External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G32790 365 / 4e-126
CID11
CTC-interacting domain 11 (.1.2)
AT4G10610 350 / 2e-120
ATRBP37, RBP37, CID12
RNA-BINDING PROTEIN 37, CTC-interacting domain 12 (.1.2)
AT3G49390 318 / 2e-107
CID10
CTC-interacting domain 10 (.1.2)
AT3G14450 289 / 1e-96
CID9
CTC-interacting domain 9 (.1)
AT1G53650 285 / 2e-95
CID8
CTC-interacting domain 8 (.1.2)
AT5G24440 243 / 2e-78
CID13
CTC-interacting domain 13 (.1)
AT3G23900 50 / 1e-06
RNA recognition motif (RRM)-containing protein (.1), RNA recognition motif (RRM)-containing protein (.2), RNA recognition motif (RRM)-containing protein (.3)
AT5G09880 49 / 5e-06
Splicing factor, CC1-like (.1)
AT5G10350 46 / 1e-05
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
AT3G55340 47 / 2e-05
PHIP1
phragmoplastin interacting protein 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G152700
468 / 2e-166
AT4G10610 401 / 5e-140
RNA-BINDING PROTEIN 37, CTC-interacting domain 12 (.1.2)
Potri.012G022000
338 / 3e-115
AT1G32790 428 / 4e-150
CTC-interacting domain 11 (.1.2)
Potri.015G005400
335 / 5e-114
AT1G32790 415 / 5e-145
CTC-interacting domain 11 (.1.2)
Potri.001G377500
288 / 2e-96
AT3G14450 424 / 2e-150
CTC-interacting domain 9 (.1)
Potri.011G095800
286 / 1e-95
AT3G14450 391 / 3e-137
CTC-interacting domain 9 (.1)
Potri.017G058300
60 / 1e-09
AT3G23900 447 / 2e-142
RNA recognition motif (RRM)-containing protein (.1), RNA recognition motif (RRM)-containing protein (.2), RNA recognition motif (RRM)-containing protein (.3)
Potri.001G319000
58 / 4e-09
AT3G23900 386 / 4e-119
RNA recognition motif (RRM)-containing protein (.1), RNA recognition motif (RRM)-containing protein (.2), RNA recognition motif (RRM)-containing protein (.3)
Potri.010G209701
57 / 7e-09
AT3G55340 265 / 2e-82
phragmoplastin interacting protein 1 (.1)
Potri.005G073200
52 / 1e-07
AT5G10350 292 / 2e-101
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033242
360 / 4e-124
AT1G32790 436 / 1e-153
CTC-interacting domain 11 (.1.2)
Lus10008275
363 / 6e-124
AT1G32790 437 / 8e-153
CTC-interacting domain 11 (.1.2)
Lus10008851
332 / 1e-112
AT3G49390 427 / 7e-150
CTC-interacting domain 10 (.1.2)
Lus10032942
317 / 7e-107
AT3G49390 421 / 2e-147
CTC-interacting domain 10 (.1.2)
Lus10032941
292 / 1e-97
AT1G32790 383 / 4e-133
CTC-interacting domain 11 (.1.2)
Lus10032590
57 / 2e-08
AT3G23900 647 / 0.0
RNA recognition motif (RRM)-containing protein (.1), RNA recognition motif (RRM)-containing protein (.2), RNA recognition motif (RRM)-containing protein (.3)
