External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G35070 148 / 6e-43
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
AT1G45976 103 / 1e-25
SBP1
S-ribonuclease binding protein 1 (.1)
AT3G12920 97 / 7e-23
BRG3
BOI-related gene 3, SBP (S-ribonuclease binding protein) family protein (.1)
AT1G79110 90 / 2e-20
BRG2
BOI-related gene 2, zinc ion binding (.1.2)
AT1G10650 90 / 2e-20
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
AT5G45100 87 / 9e-20
BRG1
BOI-related gene 1, SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
AT1G60610 85 / 1e-18
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2), SBP (S-ribonuclease binding protein) family protein (.3)
AT5G47050 81 / 1e-17
SBP (S-ribonuclease binding protein) family protein (.1)
AT1G32740 81 / 2e-17
SBP (S-ribonuclease binding protein) family protein (.1)
AT4G17680 79 / 6e-17
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G242600
397 / 8e-141
AT4G35070 135 / 3e-38
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Potri.004G175500
171 / 5e-52
AT4G35070 142 / 1e-41
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Potri.009G135400
162 / 2e-48
AT4G35070 135 / 6e-39
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Potri.001G148600
102 / 6e-25
AT1G32740 221 / 4e-70
SBP (S-ribonuclease binding protein) family protein (.1)
Potri.001G439600
95 / 3e-22
AT3G12920 256 / 1e-83
BOI-related gene 3, SBP (S-ribonuclease binding protein) family protein (.1)
Potri.003G085800
94 / 7e-22
AT1G32740 220 / 9e-70
SBP (S-ribonuclease binding protein) family protein (.1)
Potri.014G027000
92 / 3e-21
AT1G45976 380 / 2e-132
S-ribonuclease binding protein 1 (.1)
Potri.002G124700
92 / 4e-21
AT1G45976 378 / 2e-131
S-ribonuclease binding protein 1 (.1)
Potri.010G042200
89 / 2e-20
AT1G10650 393 / 2e-137
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014118
189 / 1e-59
AT4G35070 171 / 7e-54
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Lus10019790
184 / 1e-57
AT4G35070 170 / 1e-53
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Lus10017276
163 / 4e-49
AT4G35070 122 / 2e-34
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Lus10013569
157 / 4e-46
AT4G35070 125 / 1e-34
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Lus10031780
94 / 4e-22
AT3G12920 260 / 3e-85
