MOR2,Pt-MOR1.2 (Potri.001G443200) [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | MOR2,Pt-MOR1.2 | ||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info | |||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G443200.2 pacid=42790174 polypeptide=Potri.001G443200.2.p locus=Potri.001G443200 ID=Potri.001G443200.2.v4.1 annot-version=v4.1
ATGTCGGAAGAAGAGAAGTTGTTAAAGGAAGCGAAGAAATTGGCATGGGAAGATCGGTTGTTACACAAGAACTGGAAAGTTAGAAACGAAGCCAATATCG
ATTTGGCTTCTCTTTGTGATTCCATTTCAGATCCTAAAGACTCCAGACTTCGTGAATTCGCTCCGTTGTTTAGGAAAACGGTTGCGGATTCAAATGCGCC
TGTACAGGAGAAAGCGCTCGATGCTTTGATTGCGTTTTTACGTGCTGCCGATGCGGATGCTGGAAGGTATGCGAAAGAAGTGTGTGATGCGATTGTGGCT
AAGTGTCTTACTGGTAGGCCAAAGACTGTGGAGAAAGCGCAGGCAGCGTTTATGCTGTGGGTGGAGTTGGAAGCTGTGGATGTTTTTTTGGATGCAATGG
AGAAAGCAATTAAGAACAAAGTTGCCAAAGCTGTGGTGCCCGCTATAGACGTTATGTTTCAAGCCTTAAGTGATTTTGGAGCCAAAGTTGTCCCCCCAAA
GAGGATTTTAAAGATGCTTCCAGAATTGTTTGACCACCAGGATCAAAATGTCCGTGCATCATCTAAAGGACTAACTCTTGAGCTTTGTCGGTGGATCGGA
AAAGATCCTGTGAAATCAATTCTCTTTGAGAAGATGCGAGATACCATGAAAAAAGAATTGGAGGCTGAGCTTGTTAATGTTAAAGGGACAGCTAAGCCAT
CCCGCAAGATTAGATCTGAGCAAGACAAGGAGCCAGAGCCAGAAGGGGTATCAGAAGTTGTGGGTTCTGGTCCATCTGAAGAAGTTGCTGCTGAAGCTCC
TCAAGAAATAGATGAGTATGACCTTGTAGACCCGGTTGATATTTTGGGTCCTTTGGAGAAGGCTGGATTTTGGGATGGAGTGAAAGCCACCAAATGGTCA
GAAAGAAAGGAGGCTGTTGCAGAGCTGACGAAGCTTGCATCCACAAAGAGGATAGCACCTGGTGATTTCTCTGAAGTTTGTCGGACTTTGAAGAAGCTTA
TTACAGATGTGAACATCGCTGTTGCAGTTGAAGCCATACAAGCTATTGGCAACCTTGCACGTGGTCTAAGAACCCATTTTTCTGGGAGCTCTCGGTTCTT
ATTACCTGTATTGCTTGAAAAATTGAAAGAGAAAAAACCTACACTCACCGAGGCACTTGCTCAAACTCTGCAAGCGATGCATACAGCTGGATGCTTAAAT
CTTGCTGATATCATTGAAGATGTCAAAACAGCAGTGAAAAATAAAGTTCCTCTAGTGCGCTCATTAACTTTAAACTGGGTGACATTTTGTATTGAAACAA
GTAACAAGGCTGTTATTCTGAAGGTGCACAAGGATTATGTCCCTATCTGTATGGAGTGCCTCAATGATGGGACTCCTGATGTGAGGGATTCTGCCTTTTC
