External link
Symbol
Pt-SKP1.2
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G42190 249 / 6e-86
SKP1B, ASK2
Arabidopsis SKP-like 2, E3 ubiquitin ligase SCF complex subunit SKP1/ASK1 family protein (.1)
AT1G75950 244 / 3e-84
UIP1, SKP1A, ATSKP1, ASK1, SKP1
UFO INTERACTING PROTEIN 1, ARABIDOPSIS SKP1 HOMOLOGUE 1, S phase kinase-associated protein 1 (.1)
AT1G20140 216 / 3e-73
ASK4
SKP1-like 4 (.1)
AT2G25700 205 / 8e-69
ASK3
SKP1-like 3 (.1)
AT4G34210 204 / 1e-68
ASK11
SKP1-like 11 (.1)
AT4G34470 200 / 7e-67
ASK12
SKP1-like 12 (.1)
AT3G60010 191 / 4e-63
ASK13
SKP1-like 13 (.1)
AT3G21860 181 / 2e-59
ASK10
SKP1-like 10 (.1)
AT3G21850 181 / 3e-59
ASK9
SKP1-like 9 (.1)
AT2G03170 178 / 2e-58
ASK14
SKP1-like 14 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017282
276 / 1e-96
AT5G42190 259 / 6e-90
Arabidopsis SKP-like 2, E3 ubiquitin ligase SCF complex subunit SKP1/ASK1 family protein (.1)
Lus10005610
274 / 7e-96
AT5G42190 261 / 2e-90
Arabidopsis SKP-like 2, E3 ubiquitin ligase SCF complex subunit SKP1/ASK1 family protein (.1)
Lus10017281
273 / 2e-95
AT5G42190 259 / 4e-90
Arabidopsis SKP-like 2, E3 ubiquitin ligase SCF complex subunit SKP1/ASK1 family protein (.1)
Lus10013575
271 / 1e-94
AT1G75950 255 / 2e-88
UFO INTERACTING PROTEIN 1, ARABIDOPSIS SKP1 HOMOLOGUE 1, S phase kinase-associated protein 1 (.1)
Lus10019794
253 / 2e-87
AT1G75950 241 / 8e-83
UFO INTERACTING PROTEIN 1, ARABIDOPSIS SKP1 HOMOLOGUE 1, S phase kinase-associated protein 1 (.1)
Lus10014115
246 / 3e-79
AT1G68010 700 / 0.0
hydroxypyruvate reductase (.1.2)
Lus10041874
216 / 2e-73
AT4G34210 201 / 3e-67
SKP1-like 11 (.1)
Lus10019432
192 / 1e-63
AT1G20140 186 / 2e-61
SKP1-like 4 (.1)
Lus10002803
172 / 2e-55
AT1G20140 189 / 3e-62
SKP1-like 4 (.1)
Lus10031747
97 / 7e-26
AT2G03190 104 / 7e-29
SKP1-like 16 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01466
Skp1
Skp1 family, dimerisation domain
CL0033
POZ
PF03931
Skp1_POZ
Skp1 family, tetramerisation domain
Representative CDS sequence
>Potri.002G018700.1 pacid=42776912 polypeptide=Potri.002G018700.1.p locus=Potri.002G018700 ID=Potri.002G018700.1.v4.1 annot-version=v4.1
ATGTCGTCGAGCAGGAAATTTACTCTGAAAAGCTCAGACGGAGAGGCTTTTGAGGTAGACGAAGCTGTAGCGCTTGAATCGCAGACTATAAAACACATGA
TCGAAGAAGATTGCGCTGATAACGCTATTCCTTTGCCTAACGTCACAAGCAAGATTCTTTCTAAGGTGATCGAGTACTGCAAAAAGCACGTTGAGACTCC
TAAATCGGACGATCGCCCTTCTTCCGCTGATGATGATCTTAAATCTTGGGATGCTGAGTTCGTCAAAGTTGATCAGGCCACTCTCTTTGATCTCATCCTG
GCTGCCAATTACTTGAACATCAAGAACTTGCTGGACCTAACTTGTCAGACAGTGGCTGATATGATCAAGGGCAAGACTCCAGAAGAGATCCGTAAGACAT
TCAACATTAAGAATGACTTTACACCAGAGGAGGAGGAGGAGGTTAGGAGAGAGAACCAGTGGGCTTTTGAGTAG
AA sequence
>Potri.002G018700.1 pacid=42776912 polypeptide=Potri.002G018700.1.p locus=Potri.002G018700 ID=Potri.002G018700.1.v4.1 annot-version=v4.1
MSSSRKFTLKSSDGEAFEVDEAVALESQTIKHMIEEDCADNAIPLPNVTSKILSKVIEYCKKHVETPKSDDRPSSADDDLKSWDAEFVKVDQATLFDLIL
AANYLNIKNLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPEEEEEVRRENQWAFE
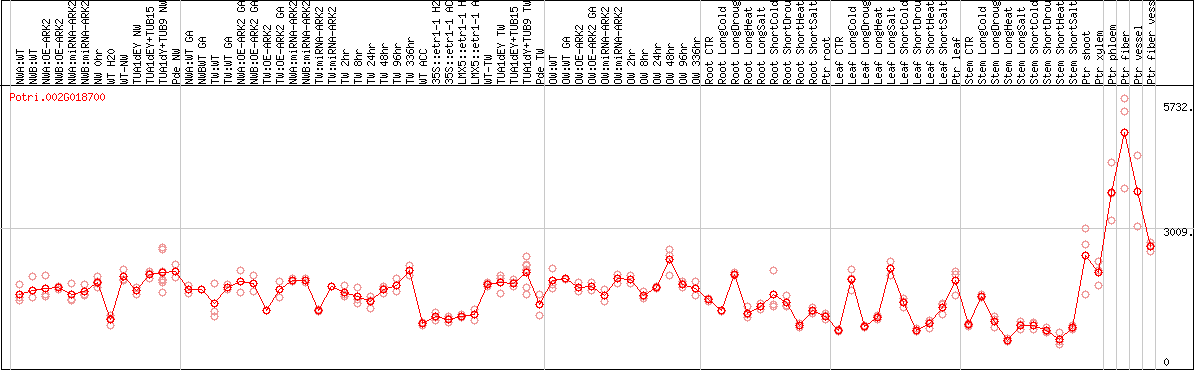
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G018700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.