BOI-related gene 3, SBP (S-ribonuclease binding protein) family protein (.1)
Lus10001131
94 / 7e-22
AT1G45976 407 / 7e-143
S-ribonuclease binding protein 1 (.1)
Lus10030891
86 / 4e-19
AT1G10650 275 / 1e-92
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Lus10013027
86 / 8e-19
AT1G10650 382 / 3e-133
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Lus10031202
85 / 1e-18
AT3G12920 255 / 3e-83
BOI-related gene 3, SBP (S-ribonuclease binding protein) family protein (.1)
Lus10030597
83 / 4e-18
AT1G10650 380 / 3e-132
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0229
RING
PF13920
zf-C3HC4_3
Zinc finger, C3HC4 type (RING finger)
Representative CDS sequence
>Potri.002G019000.3 pacid=42776992 polypeptide=Potri.002G019000.3.p locus=Potri.002G019000 ID=Potri.002G019000.3.v4.1 annot-version=v4.1
ATGCCATTAAAGAGAGGGAGCGCCAACAGAGAGAGAAAAAGTTCTTTCAATCTTGTTGTTGTCATGGCTGTTCAAGCACAGTTGTATCCAGAAAGACTTG
GGCTTTTGCCTATGGGTGGCCAACAGGATTGCATACTTAACGATCATGTTTCAGAGTTCGATGCTGACTGGGGTTTTGCTTTTCAAGAGCCGCAACAACA
AAATCTCTTTCTTGATCAAAATAACTCTCAGAACTTTTGTTTTGATTGTAATATTGGGGCTTCTTCTTCTTCTTCTTCTTATTCAACAACTTGTGATAGT
TCTTTCTCCATGTTTCTTTCTCAATGTTTGGATGTTCAGCTTGACATGCAAAGGCGAGAAGTTGATTGCATGCTTCAATTACAGGCTGAGAGGCTAAGAT
TTGCTTTGCAACAGCAAAGAAAGCAGCAGTTGGGAATCATACTGAAGAGTGTAGAATCAAAGGTCTCCTCTTTGATAAGGCAAAACGAAGAGGACTTGGC
ACAAACAACAAAGAAAACCATGGAGCTTGAGGTCTGCCTGAGAAAAGTAGAGCAGGAGAGTGAACAATGGCAGAGACTAGCAAGAGAAAAGGAAGCTGTG
GTCGTTGATTTAAGCAACACGCTGGAACGAATCAGAGAACGACTGGTCACACCCAGCAATAAAGTCCAGGATGCAGAGTCCTTTTGTTGTGGCTCATGTG
ATATAGAACAAGTGGAAAGCCAAAAAAAGGTGGTTTGTAAAGGGTGCAATTCTAGGACCTCGTGCGTTATCTTTCTCCCTTGCAGGCACCTCTGCTCGTG
CAAATCTTGTGAAGCTTTTCTTGGCTCCTGCCCTGTCTGTAAATCAGTAAAGGAAGCAAGCATGGAAGTTTTCTGGGTCTAA
AA sequence
>Potri.002G019000.3 pacid=42776992 polypeptide=Potri.002G019000.3.p locus=Potri.002G019000 ID=Potri.002G019000.3.v4.1 annot-version=v4.1
MPLKRGSANRERKSSFNLVVVMAVQAQLYPERLGLLPMGGQQDCILNDHVSEFDADWGFAFQEPQQQNLFLDQNNSQNFCFDCNIGASSSSSSYSTTCDS
SFSMFLSQCLDVQLDMQRREVDCMLQLQAERLRFALQQQRKQQLGIILKSVESKVSSLIRQNEEDLAQTTKKTMELEVCLRKVEQESEQWQRLAREKEAV
VVDLSNTLERIRERLVTPSNKVQDAESFCCGSCDIEQVESQKKVVCKGCNSRTSCVIFLPCRHLCSCKSCEAFLGSCPVCKSVKEASMEVFWV
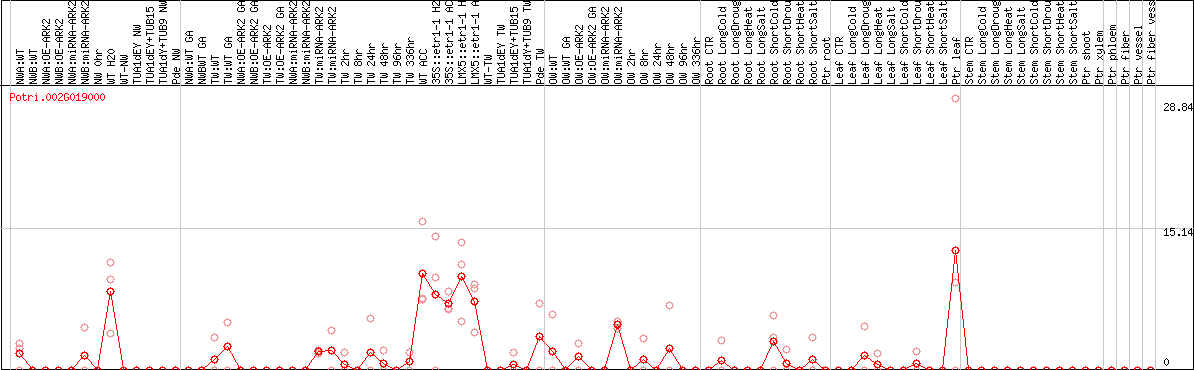
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G019000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.