AGTGTTAGCAGCAGTAGCTAAGTCGGTTGGCATGAGGCCTTTGGAGCGATCACTGGAGAAACTTGATGATGTGAGAAGAAAGAAGCTGTCTGAAATGATA
GCAGGTTCAGGAGATGGTGTGCCTGCTGTGGCATCCTCAGGTCCAGTCCAAGCTGTTCGTGGAAGCATGTCATCGGTAGAGACATCAGAAGGCTCATTTG
TTAAAAAGTCGGCGGCAAGCATGCTTTCTGGGAAGAGGCCTGCACCAGCAGCTGCTGCTAACAAAAAGGCAGCGCCGACTAAATCAGGGGTTAGCAAGAA
AGGAGATGGTGCTGGGCGGGCAGAATCTTCAAGAGCAATTGAACCACCTGAGGATGTTGAGCCAGCAGAGATGAGTCTTGAAGAAATTGAAACCAGATTA
GGTTCTCTCATTCAAGCAGATACTGTTTCCCAACTGAAGAGCGCAGTGTGGAAAGAGCGGTTGGAAGCAATATCCTCATTTAAACTACAAGTGGAGGGGT
TACAGAATCTTGATCAATCTGTGGAGATTTTGATTCGTTTACTCTGTGCTATTCCTGGATGGAACGAAAAAAATGTTCAGGTTCAGCAACAGGTCATTGA
AGTCATTACTTACCTAGCTTCAACTGCATCAAAATTTCCGAAGAAATGTGTAGTGCTCTGCCTTTTAGGTATAAGTGAACGGGTAGCTGATATTAAAACC
AGAGCTCATGCCATGAAGTGCCTTACCACTTTCTCTGAAGCAGTGGGTCCAGGATTTGTTTTTGACCGACTTTACAAAATTATGAAAGAACACAAGAATC
CAAAGGTTCTTAGTGAGGGAATAATCTGGATGGTTTCGGCAATTGATGACTTTGGTGTGTCACATTTGAAGCTTAAGGATTTAATTGATTTCTGTAAAGA
TACTGGGCTACAGTCGAGTGTAGCCGCAAGCAGAAATGCAACTATCAAACTTTTGGGTGCTTTACATAAGTTTGTTGGTCCAGATATTAAAGGGTTTCTT
GCTGATGTCAAACCTGCACTGCTGAGTGCACTTGATGCTGAGTATGATAAAAATCCATTTGAGGGTGCTTCCGCCGCTCCAAAGAAAACAGTCAGGACAT
CAGAATCCACCTCTTCTGTTTCTGGTGGTGGGCTGGATAGCCTGCCACGTGAAGACATTAGTGGAAAGATAACACCTACTCTGATCAAGAGTTTGGAAAG
TCCTGATTGGAAGGTTCGACTGGAGTCGATTGAAGCTGTCAATAAAATTTTAGAGGAGGCTAATAAACGCATTCAACCAACTGGAACTGGTGAATTATTT
GGTGCCCTTAGAGGGCGTCTCTATGACAGCAATAAAAATTTAATCATGACAGCTTTGACTACCATTGGTGGTGTTGCTTCTGCTATGGGACCGGCTGTTG
AAAAGTCAAGCAAGGGTGTTCTGTCGGATATTTTGAAATGTCTTGGGGATAATAAGAAACATATGAGGGAATGCACTCTGAATACTTTAGATTCTTGGGT
TGCTGCTGTTCATCTTGATAAAATGGTTCCTTACATCACAGCAGCTCTGATAGAAACTAAGCTTGGTGCTGAAGGCCGCAAGGATCTTTTTGATTGGCTG
TCAAAGCAACTTTCTGGATCAAGTGAGTTTTCTGATGCCATACATTTGCTGAAACCAGCCAGCTCTGCTATGACGGATAAATCATCAGATGTTCGAAAAG
CAGCTGAAGCATGCATCTCCGAGATTTTGAGAGTGTGTGGACAAGAAATGATTGAGAAGAATCTGAAAGATATTCAAGGGCCAGCGTTGGCTCTTGTTCT
