Pt-VCS.1 (Potri.001G469500) [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-VCS.1 | |||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G469500.1 pacid=42793179 polypeptide=Potri.001G469500.1.p locus=Potri.001G469500 ID=Potri.001G469500.1.v4.1 annot-version=v4.1
ATGGCATCCCCAAATCCCCAACATCAACATCAACAACCGCAACCACAACCACAACCTCCTCCTTTTGACATGCACAAGTTCTTCATGCCCACATCCACAC
CACCACCACCTCAAAACCCTTCTTCCTCCCCTTCTCCTCCTCCTCCCAATCTCATCATGATTCCTCCACCCCAACAAATCCCTTCTTCTTACCCTCCATC
CACTGGGACCCACCACTTCCCTCACTACCCCAACTTCCCGTTTCCCTTCCCACCCCAACAACAACAATTTCAACCTCCTTACCCTCTCCAACAAAACCCT
AACCCTAGCAACCCACCTCCTCTTGCTAACCCCCAACGTTCCCTCTCTTATCCCACACCCCCTCTCACCCCAAACCAACAACTCCAAGACCGTTCTGGTG
CCGAAATCATGGCACTCCTCCGCACCCCTCAAAATCAAGAACCCCCACCAATCCCACCGCCGGCCGCGCCTGCGCAAGATTTTTCTGGAGCTGTTAATAA
TAGTAATAATATTTTTTCGGGGCCTATTAGGATGCCTAGCAGTACTAGTAGCAAGATGCCCAAGGGAAGGAGGGTAGCGGGTGAAAATGTAGTGTATGAT
GTGGATGTTAGATTGCAAGGAGAGGCGCAACCCCAGCTTGAAGTTACGCCTATAACTAAGTACTTTTCTGATCCTCAGCTGTGTTTAGGAAGGCAAATTG
CTGTTAATAGGACTTATATTTGTTATGGGTTGAAGCAAGGGAATATTAGGATTCTTAATATTAATACCGCGCTGAGGTCTTTGTTTCGGACACATTCTCA
GAGGGTCACTGACATGGCTTTTTTTGCCGAGGATGTTCATCTTTTGGCTAGTGCTGGCATAGATGGACGGATTAATGTCTGGAAGATCTCTGAAGGTCCG
GATGTGGAAGATAAGCCACAGATTACAGCACAGACAATTATTGCTGTCCAAATAGTTGGTGAGGGGGAAATTAAAAACCCAAGGGTTTGTTGGCATTGCT
ATAAACAGGAAATTTTGGTTGTTGGAGTTGGTAAACGAGTTCTGAGAATTGACACTAATAAAGTTGGAAAGGGCGAAGTCTATTCATCTGAGGCACCTCT
TCAATGTACTGTTGACAAGCTGATTGATGGGGTCCAATTCGTTGGTAAACATGATGGAGAAGTCACTGATTTGTCAATGTGCCAATGGATGACCACCCGT
TTGGTATCTGCTTCTATGGATGGCACGATAAAGATTTGGGAGGATCACAAGGCATCGCCCCTTGTGGTTTTGAGACCGCACGATAGCCAACCTGTTTATT
CAGCCACGTTCATGACTGCTACTGACCGGCCAGATCACATCATACTTGTCACAGCAGGGCCTCAGAACAGGGAAATGAAGATTTGGGCCTCAGCTAGTGA
AGAAGGCTGGCTCCTGCCTAATGATTCTGATTTGTTGAATTGCACCCAGACATTAGAGCTGAAGAGTTCAGCTGAACCTCGAGCTGAGGAGGCATTCTTC
AATCAAGTAGTAGCGTTGTCTCAAATGGGCCTTATTTTACTTGCAAATGCAAAGAGGAGTGCTATATATGCTGTGCATTTAGATTACGGTCCTAATCCAG
CATCAACTAGTATGGACTACATATCAGAGTTTACTGTCACTATGCCTATTTTGAGTTTAACTGGGACAAGTGACGTTGTGCATGGCCAATCTGTTGCTCA
AGTTTATTGCGTACAGACACAGGCAATTCAACAGTATACTTTGGAATTATGTCAGTGCCTGCCACCTTTGATGGAAAATGTGGGTTCAGAGAGGTCAGAT
