Potri.001G472400 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info | |||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G472400.1 pacid=42789023 polypeptide=Potri.001G472400.1.p locus=Potri.001G472400 ID=Potri.001G472400.1.v4.1 annot-version=v4.1
ATGTTATCCATTGAAAACCCTCCAGTACCAGATCCCTCATGTTCTTCTTCCCAGCTGAATAGTAGTGATGAGAGGGCTTACCAGCTTCCCACCAGTACTA
ATAATAAGCTTCCTTCTCCTAATTTGTCAGAGGTAGTAGTAGTAAATCTGCCAAACACAAACCCATCTCTTCATCATCATCATCATACCCCACTTCCTAA
TTTCTCCATAAGAGATTATGTATTTAAAGCCAGGAGCAAGGATATCAAGAATAGCTGGCCTTTTTCTCAAAACAATTTGCAACTTTGTTTGAAACATGGC
GTGAAGGACGTGTTGCCAAAATTTCAGCCTCATGATACTGTAAGAAACCAGTTCTTTAAGAGATGTACAGGTGAAACCAGTTCAGTTGAGAAGGAAAACA
ATTTTGACAAGGAGGCTTCGAGGCCGGACAATCGTGTGCTGCTAGACTCGTCTGATGATGCTCAATTAAATAACAAGCTAGCAGAATCCTGTGTAGACAT
TAGTTCATGTAGATCTGGAGAAGAAAATGATTTCCCATCGACAACAACAAGTGAGATAAACTCAGTTCCTGATAACAGGCAACGTAGATCACCATTAGAA
ACTCAATCTTTGGCCAAAGCCGCTGTTGAAGTTGAAGCTCCTGTGACCCACAAGACTGAAAGCACCTCCAGGCCGTTGGCTAAGAAGTGCAGGTTGATTG
TGAAATTTGGTGGCAGCTCTGATCGTAGCTCAGCTGAAGATATTGCTTCTAATTGTACTACTACATCAGAAACAATGGCTTCAAAAGTTTGTCCTGTTTG
CAAAACTTTCTCATCCTCATCAAACACCACCTTGAATGCTCACATTGATCAGTGCCTTTCTGTGGAGTCAACACCCAAGTGGACATCAGATTCTAAGCCA
ACAAGGTATAGAATTAAGCCGAGAAAGAATCGGTTGATGGTGGATATTTACGCTACAGCTCAATATTGCACGTTAGAAGACCTTGATCGAAGAAATGGTA
CGAGCTGGGCCACCATGTCAAGCTTGCCTGCCCAAGAGACTGAGAAGAGTGACGCACCTAATGAAGGGAAAAAGCAAAGGGTGTCCCCAATTCATCCTGA
GGATGCTGCTGATGTAGGTCCAGTTTATATTGATGCAGATGGTACCAAAGTTAGGATTTTATCTCAGTTTAATGATACACCACCAGTAGAAAAAGTAAGC
GAGGATATTGGTGCAAGAAGGGAGGATATTGGAGCAAAGAAATCTTTGAAAGGAGGCAAAGCAAGCAAATACATTTCAAAGAAAAAGAAAAAACGCTTAG
CACAAAAGCATCAAAAGTATCTAAGACTTGCATCTCAGAGCAAAAAGATTTTCTTCCACAAGGCGCCTTGCGCTCAGATTTCTGGAGGTCAAGAAGAATT