TGAACGAGTGAGACCTGCAGGTGGTTTTCAAGAATCGTTTGAATCTACAAAAACAATTTCAATGGGGCCATCATCCAAAACAAGTGTAAAGGTTGGAAAA
GCTGCTTCCAATGGCATCTCAAAACATGCAAATAGATCTATATCTGCGAGAGTCATACCAATGAAAGGGTCAAAGCCAGAGCCAACTATGTCTTTTCAGG
ACAGAGCTGTTCAGTCACAAGCATTATTGAATGTCAAGGACTCAAACAAGGAGGATAGGGAGAGAATGGTAGTACGGAGGTTTAAGTTTGAAGAGCCACG
GATGGAACAGGTTCAAGATCTTGAGAGTGACATGATGAAATACTTCAGAGAGGATTTGAACAGGCGACTTCTAAGTCCGGATTTTAAGAAACAAGTCGAC
GGGCTAGAGATGTTACATAAGGCCCTTCCATCCATTGGAAAGGAAATAATTGAAGTTTTGGACATTCTTTTGAGGTGGTTTGTTTTGCAGTTTTGTAAAT
CTAACACAACTTGCCTATTGAAGGTGCTGGAATTTCTCCCTGATCTTTTTGACCGGCTCCGGGATGAGGCCTACACATTGAGTGAATCTGAAGCAGCTAT
ATTTCTTCCATGCTTGATTGAAAAGTTGGGGCATAACATTGAGAAAGTGCGAGAAAAAATGCGTGAGCTTACCAAACAAATTGTGCAAGCATATTCAGCA
GCAAAATCTTTCCCTTACATTCTGGAGGGTTTGCGCTCTAAAAACAACCGTACTCGAATTGAGTGTGCAGATCTAGTTGGGTTCTTGATTGATCATCATG
GGGCTGAGATAAGTGGACAGTTAAAATCCTTGCAAATTGTTGCAAGCTTGACAGCAGAGCGTGATGGCGAAACTAGAAAAGCTGCGTTAAATACACTGGC
TACAGGTTACAAGATTCTTGGTGAGGACATTTGGAGATTTCTTGGGAAGCTGACAGATGCTCAGAAAAGCATGATAGATGATAGATTTAAGTGGAAGGTA
CGAGAGATGGAAAAAAGGAAGGAAGGTCGCCCTGGTGATGCTAGAGCTGCTTTAAGGCGCTCTGTTAGGGAAAATGGGTCTGATATAGCAGAACAGAGCG
GGGAACTTTCACAATCAGTCTCTGGTCCTATTATAGCTAGGAAAAACTATGGGACGCAGGAGCTTCATATGGAGGGACATATGATGCCCCGTGCACTCGT
GAGTGTCAATGGCCCTGCAGACTGGAATGAAGCTCTGGACATCATTTCTTTTGGTTCCCCGGAGCAGTCAGTTGAAGGTATGAAAGTTGTTTGCCATGAG
TTGGCACAGGCCACAAACGATGCAGAAGGCAGTGCTATGGATGAACTTGTGAAAGATGCAGATAAACTTGTTTCATGCTTGGCAAACAAGGTATCGAGGA
CTTTTGACTTCAGTTTGACTGGAGCCTCATCACGGGCTTGTAAATATGTACTTAATACACTCATGCAGACTTTTCAAAATAAAATTCTTGCTTATGCTGT
CAAAGAGAGTACTCTTGACAGTTTGATAACGGAGCTTCTACTTTGGCTTTTGGATGAAAGGGTCCCACATATGGATGATGGCAGCCAGCTCCTAAAGGCA
TTGAATGTGCTGATGCTTAAGATTCTGGATAATGCAGATCGAACTTCATCCTTTGTTGTGCTGATTAATCTGTTGCGACCACTGGATCCCACAAGGTGGC
