Potri.001G472900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G472900.2 pacid=42787717 polypeptide=Potri.001G472900.2.p locus=Potri.001G472900 ID=Potri.001G472900.2.v4.1 annot-version=v4.1
ATGAAGAAACAATGGAGGATATCATCACAAAGGCCACCACACAAAGCCATGATTAGAATCTTTGGTTATGTGTTGCTGTTGCTGTTGTTATTTATGCCAT
CATCATCACAAACCAGAGAGTTGTCATCGCAGCAAAGTACCAATAATGAAGTTGTGGGATTATTAGCCTTCAAGAAATCCTCTGTTCAATCTGATCCAAA
TAATTTATTAGCCAACTGGTCTCCGAATTCTGCTACTCCATGTTCATGGTCAGGCATCTCTTGCTCTCTTGATTCCCATGTCACCACTCTCAACCTCACC
AACGGTGGTCTCATTGGAACCCTAAACCTCTATAACCTAACTGGGGCCTTGCCATCTCTCAAACATCTTTATTTGCAAGGCAATTCTTTCTCTGCTTCTG
ATCTTTCTGCTTCCTCTTCTTGTGTTCTTGAGAGTCTTGACTTGTCTTCAAACAACATCTCGGACCCTCTTCCAAGAAAGTCCTTTTTTGAGTCTTGCAA
TCACCTCTCTTATGTTAATCTCTCTCATAATTCCATTCCTGGTGGTTCCCTTCGATTTAGTCCTTCCTTGCTACAGCTTGACCTCTCTCGCAATACTATT
TCTGATTCTACCTGGCTGGCCTACTCTCTTTCCACCTGCCAGAACTTGAATCTTCTTAACTTCTCCGATAACAAGCTTGCTGGAAAACTTGCGGTAACTC
CCTTGTCCTGCAACAGTCTATCGGTTTTAGACCTCTCATACAATCTATTATCTGGGGAGATACCACCTAATTTTGTTGCAGACTCTCCATCTCTCAAATA
TTTGGATCTCTCACACAACAACTTCTCTGCCAACTTCTCCAGCCTTGATTTTGGGCACTATTGCAATCTTACTTGGCTCAGTTTGTCACAAAATAGGCTC
TCCGGCATTGGATTTCCATTAAGTTTGAGGAACTGTGTGCTTCTACAGACGCTTAACCTCTCCCGTAACGAGCTACAATTAAAGATTCCTGGTAACTTTT
TGGGGAGTTTCACGAATCTAAGGCAGTTGTCTCTGGCTCACAATCTGTTTTATGGTGACATTCCTCTTGAGTTGGGACAGACCTGTGGAACTCTACAGGA
ATTGGATCTATCAGCAAACAAACTCACTGGTGGCTTGCCGCTGACTTTTGCATCGTGTTCTTCTATGCAGAGTCTGAATCTTGGCAACAATCTACTCTCC
GGGGATTTCCTCACTACTGTTGTAAGCAATCTTCAAAGTCTGATATATCTGTATGTTCCATTCAACAACATTACTGGTACCGTGCCTCTGTCTCTCGCAA
ATTGTACTCATCTTCAAGTGCTTGACCTCAGTTCTAATGGCTTCACGGGGGATGTTCCTTCTAAGTTGTGTTCCTCCTCGAACCCAACCGCACTCCAAAA
GTTACTCCTAGCTGATAATTACCTATCAGGAAAAGTTCCATCAGAGCTTGGAAGCTGCAAAAACCTGAGGAGCATTGATCTTAGTTTCAACAGCTTGAAT
GGACCAATTCCCTTGGAGGTTTGGACCTTGCCAAATCTTTTGGACTTGGTTATGTGGGCAAACAATCTCACTGGTGAGATCCCGGAAGGCATTTGTGTTA
