External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G06060 283 / 1e-97
LisH and RanBPM domains containing protein (.1)
AT1G61150 82 / 3e-19
LisH and RanBPM domains containing protein (.1.2.3.4.5.6.7)
AT1G35470 70 / 8e-14
SPla/RYanodine receptor (SPRY) domain-containing protein (.1), SPla/RYanodine receptor (SPRY) domain-containing protein (.2)
AT4G09340 55 / 6e-09
SPla/RYanodine receptor (SPRY) domain-containing protein (.1)
AT5G09630 43 / 9e-05
LisH/CRA/RING-U-box domains-containing protein (.1)
AT4G09310 41 / 0.0003
SPla/RYanodine receptor (SPRY) domain-containing protein (.1)
AT4G09200 41 / 0.0003
SPla/RYanodine receptor (SPRY) domain-containing protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G233000
308 / 8e-108
AT1G06060 201 / 3e-65
LisH and RanBPM domains containing protein (.1)
Potri.011G046200
84 / 9e-20
AT1G61150 383 / 3e-136
LisH and RanBPM domains containing protein (.1.2.3.4.5.6.7)
Potri.004G037600
82 / 5e-19
AT1G61150 384 / 3e-137
LisH and RanBPM domains containing protein (.1.2.3.4.5.6.7)
Potri.013G108400
67 / 7e-13
AT1G35470 607 / 0.0
SPla/RYanodine receptor (SPRY) domain-containing protein (.1), SPla/RYanodine receptor (SPRY) domain-containing protein (.2)
Potri.019G080800
60 / 1e-10
AT1G35470 610 / 0.0
SPla/RYanodine receptor (SPRY) domain-containing protein (.1), SPla/RYanodine receptor (SPRY) domain-containing protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004152
320 / 2e-112
AT1G06060 275 / 2e-94
LisH and RanBPM domains containing protein (.1)
Lus10005633
318 / 1e-111
AT1G06060 275 / 2e-94
LisH and RanBPM domains containing protein (.1)
Lus10011219
81 / 2e-18
AT1G61150 392 / 6e-140
LisH and RanBPM domains containing protein (.1.2.3.4.5.6.7)
Lus10018464
81 / 3e-18
AT1G61150 392 / 9e-140
LisH and RanBPM domains containing protein (.1.2.3.4.5.6.7)
Lus10009451
69 / 1e-13
AT1G35470 601 / 0.0
SPla/RYanodine receptor (SPRY) domain-containing protein (.1), SPla/RYanodine receptor (SPRY) domain-containing protein (.2)
Lus10001544
67 / 1e-12
AT1G35470 604 / 0.0
SPla/RYanodine receptor (SPRY) domain-containing protein (.1), SPla/RYanodine receptor (SPRY) domain-containing protein (.2)
Lus10008254
42 / 0.0003
AT1G35470 309 / 3e-102
SPla/RYanodine receptor (SPRY) domain-containing protein (.1), SPla/RYanodine receptor (SPRY) domain-containing protein (.2)
Lus10002832
40 / 0.0007
AT4G37880 407 / 1e-141
LisH/CRA/RING-U-box domains-containing protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF10607
CLTH
CTLH/CRA C-terminal to LisH motif domain
Representative CDS sequence
>Potri.002G029700.6 pacid=42777073 polypeptide=Potri.002G029700.6.p locus=Potri.002G029700 ID=Potri.002G029700.6.v4.1 annot-version=v4.1
ATGGATGTTGATCCTCGTCACTACGAACAAATTGCCATCAAAGATAGTGATATCCACAATATTGTGCTATCCTATCTGGTGCACAATTGCTATGGAGAAA
CTCTGGAATCGTTCGTGGCTTGTTCCGGGATGCCTGAGCCTGCAGATTTTATCGAGGATATGGAGAAAAGAAAAGGAATTGTCCGTTGTGCTTTAGAGGG
GAATGCACTCAAGGCAGTTGAGCTGACAGAACAGGTGGCAGGTGACTTACTGGAGAATAACAAGGACTTGCATTTTGATCTTTTGAGCCTTCATTTTGCC
GATCTTGTTTGTGCTAAGAAGTGCACGGAAGCTTTGGAATTTGCTCAAAAGAAGTTGACACCATTTGGGAAGGAGAAGAAGTACGTTGAAAAACTTGAAG
ACTTTATGGCTCTGCTTGCTTACGAGGAACCTGAAAAATCACCAGTGTTTCATCTGCTTGGCTTGGAGTACCGTCAGCATGTAGCAGATAAATTGAATCG
GGCTATTCTTGCTCATACAAACTTGCCAAGCTACACAGCAATCGAGAGGCTGATACAACAGACCACAGTAGTCAGGCAAAGCTTAAATCAAGATCACGGT
AAGGATGGGATTCCACTATTTTCTCTGAAAGACTTTCTCAAAGGCTAG
AA sequence
>Potri.002G029700.6 pacid=42777073 polypeptide=Potri.002G029700.6.p locus=Potri.002G029700 ID=Potri.002G029700.6.v4.1 annot-version=v4.1
MDVDPRHYEQIAIKDSDIHNIVLSYLVHNCYGETLESFVACSGMPEPADFIEDMEKRKGIVRCALEGNALKAVELTEQVAGDLLENNKDLHFDLLSLHFA
DLVCAKKCTEALEFAQKKLTPFGKEKKYVEKLEDFMALLAYEEPEKSPVFHLLGLEYRQHVADKLNRAILAHTNLPSYTAIERLIQQTTVVRQSLNQDHG
KDGIPLFSLKDFLKG
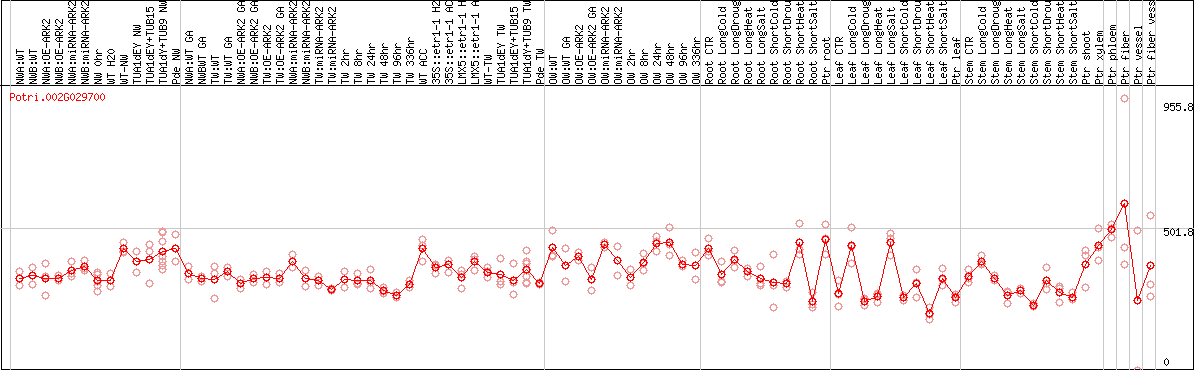
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G029700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.