CTTCACCTGCATCAGCTGAGACCTTTGCTATCAGGAATCAGAAGTTTTCTGACCTTGTTGTAAAATGCCTTATCAAACTCACAAAGGTTCTTCAAACTAC
TATATATGATGTTGATCTTGACCGCATCCTTCAGAGCATCCATATTTACCTACAAGAATTGGGGATGGAAGAAATCAGGAGAAGAGCTGGAGCAGATGAC
AAGCCTTTGCGTATGGTGAAAACTGTTCTCCATGAACTTGTTAAGCTTCGTGGAGCTGCAATAAAGGGTCACCTATCGATGGTTCCTATCGACATGAAAC
CCCAGCCAATTATTCTTGCCTACATTGATCTTAATCTTGAGACTTTAGCTGCAGCAAGAATGTTGACCTCAACAGCTCCTGTGGGCCAAAATCATTGGGG
TGATTCTGCAGCCAACAATTCATCACCTGCCGCGCATTCTGCTGAGGCTCAGTTGAAGCAAGAACTTGCTGCCATTTTCAAGAAAATTGGTGACAAACAA
ACCTGCACCATTGGTCTATATGAGCTCTATCGCATCACCCAGTTATATCCTAAGGTTGATATATTTGCTCAACTCCAAAATGCTAGTGAGGCATTTCGCA
CTTACATCAGAGATGGTCTAGCTCAGATGGAGAAGAATACTGCTGCTGGAAGGACACCTTCTAGTTTGCCAATTTCAACTCCCCCGCCATCCGCCTTAAA
TGTTTCTTCCCCTGATTTGCAACCCCTTTCCCCTGTGCATACAAACTCCTTAAATGATGCTAAACCATTGCATGTTAAACCCGAAACAACAAACTTTCAT
CTTCCTCCCTCGTATGCTGAAGACAACAGAGCAGTTAGTGCCTTTTTATCTAGAGGTCTCGTCTCAGAGAATTCTTTGGGTGACCAAAGAAATGAGAAGC
TTATTGGTGGAGTAACCAGTGGAACATTAGATGCAATTAGAGAGAGAATGAAGAGTATGCAGTTAGCTGCTGCTACTGGGAATCCCGATTCAGGAAGCAG
GCCATTGATGTCTATGAATGAGAACCTAAATAATGGACTGTCAAGTCAGATTCTTCGTGCGCCAGACTCGACGGGTATGGAGAATCCTTTACATAGCGGA
GTTCTTCCCATGGATGAGAAGGCATTGTCTGGTCTTCAAGCCAGGATGGAGAGACTTAAAAGTGGATCTCTTGAACCCCTGTAG
|
||||||||||||||||||||
|
AA sequence
|
>Potri.001G443200.2 pacid=42790174 polypeptide=Potri.001G443200.2.p locus=Potri.001G443200 ID=Potri.001G443200.2.v4.1 annot-version=v4.1
MSEEEKLLKEAKKLAWEDRLLHKNWKVRNEANIDLASLCDSISDPKDSRLREFAPLFRKTVADSNAPVQEKALDALIAFLRAADADAGRYAKEVCDAIVA
KCLTGRPKTVEKAQAAFMLWVELEAVDVFLDAMEKAIKNKVAKAVVPAIDVMFQALSDFGAKVVPPKRILKMLPELFDHQDQNVRASSKGLTLELCRWIG
KDPVKSILFEKMRDTMKKELEAELVNVKGTAKPSRKIRSEQDKEPEPEGVSEVVGSGPSEEVAAEAPQEIDEYDLVDPVDILGPLEKAGFWDGVKATKWS
ERKEAVAELTKLASTKRIAPGDFSEVCRTLKKLITDVNIAVAVEAIQAIGNLARGLRTHFSGSSRFLLPVLLEKLKEKKPTLTEALAQTLQAMHTAGCLN