TCCAGTGTTTTACATGGTGTGCCTAATGCTGATGGATATGCTGCCCTGGAGTCACATGGACGTAAATATTCTGATGTTCTTATGAGCTCCACATCAGTTG
ATGTGACCACTCCGCAACAAGATACTCCTGCTTCAAATATGGACCCCAGAACTATTGCTTCAAATATGTCCTCATCAACTAGTGATGCTGATATAGTTTG
TGTTCCATCACCTCCTCTTCCTTTGAGACCTAGATTATCTAGAGGTCTTGCTGAGTTTGCAGTTGGGGGAATGTTTGAGCCAAGTCCTGCATCCAGCAAC
CAAGGTTTTAATCAACCAGCTATCGATTATTCAGTTGACCAGCAAATGGACACAATTTGTTCAAATTTGTCCGATGTGCCTTCCTTGGATGGTGACTCGA
GGAATGAGAAAATTGTACAAGATAATTCTACCACTCTTAATCCTCCTGTTACATTTAAACACCCAACTCATCTAATAACCCCTTCTGAGATTTTAATGGG
TGCTTCATCCTCTGAAATTACCAATGTTAATGAGGGCAAGAGCGAGGTTGACTCAAATGTTCAAGATGTGGTTGTAAATAATGATGTTGTCAATGCTGAG
GTGGAAGTTAAAACAGTTGGTGAAACAATAGCCACTCGAGATGACGGATTTAGTCTTCGAGGGGAATCCAAAAGGCCTGTTTTTGAAAACAAGGAAAAAA
TCTTTTGCTCCCAGGCATCAGATCTTGGTGTTGAGATGACTAGAGAGTGTTGTGCATTACAGCCAGACAAAAATATTGAGGAAAGTGGACAAGTTGATGG
TGTTGGCATTAGCGAGTCTCTTGCCCCACCTTCTCATGCAGGTGGGGATGAAGTCCACGACTCAACAAAAGATATATCTGGGAAAGTTTCTGAGTCCACT
GTGTCGACAGTTGTTCCACTGTCAACAACTCCAAGTACAAAAGGGAAGAAGCAGAAGGGGAAAAATTCTCAAGCATCAGGTTCATCTTCTCAATCTCCAC
CTGCCTTTAATTCAACAGATTCTTTGAACGAACCTGTCGGGGCTTCAAATCTCCTCTCCGTGGAGGGTGCCTTTCCTCGGTTTTTGGCTATGCAGGAGGC
AATTAATCAGCTAGTGATCACACAAAAGGAAATGAATAAGCAGATGTCAAATATGGTTGCAATTCCTGTTTCTAAAGAATGTAGAAGACTAGAGACCACT
TTGGGGCGGAACATTGAGAAAGCCATCAAAGCCAATACTGATGCATTGTGGGCTCGATTTCAAGAAGAGAATGCAAAGAATGAGAAGTTATTGCGAGATC
GCACACAGCAGATAACAAGTTTGATCTTGAATTTTATCAATAAGGACTTGGCAGCCATGTTAGAGAAAGCACTGAAGAAGGAATTGGCTTCAGTTGGACA
AGGTGTAGTTCGCACAATATCTCCAGTCATCGAGAAAACAGTATCTTCTGTTATTGTCGAGTCTTTCCAGAGAGGAGTTGGTGACAAGGCAGTGAATCAA
CTGGAGAAATCAGTTAACTCAAAACTTGAGGCTACTGTTGCTAGGCAAATCCAAGCTCAGTTTCAAACTTCTGGCAAGCAAGCTCTCCAGGATTCCTTGA
AGGCTGGTCTGGAAGCGTCAGTAATCCCTGCCTTTGAGATGTCTTGCAAAGCTATGTTTGATCAAGTAGATGCTGCCTTTCGGAAAGGGATGGTCGAGCA
TACAGCTACAACCCAGCAGCATTTTGAATCTGCACATTCTTCATTGGCACTCACTTTAAGGGATTCCATCAACTCAGCATCATCATTGACGAAAACCTTG
AGTACAGAGTTGGCTGATAGTCAAAGACAGCTGTTAGCTATTGCAGCAAACTCAGGTGCAACAAATTCGTTGGCTACACAACCTAGCAATGGACCTTTGG
CTAGCTTCCATGAGAAGGTTGAAACACCTTTGGATCCAACAAAAGAGCTAACACGATTAATATCTGAACAACAATACGAGGCGGCATTCACTATTGTGCT