TAACGGAGAGGGGAAAAGTTGTGAGAAAGAACAACGGATGTTGAAGCAAATCAATCCCAATGATGGCGGGACTTTAAGACCATGGATTTGCTCCAAACGA
AGAGGTTTTCCAAAGAAGATTCCAACTCAAGAGGATCATCAACCTGTGAGATGTAAATGGCATTTGGCTCAAGACTTGCTGGTTGAGAATGATAGTCTTT
CAGAGAGGAGCCGCACTCAGAAATCTGTCATTTTGTCTGATAACCCAATATCTTCTCATAGAAACATTGAGAGGACGGAGAAGCCATTTCATAAAGACCA
AGTAAATGAGAGTATGGAGCACTCTCCTGGAAGAAAGATGGTAACAAATCTCCCAGTGAGAGACAGGATTAATGGCAAAGTGGATAAGTTGTTTCCACCA
ATGAAGTTGAGCAAAGATGGCACTTCTATCCGTGATACCTGTTTGCTAAGACCTCCAGACTCTCCCAGGATAAAGGTCTCTTCACTAACCAAGAAGACAA
TCTATACTGATGCTGATACAAGTAATAATTCAGATACCTCTCCCATTGCTAGCACAAAATCATCCCGGAGTTCTCGTACAGTTGTATCTAAAGCATTGAG
ATTCTGCTCCTTTAGGAAAAGTGTACTGTCTGTCAGCAGCCAATCTTCTGTGACTGAATCCAGGCCCAGCGAAGTTAGGAAATGGTCAACCCTTGACAAG
TCTGAAGACCCTTCGACAACAGAAATAGATGAGGATGCAATGGGCAGGCATTCTGAGGTTGATGAACAGTATGATTTGATGCAGGATCATACAGAAAATG
TACTTGAAAGAGAAGAAATAACAGATGAGGTGTCTCTTGGTGGAAGCTCCATCCGGGAAACTAGGCAAGAAAAAAGATTGTCATGTAGTTCTGAAAGGCT
GGAAGTCTTGTCTTTGAGGAGCTCAAAATCTACACCTCGCTATGGTCATGATGAGGAGATAAATGTAGATTCTTCAGCTAGATTTGATGATGATGATTAT
TTGCGCAAAATTGATCCCCTAGAATCTCCTGGAACACAAGTTCGAATACATGAGGACATTGTTGTTGAACCATCTTCCAAGACTTTGGATGGGAGGACGA
GCACCTCTGGTACGAGTAAATCCGTTGACACTGGATTCTATGAGCTAGGTGTTTCTTCAAAGGTCCCATCAAAATGTCTTCGGTCTATTGAACATTATGA
AGGACTTTCAAGGCAAAATGATGGGTCAACTGGTCCAACTGAGCCAGGTTTTGTTCACGACCAAGGGATGTTTTCTGCTGCTGAAGCTGGCAATGGCATG
ATGGGCCATAATGCTGATATGAGGGTGGTGGAATTGGATTCTGAAGCTGCGAAAGTAGACTCTTTTCCTGAGGTTGATCCCATTCTGATTCCAGGACCAC
CGGGGTCATTTTTGCCAAGTCCTAGAGATATGGGCTCTGAAGATTTTCAAGGAAACTCGTCATTGACTAGTAGCCAGGTTCAGTCATCTCCAGATCAGTA
TGATGTGATTGATGGAGATTCATCAGATTCCCCTCTATCTGCAGCATCGACCATCTCTAACTCCATGGCTGGCAGACCGGATTTCAATTATTCAGAACCG