LADIIEDVKTAVKNKVPLVRSLTLNWVTFCIETSNKAVILKVHKDYVPICMECLNDGTPDVRDSAFSVLAAVAKSVGMRPLERSLEKLDDVRRKKLSEMI
AGSGDGVPAVASSGPVQAVRGSMSSVETSEGSFVKKSAASMLSGKRPAPAAAANKKAAPTKSGVSKKGDGAGRAESSRAIEPPEDVEPAEMSLEEIETRL
GSLIQADTVSQLKSAVWKERLEAISSFKLQVEGLQNLDQSVEILIRLLCAIPGWNEKNVQVQQQVIEVITYLASTASKFPKKCVVLCLLGISERVADIKT
RAHAMKCLTTFSEAVGPGFVFDRLYKIMKEHKNPKVLSEGIIWMVSAIDDFGVSHLKLKDLIDFCKDTGLQSSVAASRNATIKLLGALHKFVGPDIKGFL
ADVKPALLSALDAEYDKNPFEGASAAPKKTVRTSESTSSVSGGGLDSLPREDISGKITPTLIKSLESPDWKVRLESIEAVNKILEEANKRIQPTGTGELF
GALRGRLYDSNKNLIMTALTTIGGVASAMGPAVEKSSKGVLSDILKCLGDNKKHMRECTLNTLDSWVAAVHLDKMVPYITAALIETKLGAEGRKDLFDWL
SKQLSGSSEFSDAIHLLKPASSAMTDKSSDVRKAAEACISEILRVCGQEMIEKNLKDIQGPALALVLERVRPAGGFQESFESTKTISMGPSSKTSVKVGK
AASNGISKHANRSISARVIPMKGSKPEPTMSFQDRAVQSQALLNVKDSNKEDRERMVVRRFKFEEPRMEQVQDLESDMMKYFREDLNRRLLSPDFKKQVD
GLEMLHKALPSIGKEIIEVLDILLRWFVLQFCKSNTTCLLKVLEFLPDLFDRLRDEAYTLSESEAAIFLPCLIEKLGHNIEKVREKMRELTKQIVQAYSA
AKSFPYILEGLRSKNNRTRIECADLVGFLIDHHGAEISGQLKSLQIVASLTAERDGETRKAALNTLATGYKILGEDIWRFLGKLTDAQKSMIDDRFKWKV
REMEKRKEGRPGDARAALRRSVRENGSDIAEQSGELSQSVSGPIIARKNYGTQELHMEGHMMPRALVSVNGPADWNEALDIISFGSPEQSVEGMKVVCHE
LAQATNDAEGSAMDELVKDADKLVSCLANKVSRTFDFSLTGASSRACKYVLNTLMQTFQNKILAYAVKESTLDSLITELLLWLLDERVPHMDDGSQLLKA
LNVLMLKILDNADRTSSFVVLINLLRPLDPTRWPSPASAETFAIRNQKFSDLVVKCLIKLTKVLQTTIYDVDLDRILQSIHIYLQELGMEEIRRRAGADD
KPLRMVKTVLHELVKLRGAAIKGHLSMVPIDMKPQPIILAYIDLNLETLAAARMLTSTAPVGQNHWGDSAANNSSPAAHSAEAQLKQELAAIFKKIGDKQ
TCTIGLYELYRITQLYPKVDIFAQLQNASEAFRTYIRDGLAQMEKNTAAGRTPSSLPISTPPPSALNVSSPDLQPLSPVHTNSLNDAKPLHVKPETTNFH
LPPSYAEDNRAVSAFLSRGLVSENSLGDQRNEKLIGGVTSGTLDAIRERMKSMQLAAATGNPDSGSRPLMSMNENLNNGLSSQILRAPDSTGMENPLHSG
VLPMDEKALSGLQARMERLKSGSLEPL
|
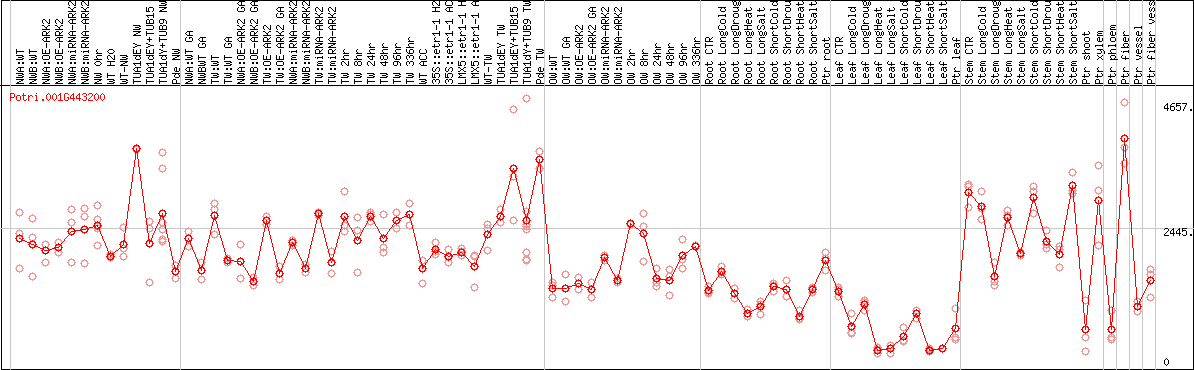
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G443200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