ACAAAGAAGTGATGTGGCCATGGTATCCTGGCTGTGTTGTCAGGTTGATCTGCAGGGTATCATGGCTATGTCTCCTCCTCCACTAAGCCAAAGAGTATTA
CTCGCACTTCTACAGCAGCTAGCCTGTGATATAAGCAAAGAGACTCCTCGAAAGCTTGAGTGGATGACAGTGGTGGCGGCTGCCATACAGCCAACAGACC
AAATGATTGCTTACTATGCACGGCCTATTGTCGAGCAAGTATGTGAGATTTTGAACCAGCTAAGGAGATCAGCTGGCGTTACTGGCGCTGACATCACAAG
CATCCGTATTCTCATGCATGTCGTTAATTACTTGCTGGTGACCTGTAAATGA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G469500.1 pacid=42793179 polypeptide=Potri.001G469500.1.p locus=Potri.001G469500 ID=Potri.001G469500.1.v4.1 annot-version=v4.1
MASPNPQHQHQQPQPQPQPPPFDMHKFFMPTSTPPPPQNPSSSPSPPPPNLIMIPPPQQIPSSYPPSTGTHHFPHYPNFPFPFPPQQQQFQPPYPLQQNP
NPSNPPPLANPQRSLSYPTPPLTPNQQLQDRSGAEIMALLRTPQNQEPPPIPPPAAPAQDFSGAVNNSNNIFSGPIRMPSSTSSKMPKGRRVAGENVVYD
VDVRLQGEAQPQLEVTPITKYFSDPQLCLGRQIAVNRTYICYGLKQGNIRILNINTALRSLFRTHSQRVTDMAFFAEDVHLLASAGIDGRINVWKISEGP
DVEDKPQITAQTIIAVQIVGEGEIKNPRVCWHCYKQEILVVGVGKRVLRIDTNKVGKGEVYSSEAPLQCTVDKLIDGVQFVGKHDGEVTDLSMCQWMTTR
LVSASMDGTIKIWEDHKASPLVVLRPHDSQPVYSATFMTATDRPDHIILVTAGPQNREMKIWASASEEGWLLPNDSDLLNCTQTLELKSSAEPRAEEAFF
NQVVALSQMGLILLANAKRSAIYAVHLDYGPNPASTSMDYISEFTVTMPILSLTGTSDVVHGQSVAQVYCVQTQAIQQYTLELCQCLPPLMENVGSERSD
SSVLHGVPNADGYAALESHGRKYSDVLMSSTSVDVTTPQQDTPASNMDPRTIASNMSSSTSDADIVCVPSPPLPLRPRLSRGLAEFAVGGMFEPSPASSN
QGFNQPAIDYSVDQQMDTICSNLSDVPSLDGDSRNEKIVQDNSTTLNPPVTFKHPTHLITPSEILMGASSSEITNVNEGKSEVDSNVQDVVVNNDVVNAE
VEVKTVGETIATRDDGFSLRGESKRPVFENKEKIFCSQASDLGVEMTRECCALQPDKNIEESGQVDGVGISESLAPPSHAGGDEVHDSTKDISGKVSEST
VSTVVPLSTTPSTKGKKQKGKNSQASGSSSQSPPAFNSTDSLNEPVGASNLLSVEGAFPRFLAMQEAINQLVITQKEMNKQMSNMVAIPVSKECRRLETT
LGRNIEKAIKANTDALWARFQEENAKNEKLLRDRTQQITSLILNFINKDLAAMLEKALKKELASVGQGVVRTISPVIEKTVSSVIVESFQRGVGDKAVNQ
LEKSVNSKLEATVARQIQAQFQTSGKQALQDSLKAGLEASVIPAFEMSCKAMFDQVDAAFRKGMVEHTATTQQHFESAHSSLALTLRDSINSASSLTKTL
STELADSQRQLLAIAANSGATNSLATQPSNGPLASFHEKVETPLDPTKELTRLISEQQYEAAFTIVLQRSDVAMVSWLCCQVDLQGIMAMSPPPLSQRVL
LALLQQLACDISKETPRKLEWMTVVAAAIQPTDQMIAYYARPIVEQVCEILNQLRRSAGVTGADITSIRILMHVVNYLLVTCK
|
DESeq2's median of ratios [POPLAR]
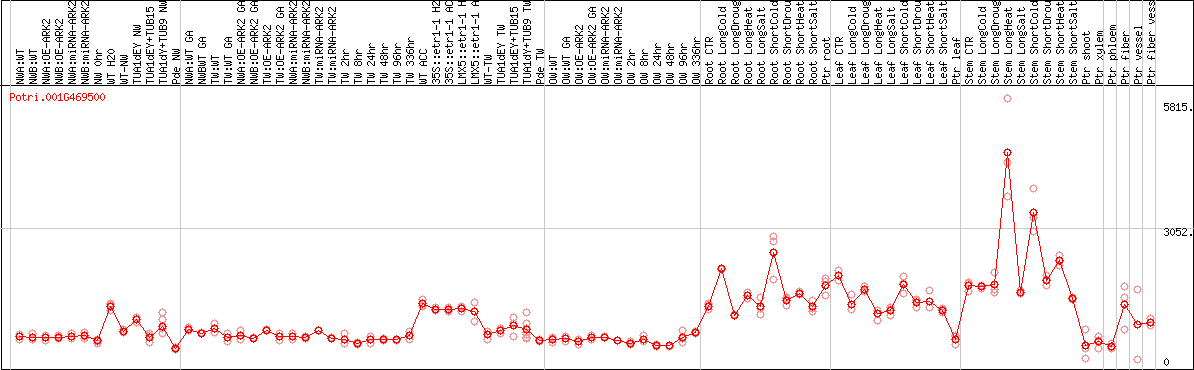
Coexpressed genes
Potri.001G469500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