ATGGAGGAAACCTCGAGACTTTGATTCTCAATAACAATCTCATCACAGGAAGCATTCCTCAGTCCATTGGCAATTGCACTAATATGATTTGGGTTTCCCT
TTCCAGCAACCGGCTTACTGGAGAAATTCCTGCTGGCGTTGGAAATCTAGTTAATCTGGCTGTTCTCCAAATGGGTAACAATTCACTTACCGGGAAAATA
CCACCAGAGATTGGCAATTGTCGGAGCCTCATTTGGCTTGATTTGAACAGCAACAACCTAAGTGGTCCTCTTCCACCCGAGCTTGCAGACCAAGCTGGTC
TAGTTGTTCCTGGAATTGTATCCGGGAAGCAGTTTGCATTTGTAAGAAATGAGGGTGGGACATCATGCCGAGGAGCTGGGGGACTGGTTGAATTTCAGGG
GATCCGGGCAGAAAGGTTGGAAAATCTTCCTATGGTTCACTCTTGTCCAACAACAAGAATTTATTCTGGCATGACAGTTTATACATTCGTCACAAATGGC
AGCATGATATTCCTTGATCTTGCCTACAATTCGTTGTCAGGAACTATCCCTCAAAATTTTGGTTCGATGAGCTATTTGCAGGTCTTGAACTTGGGGCACA
ATAAGCTAACCGGAAATATTCCTGATAGTTTTGGAGGTTTGAAAGCAATTGGAGTTCTGGATCTCTCACACAATGATCTCCAAGGATTTCTACCAGGGTC
TTTAGGGACTCTCTCATTTCTTAGTGATCTTGATGTGTCGAACAACAACCTCACTGGTCCTATCCCTTCTGGAGGACAGCTAACTACTTTCCCCCAATCC
AGGTATGAAAACAATTCTGGCCTCTGCGGTGTACCATTGCCCCCTTGTAGCTCTGGAGGTCACCCCCAAAGTTTTACCACTGGGGGAAAGAAGCAATCTG
TGGAAGTAGGGGTGGTCATTGGCATCACATTTTTTGTCTTATGCCTCTTTGGTCTTACATTGGCTCTTTATCGAGTAAAGAGGTACCAGCGGAAGGAAGA
ACAGAGGGAGAAGTACATCGACAGCCTCCCAACTTCAGGTAGCAGCAGCTGGAAACTTTCAGGTGTTCCTGAGCCTTTGAGCATCAACATTGCCACTTTC
GAGAAACCTCTAAGAAAGTTGACTTTTGCCCATCTACTTGAAGCTACGAATGGTTTCAGTGCTGATAGCTTAATAGGGTCTGGTGGATTTGGAGAGGTAT
ACAAGGCACAACTAAAAGATGGATGTGTTGTTGCTATAAAGAAGCTGATTCATGTCACAGGTCAGGGGGACAGAGAGTTTATGGCAGAAATGGAAACTAT
TGGGAAGATCAAGCACCGCAACCTGGTTCCTTTGCTGGGTTACTGCAAAATTGGAGAGGAGAGGCTTCTTGTGTATGAATACATGAAATGGGGGAGTTTA
GAGTCTGTTCTTCATGACAGGTCCAAAGGAGGATGCTCAAGGCTTGATTGGGCAGCAAGAAAGAAGATTGCAATAGGCTCAGCAAGAGGACTTGCTTTCC
TACATCACAGCTGCATACCTCACATTATCCACCGTGACATGAAGTCTAGCAATGTTCTTTTAGATGAAAACTTTGAAGCTCGGGTTTCGGATTTTGGAAT
GGCAAGATTGGTGAACGCCCTTGACACACATCTTAGCGTGAGTACACTTGCAGGAACTCCAGGTTATGTGCCCCCTGAGTATTATCAGAGCTTTAGATGC
ACATCGAAAGGGGATGTCTACAGCTATGGTGTCATACTTCTGGAGCTCCTCTCAGGCAAGAAGCCAATAGATTCCGCTGAGTTTGGTGATGATAACAACC
