGCTGTCAGGTTTACCTGAGCTCAAATGCATGTGGAAGGGGCCCACCAGACATGTCAGCCTCCAAAGTCTTGCTTATCTGGATTTG
TGGTCTCTGGACAAACTGACATTTATATTCACACCGTCCCTCGCTCGAAGTCTGCCAAAGCTAGAAAGACTTTACATAGGTAAATGCGGTCAATTGAAGC
ATATTATCAGAGAAGAGGATGGTGAAAAGGAAATAATTCCAGAGCCTCCTGGGCAGGATGGTCAAGCTTCACCTATCAACGTTGAGAAGGAGATAGTGCT
TCCTAATCTAAAGGAGTTGTCGATACAACAATTATCAAGTATTGTCTGTTTTAGTTTCGGATGGTGTGATTATTTGTTATTCCCTCGTTTGGAGAAGTTG
GAGGTCCATCTATGTCCAAAGCTGACCACAAAATTTGCTAGTACACCAGATGGTTCAATGAGTGCTCAATCAGAGGTATCTGAAGTAGCTGAAGATTCAA
GCATTTATAGAGAGTGGACCCGAGATAAGGGGTGGAAAGAACGATGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G427210.1 pacid=42792678 polypeptide=Potri.001G427210.1.p locus=Potri.001G427210 ID=Potri.001G427210.1.v4.1 annot-version=v4.1
MSCLSIEELPDEIGELKELRLLDVTGCQRLRRIPVNLIGRLKKLEELLIGHLSFKGWDVVGCDSTGGMNASLTELNSLSQFAVLSLRIPKVECIPRDFVF
PVSLRKYDIILGNAFGYGYYLSTSTRLYWAVTSLNAKTFEQLFLHKLESVVVRGCGDVFTLFPARLRQGLKNLKEVVIDNCESLEEVFELGEEKELPLLS
SLTKLELSRLPELKCIWKGPTSHVSLQSLNFLELGYLDKLAFIFTPSLAQSLPQLQTLDITACGELKHIIREEDGEREIIPESPCFPQLKNIFIQRCGKL
EYVFPVSMSPSLPNLEQMTIYYADNLKQIFYSGEGDALTTDGIIKFPQLRELSLGLRSNYGFIGPRNFDFQLPSLQNLNIKGHEEVGNWLAQLQQNGFLQ
RLKSVAVDDCGDVRTPFPAKLLRAVKNLSSVNINGCKSLEEVFELGEPDEGSREEKELPLLSSLTGLRLSRLPELKCIWKGPTRHISLQSLAHLYLNSLD
KLIFILTPSLARSLPKLEILEISECGELKHIIREEDGEREIIPESPCFPQLKNIFIERCGKLEYVFPVSMSPSLPNLEQMTIYYADNLKQIFYSGEGDAL
TTDGIIKFPRLSDLVLSSISNYSFFGPTNLAAQLPSLRFLKINGHKELGNLFAQLQCTKFLLMQGFTNLKELSLESVPDLRGLLLSKLTTLEMAAHGEQN
GSLHRLERVRVDDCGDVRAPFPAKLLRALKNLSSVNINGCKSLEEVFELGEPDEGSREEKELPLLSSLTGLRLSGLPELKCMWKGPTRHVSLQSLAYLDL
WSLDKLTFIFTPSLARSLPKLERLYIGKCGQLKHIIREEDGEKEIIPEPPGQDGQASPINVEKEIVLPNLKELSIQQLSSIVCFSFGWCDYLLFPRLEKL
EVHLCPKLTTKFASTPDGSMSAQSEVSEVAEDSSIYREWTRDKGWKER
|
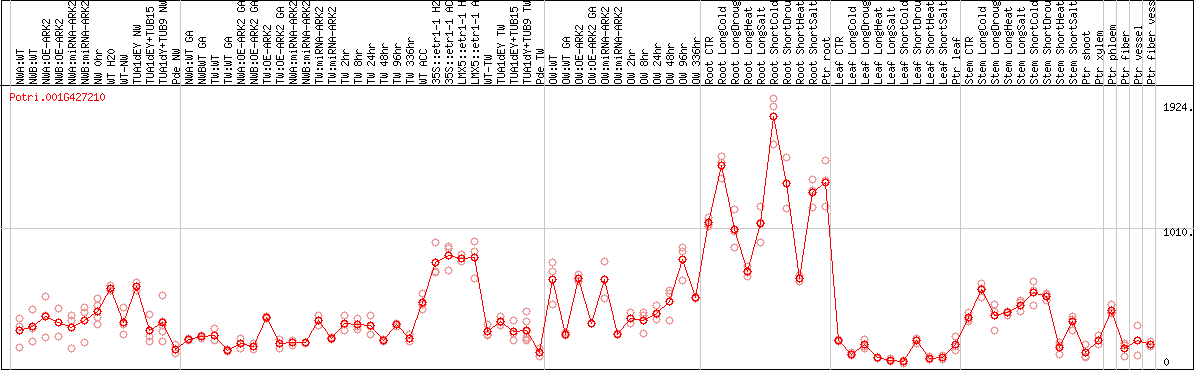
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G427210 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