Lus10043159
56 / 2e-08
AT3G23900 635 / 0.0
RNA recognition motif (RRM)-containing protein (.1), RNA recognition motif (RRM)-containing protein (.2), RNA recognition motif (RRM)-containing protein (.3)
Lus10027886
54 / 1e-07
AT4G34110 922 / 0.0
ARABIDOPSIS POLY\(A\) BINDING 2, poly(A) binding protein 2 (.1)
Lus10002835
54 / 1e-07
AT4G34110 918 / 0.0
ARABIDOPSIS POLY\(A\) BINDING 2, poly(A) binding protein 2 (.1)
Lus10010042
53 / 2e-07
AT4G34110 865 / 0.0
ARABIDOPSIS POLY\(A\) BINDING 2, poly(A) binding protein 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
CL0221
PF07145
PAM2
Ataxin-2 C-terminal region
Representative CDS sequence
>Potri.001G448800.3 pacid=42790748 polypeptide=Potri.001G448800.3.p locus=Potri.001G448800 ID=Potri.001G448800.3.v4.1 annot-version=v4.1
ATGGCTGTTGGAGAGAATGTTAGTGTTGATTTGGGGAATTTTAGGGCGGAGTCAAGTGTATCATCATCATCAAATGATCAGGATCATCACATCCACAACA
ACAATGTTCTGATTCAGCCTCTGTACATGAAGGTTTCCCAGGTGGGCCATCATCAGGCTCCTCCTTATAATCATCATCATCAACAGAGATCAAATGGTGG
GGAGAGTTATAAGAGGGAGATTAGAGAGTTGCAGGAGTTATTCTCTAAGTTAAACCCCATGGCTGCTGAGTTTGTGCCTCCTTCACTTTCCAACAACAAT
AGTTTCGGTACAGTTAATGGGCTCAATGGAGTTAATGGGGGTTTTTATGGCAACAGCAGCAGCAGCAACAACTTGGTTGTGAATGGAAATGGCTTCGACA
GAAATGGACAAGTTAATGGAAATGCTGCTAGAAGAAAGAAAAATTATGGTCAAGTGAAACGTAGAATCAGTAGCAGGACAAGTATGGCTCAACGGGAGGA
TACAGTTCGCAGGACTGTGTATGTCTCTGACATCGATCAACAGGTTACTGAAGAGCAACTAGCAGCTCTATTTATTAATTGTGGACAGGTTGTGGACTGT
CGTATCTGTGGCGATCCCAATTCTGTACTTCACTTTGCCTTTATTGAGTTCACTGATGAGGAAGGTGCACGGGCTGCCTTGAGCCTGTCTGGGACGATGC
TTGGATACTACCCTGTTAAAGTGCTACCCTCCAAAACAGCCATTGCACCAGTTAATCACACCTTTTTGCCCAGGAATGATGATGAGCGTGAAATGTGTGC
AAGAACTATCTACTGTACCAATATTGATAGGAATTTTACTCAATCAGATATAAAACTCTTTTTTGAATCACTCTGCGGGGAGGTTTATCGCCTGAGGCTG
CTAGGAGATCATCATCATCCAACTCGTATTGCTTTTGTGGAGTTTGTGATGGTAATTGCTTCACTCTTTATTTCTAATGCTGAACTTTCTCTCTCTTTCA
ACATCAAACAAATTTAG
AA sequence
>Potri.001G448800.3 pacid=42790748 polypeptide=Potri.001G448800.3.p locus=Potri.001G448800 ID=Potri.001G448800.3.v4.1 annot-version=v4.1
MAVGENVSVDLGNFRAESSVSSSSNDQDHHIHNNNVLIQPLYMKVSQVGHHQAPPYNHHHQQRSNGGESYKREIRELQELFSKLNPMAAEFVPPSLSNNN
SFGTVNGLNGVNGGFYGNSSSSNNLVVNGNGFDRNGQVNGNAARRKKNYGQVKRRISSRTSMAQREDTVRRTVYVSDIDQQVTEEQLAALFINCGQVVDC
RICGDPNSVLHFAFIEFTDEEGARAALSLSGTMLGYYPVKVLPSKTAIAPVNHTFLPRNDDEREMCARTIYCTNIDRNFTQSDIKLFFESLCGEVYRLRL
LGDHHHPTRIAFVEFVMVIASLFISNAELSLSFNIKQI
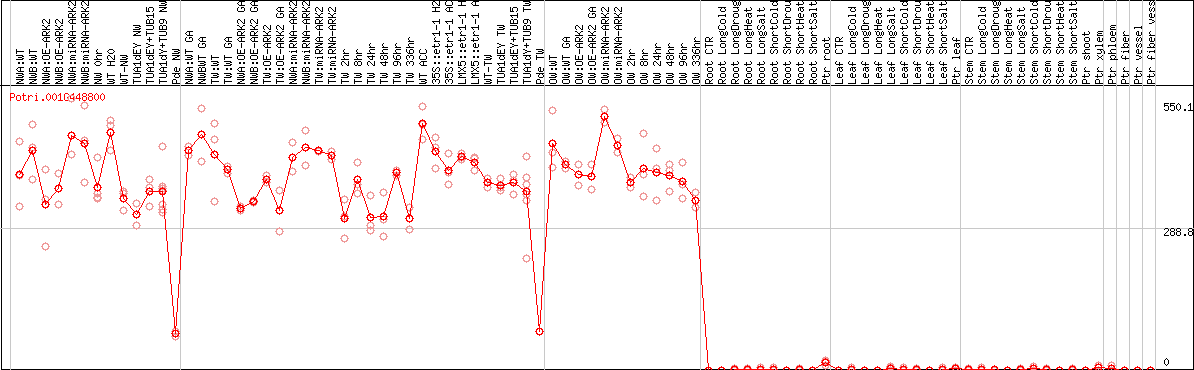
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G448800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.