External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G31190 333 / 1e-113
WXR1, RUS2
weak auxin response1, ROOT UV-B SENSITIVE 2, Protein of unknown function, DUF647 (.1.2)
AT5G49820 106 / 4e-26
RUS6, EMB1879
ROOT UV-B SENSITIVE 6, Protein of unknown function, DUF647 (.1)
AT1G13770 87 / 6e-20
RUS3
ROOT UV-B SENSITIVE 3, Protein of unknown function, DUF647 (.1.2)
AT3G45890 87 / 3e-19
RUS1
ROOT UVB SENSITIVE 1, Protein of unknown function, DUF647 (.1)
AT5G01510 69 / 3e-13
RUS5
ROOT UV-B SENSITIVE 5, Protein of unknown function, DUF647 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G224000
372 / 2e-129
AT2G31190 691 / 0.0
weak auxin response1, ROOT UV-B SENSITIVE 2, Protein of unknown function, DUF647 (.1.2)
Potri.001G193300
108 / 1e-26
AT3G45890 671 / 0.0
ROOT UVB SENSITIVE 1, Protein of unknown function, DUF647 (.1)
Potri.004G229500
95 / 3e-22
AT5G49820 674 / 0.0
ROOT UV-B SENSITIVE 6, Protein of unknown function, DUF647 (.1)
Potri.008G095700
80 / 6e-17
AT1G13770 624 / 0.0
ROOT UV-B SENSITIVE 3, Protein of unknown function, DUF647 (.1.2)
Potri.006G099700
54 / 4e-08
AT5G01510 520 / 0.0
ROOT UV-B SENSITIVE 5, Protein of unknown function, DUF647 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10003558
335 / 8e-115
AT2G31190 649 / 0.0
weak auxin response1, ROOT UV-B SENSITIVE 2, Protein of unknown function, DUF647 (.1.2)
Lus10033888
303 / 5e-102
AT2G31190 588 / 0.0
weak auxin response1, ROOT UV-B SENSITIVE 2, Protein of unknown function, DUF647 (.1.2)
Lus10038136
112 / 3e-28
AT5G49820 706 / 0.0
ROOT UV-B SENSITIVE 6, Protein of unknown function, DUF647 (.1)
Lus10010692
112 / 5e-28
AT5G49820 701 / 0.0
ROOT UV-B SENSITIVE 6, Protein of unknown function, DUF647 (.1)
Lus10001413
109 / 4e-27
AT3G45890 658 / 0.0
ROOT UVB SENSITIVE 1, Protein of unknown function, DUF647 (.1)
Lus10001047
108 / 8e-27
AT3G45890 653 / 0.0
ROOT UVB SENSITIVE 1, Protein of unknown function, DUF647 (.1)
Lus10036909
85 / 1e-18
AT1G13770 627 / 0.0