TTGTTGGATGGGCAAAGCAGCTATATAGAGAGAAGAGAAGCAATGGGATACTGGACCCTGAGTTAATGACACAAAAATCTGGAGAAGCTGAATTATACCA
GTATTTGAGAATCGCATTCGAGTGCCTTGATGACAGGCCTTTCAGACGGCCGACCATGATACAGGTTATGGCTATGTTCAAAGAGCTTCAGGTTGACTCT
GAAAGCGATATCCTAGATGGTTTCTCTTTGAAAGATGCAAGCATTGATGAATTAAGGGAGAAAGAGTCTAGCTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G472900.2 pacid=42787717 polypeptide=Potri.001G472900.2.p locus=Potri.001G472900 ID=Potri.001G472900.2.v4.1 annot-version=v4.1
MKKQWRISSQRPPHKAMIRIFGYVLLLLLLFMPSSSQTRELSSQQSTNNEVVGLLAFKKSSVQSDPNNLLANWSPNSATPCSWSGISCSLDSHVTTLNLT
NGGLIGTLNLYNLTGALPSLKHLYLQGNSFSASDLSASSSCVLESLDLSSNNISDPLPRKSFFESCNHLSYVNLSHNSIPGGSLRFSPSLLQLDLSRNTI
SDSTWLAYSLSTCQNLNLLNFSDNKLAGKLAVTPLSCNSLSVLDLSYNLLSGEIPPNFVADSPSLKYLDLSHNNFSANFSSLDFGHYCNLTWLSLSQNRL
SGIGFPLSLRNCVLLQTLNLSRNELQLKIPGNFLGSFTNLRQLSLAHNLFYGDIPLELGQTCGTLQELDLSANKLTGGLPLTFASCSSMQSLNLGNNLLS
GDFLTTVVSNLQSLIYLYVPFNNITGTVPLSLANCTHLQVLDLSSNGFTGDVPSKLCSSSNPTALQKLLLADNYLSGKVPSELGSCKNLRSIDLSFNSLN
GPIPLEVWTLPNLLDLVMWANNLTGEIPEGICVNGGNLETLILNNNLITGSIPQSIGNCTNMIWVSLSSNRLTGEIPAGVGNLVNLAVLQMGNNSLTGKI
PPEIGNCRSLIWLDLNSNNLSGPLPPELADQAGLVVPGIVSGKQFAFVRNEGGTSCRGAGGLVEFQGIRAERLENLPMVHSCPTTRIYSGMTVYTFVTNG
SMIFLDLAYNSLSGTIPQNFGSMSYLQVLNLGHNKLTGNIPDSFGGLKAIGVLDLSHNDLQGFLPGSLGTLSFLSDLDVSNNNLTGPIPSGGQLTTFPQS
RYENNSGLCGVPLPPCSSGGHPQSFTTGGKKQSVEVGVVIGITFFVLCLFGLTLALYRVKRYQRKEEQREKYIDSLPTSGSSSWKLSGVPEPLSINIATF
EKPLRKLTFAHLLEATNGFSADSLIGSGGFGEVYKAQLKDGCVVAIKKLIHVTGQGDREFMAEMETIGKIKHRNLVPLLGYCKIGEERLLVYEYMKWGSL
ESVLHDRSKGGCSRLDWAARKKIAIGSARGLAFLHHSCIPHIIHRDMKSSNVLLDENFEARVSDFGMARLVNALDTHLSVSTLAGTPGYVPPEYYQSFRC
TSKGDVYSYGVILLELLSGKKPIDSAEFGDDNNLVGWAKQLYREKRSNGILDPELMTQKSGEAELYQYLRIAFECLDDRPFRRPTMIQVMAMFKELQVDS
ESDILDGFSLKDASIDELREKESS
|
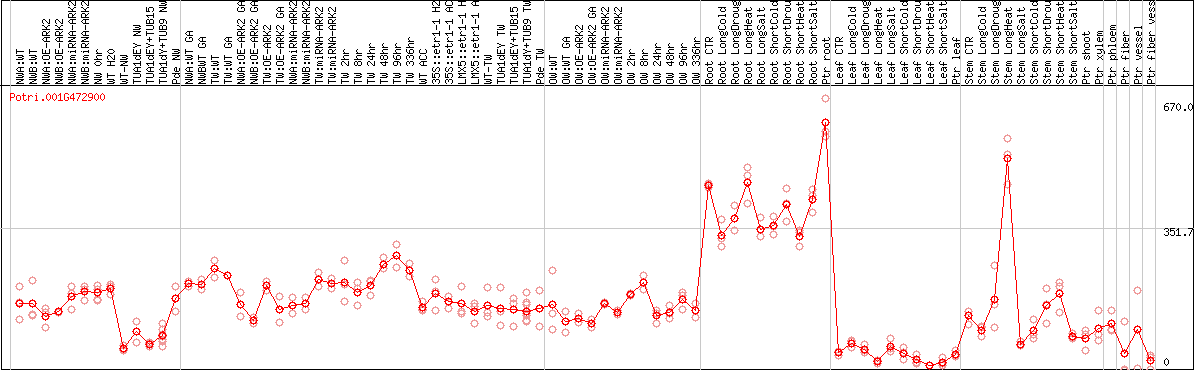
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G472900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