CCATCATCAGCAGGACATTATGTTTTTCAAGACAATATGAGGTCAGGCTTAATTTCTGCTGGAATTGAACCTTTGGCACAGAATGCTGATGCAGTTCCAC
AAGCAGCAACTACAAGAGTTGAAAGAGCTACTTTCTTGGGAGAGCACGTGAAACTTGATGGGATCCCCATTGAGAAGGAATCTTTTGGCTTAAAAAATGA
TCAGCCATGCTGTTGCCAAAGAAAGGAGAGGTTTGCTGAGAGTGTTGCTCTAAATCACCAAGAATCACAGCTACTAAGGCGGAGGAAAACGCCATCGATG
ACATTTCCTTCTGTTAGCAAGCAAATGGGTTGCAATTCAAACCCAATGCCGATTAATTTGGATGTCAGGCCTGAATTAGTTTCCCTGAACAGTTACTCAG
CTTCAGGATCTGAAAAGATGGTTCTCCCACTCATCAAGCCCCCCGGAGATCCTATTCCTTTGAAGGACTCTCCCAATAATTCTGCAGTGAGATCCTTAGC
TCGTGCAGATGGTGATTCTGCTAGTCCATCAGCTTCTAATCCTATACTCAGACTGATGGGGAAGAACTTAATGGTGGTCAACAAAGATGATCATGTAGCC
ATGCCAATTGGGCAGGTCCAACCTTGTGCTCAAACAATCAATCGAACTCCTCACTTTCCAACAATTTCTGCAGTCTCTCCTGGCAATATTCAGAATCAGG
ACAGTCATTCATTTCATCGCGTCACCCCTCAAGGTTTTGCCATCTTTAGTCGGGATCCATATTACAAGACAGCAGTTCAACGTTTTGATGTTGGGTTGTC
CAACAGTTTTGGAAGCCACACTGATTCTAAGCTACCCCGGGCACCATCGCAGTTACCAGCAGGCATGTTTTGTGACCAACAAAATGATGGTGGTTTTGTA
ACATCCATGAAGCCTCAACAGTGCAAAGATGATTATAATTTCTCAAGTTCTCAGAACAGGCTAAAGAGAAGACTAGATGCGTTTCCTACGTGTACTATGC
AGAAAGCTACAGAAACTCCTGACCGCCAGTGCAAGCGTGCAGATTCATCTGCTCATCCTGTAAAAGAAATCATCATCATTGATGATGTTCCTGAAAGTCA
AACTGTTGTGATCAGTGACATTACAAGATACAATGAAGGCTGGAGGGAAAGACAAGCAGTTCCATCTGGTATCTCTGTTCCAACCATCCCAGTTTATAAC
ATGAGTAATGTGAATCCCTTTACTTGTTATCAGTCACAAGACCACCCTCCACTTGGTGGAACACCATTGTTGCACAATGGCAACTTTCATGCAACTGCCA
CTAGGCTAGTTAATACGAGTCCTGTTAGGTGGGGTTGTCCATCGGAAGGTCCCAGCGTGTTGCAACAGAATCCTTTCGTGGCTGCATCAAACTCATCTGG
TCATCCAAGATCAGCATCGCTGTATTATTCTCCAAGCTTTTCATAG
|
||||||||||||||||||||
|
AA sequence
|
>Potri.001G472400.1 pacid=42789023 polypeptide=Potri.001G472400.1.p locus=Potri.001G472400 ID=Potri.001G472400.1.v4.1 annot-version=v4.1