ROOT UV-B SENSITIVE 3, Protein of unknown function, DUF647 (.1.2)
Lus10022474
69 / 5e-13
AT2G23470 566 / 0.0
ROOT UV-B SENSITIVE 4, Protein of unknown function, DUF647 (.1)
Lus10016778
67 / 1e-12
AT2G23470 570 / 0.0
ROOT UV-B SENSITIVE 4, Protein of unknown function, DUF647 (.1)
Lus10037076
64 / 1e-11
AT1G13770 583 / 0.0
ROOT UV-B SENSITIVE 3, Protein of unknown function, DUF647 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04884
DUF647
Vitamin B6 photo-protection and homoeostasis
Representative CDS sequence
>Potri.002G038600.2 pacid=42778077 polypeptide=Potri.002G038601.1.p locus=Potri.002G038600 ID=Potri.002G038600.2.v4.1 annot-version=v4.1
ATGCAAACAACAGTGAATGTAGTTGATGATGCAAGACCAGTGTATCACAGAGTCGTTGAATCTTTCTCGAACAAGTTTTTTCCTTCAGGATATCAATGCA
GTCGCTACTATTTGCTGCAGGCTTGTGTCCTACCTGCACAAGCAGCTGCAGTGAGCTGGGTAAACTTCATTAATGATGGTAGACTATTAGCCCTAATTTG
CGTTTCTCTTCTGGTCTTCATTGATATTGTTGATGAGCAGATACTGAAGGATGAGATGCAGCATGTAGGGAAGCTTATATGTAGCAATTTGGGTGCTAGA
ATGGACTCAGAGCCAAAACATTGGAGAATCCTAGCTGATGTGCTTTACGATTTGGGCACCGGCTTAGAAGTTCTTTCTCCTTTATGTCCACATCTTTTTC
TTGAGGTGGCTGGACTTGGCAATTTTGCAAAGGGGATGGCAGTAGTTGCAGCAAGGGCAACAAGGTTGCCAATATATTCTTCATTTGCAAAAGAAGGCAA
TCTAAGTGACCTATTTGCAAAAGGGGAGGCCATCTCAACACTTTTCAATGTTCTTGGACCAGGAGTAGGAATTCAATTGGCATCCACTGTCTGCTCATCA
ATGCAGGGGAAGTTAGTTGTTGGGCCTCTTCTTTCTATTGTACACGTTTGTTGTGTTATTGAAGAAATACGAGCTACCCCTGTTAATACATTGAATCCAC
AGAGGACGGCAATGGTTGTGGCTGATTTTGTCAAGACAGGGAAAATATGA
AA sequence
>Potri.002G038600.2 pacid=42778077 polypeptide=Potri.002G038601.1.p locus=Potri.002G038600 ID=Potri.002G038600.2.v4.1 annot-version=v4.1
MQTTVNVVDDARPVYHRVVESFSNKFFPSGYQCSRYYLLQACVLPAQAAAVSWVNFINDGRLLALICVSLLVFIDIVDEQILKDEMQHVGKLICSNLGAR
MDSEPKHWRILADVLYDLGTGLEVLSPLCPHLFLEVAGLGNFAKGMAVVAARATRLPIYSSFAKEGNLSDLFAKGEAISTLFNVLGPGVGIQLASTVCSS
MQGKLVVGPLLSIVHVCCVIEEIRATPVNTLNPQRTAMVVADFVKTGKI
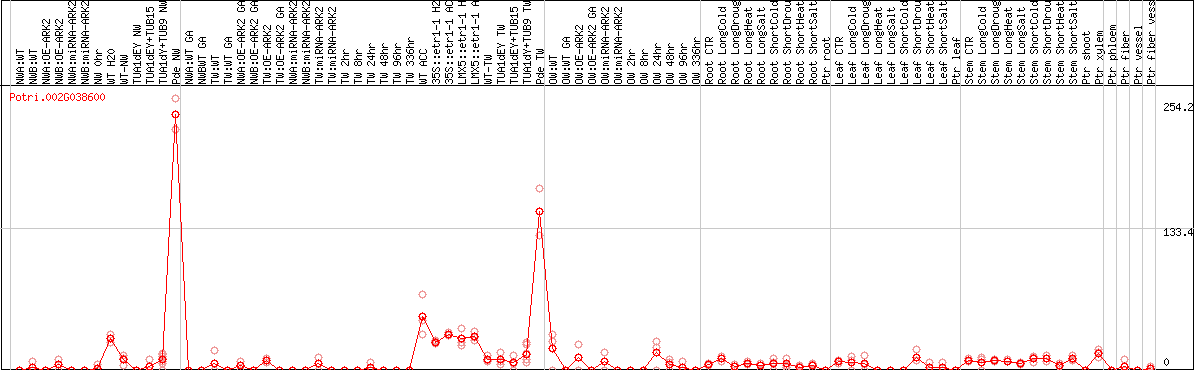
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G038600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.