MLSIENPPVPDPSCSSSQLNSSDERAYQLPTSTNNKLPSPNLSEVVVVNLPNTNPSLHHHHHTPLPNFSIRDYVFKARSKDIKNSWPFSQNNLQLCLKHG
VKDVLPKFQPHDTVRNQFFKRCTGETSSVEKENNFDKEASRPDNRVLLDSSDDAQLNNKLAESCVDISSCRSGEENDFPSTTTSEINSVPDNRQRRSPLE
TQSLAKAAVEVEAPVTHKTESTSRPLAKKCRLIVKFGGSSDRSSAEDIASNCTTTSETMASKVCPVCKTFSSSSNTTLNAHIDQCLSVESTPKWTSDSKP
TRYRIKPRKNRLMVDIYATAQYCTLEDLDRRNGTSWATMSSLPAQETEKSDAPNEGKKQRVSPIHPEDAADVGPVYIDADGTKVRILSQFNDTPPVEKVS
EDIGARREDIGAKKSLKGGKASKYISKKKKKRLAQKHQKYLRLASQSKKIFFHKAPCAQISGGQEEFNGEGKSCEKEQRMLKQINPNDGGTLRPWICSKR
RGFPKKIPTQEDHQPVRCKWHLAQDLLVENDSLSERSRTQKSVILSDNPISSHRNIERTEKPFHKDQVNESMEHSPGRKMVTNLPVRDRINGKVDKLFPP
MKLSKDGTSIRDTCLLRPPDSPRIKVSSLTKKTIYTDADTSNNSDTSPIASTKSSRSSRTVVSKALRFCSFRKSVLSVSSQSSVTESRPSEVRKWSTLDK
SEDPSTTEIDEDAMGRHSEVDEQYDLMQDHTENVLEREEITDEVSLGGSSIRETRQEKRLSCSSERLEVLSLRSSKSTPRYGHDEEINVDSSARFDDDDY
LRKIDPLESPGTQVRIHEDIVVEPSSKTLDGRTSTSGTSKSVDTGFYELGVSSKVPSKCLRSIEHYEGLSRQNDGSTGPTEPGFVHDQGMFSAAEAGNGM
MGHNADMRVVELDSEAAKVDSFPEVDPILIPGPPGSFLPSPRDMGSEDFQGNSSLTSSQVQSSPDQYDVIDGDSSDSPLSAASTISNSMAGRPDFNYSEP
PSSAGHYVFQDNMRSGLISAGIEPLAQNADAVPQAATTRVERATFLGEHVKLDGIPIEKESFGLKNDQPCCCQRKERFAESVALNHQESQLLRRRKTPSM
TFPSVSKQMGCNSNPMPINLDVRPELVSLNSYSASGSEKMVLPLIKPPGDPIPLKDSPNNSAVRSLARADGDSASPSASNPILRLMGKNLMVVNKDDHVA
MPIGQVQPCAQTINRTPHFPTISAVSPGNIQNQDSHSFHRVTPQGFAIFSRDPYYKTAVQRFDVGLSNSFGSHTDSKLPRAPSQLPAGMFCDQQNDGGFV
TSMKPQQCKDDYNFSSSQNRLKRRLDAFPTCTMQKATETPDRQCKRADSSAHPVKEIIIIDDVPESQTVVISDITRYNEGWRERQAVPSGISVPTIPVYN
MSNVNPFTCYQSQDHPPLGGTPLLHNGNFHATATRLVNTSPVRWGCPSEGPSVLQQNPFVAASNSSGHPRSASLYYSPSFS
|
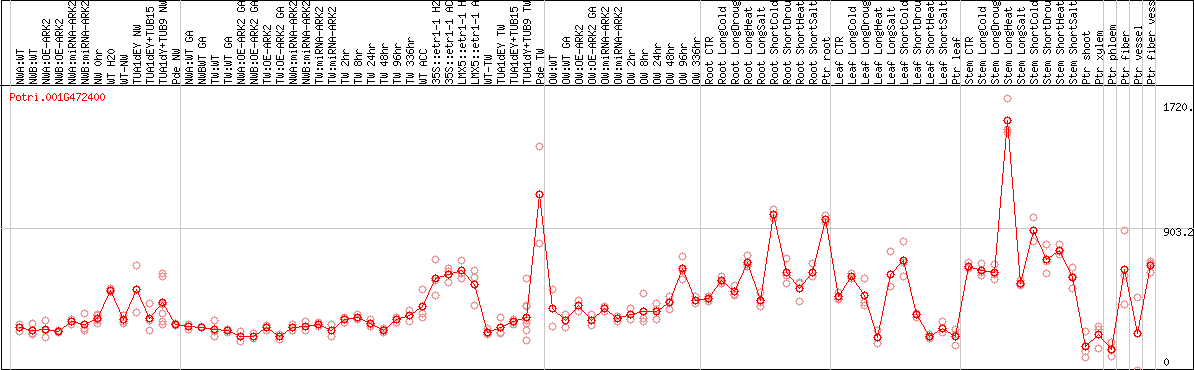
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G472400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
