Potri.002G047500 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G047500.1 pacid=42778621 polypeptide=Potri.002G047500.1.p locus=Potri.002G047500 ID=Potri.002G047500.1.v4.1 annot-version=v4.1
ATGACAAGCAGCAAGCCAACACATATCGACAGCAGCATATTGCTTGCCAGCACCATTAGCCAAGCCCAGAAGAGGTGGCGCATTGCTTATCTAGCTATCT
GCTCGGTCAGGGCAATGCTTTCTCTCGTAAGAGAGATGACATCAGAAACTAACAGCCATCAGTACTCTGGGATTTTGCACTCTGTTTCCTACACTGTGCT
TGATACGGAGCCAACCGGTTCCAAAAATCAGAAGAAAGGGCGTGAATCGACTTTCAATATATCTGACGATGATCAAATGAAATTCACTAAAATGGTGAAG
GAGAAAGACTTGGCATCTCTTAACAATCTTGGAGGAGTTGAAGGTGTCGCCACAGCTTTTGGGATAAACTCAAAAACTGGAATCACTGGTCATGACGAAG
AAGTCAGAAGGCGGCGTGAGATGTTCGGTCCCAACACTTACCATAAGCCACCGCCAAAAGGATTTTTATTTTTCGCGCTGGAAGCTTTTAGAGACACCAC
CATTCTTATTCTGCTGGTGTGCGCTGCTCTTGCCCTTGGTTTTGGCATCAAACAACACGGCGTAAAGGAGGGCTGGTACGAAGGTGGGAGTATCTTTGTT
GCAGTCTTTCTGGTCATTGTTGTCTCTGCGTCCAGCAACTTTAGACAGGAAACACAATTCGACAAGCTATCGAAGATAAGTAACAATATTAAGGTCGATG
TTCTTAGAAATGAAAGGCGACAGCAAATATCCATATTTGATATTGTTGTCGGAGATATTGTCTTCTTGAATATTGGTGATCAAATTCCTGCTGATGGCTT
GTTCTTGGATGGACACTCTTTGGAAGTGGACGAGTCAAGCATGACGGGAGAGAGTGACCATGTAGCAGTCAACACCCAAGAGAATCCCTTCTTATTTTCT
GGTTCGAAAATTGCTGATGGATATGCTCGAATGCTTGTGACATCTGTGGGGATGAACACGGCATGGGGGGAAATGATGAGCTCAATAACTCGTGACTCCA
ATGAACGAACACCACTACAAGCTCGGCTCGACAAACTAACTTCTTCTATTGGGAAGGTCGGCCTTTCAGTTGCTTTTGTAGTGCTTGTAGTCATGTTGGT
TCGTTACTTCACTGGAAATACAAAAGATGACAAGGGGAAGAAGGAGTACATCGGTAGCAGAACAGATACGGATGATGTGTTAAATGCAGTTGTACGCATT
GTTGCTGCTGCAGTCACTATTGTGGTTGTTGCAATTCCTGAAGGTTTGCCGTTGGCAGTCACCTTAACACTAGCTTACTCTATGAAGAGAATGATGGCTG
ACCAAGCAATGGTCAGAAAACTTTCAGCTTGTGAAACAATGGGCTCAGCCACAGTAATTTGCACTGACAAAACAGGCACACTGACACTAAATAAGATGAA
AGTAACCAAGTTCTGGCTTGGCCAGGAACCCATTGAGGAAGATTCTTACAAAACAATTGCTCCATCAATTCTTGAAGTATTCCACCAAGGAGTCAGTTTA
AACACGACTGGTAGTGTCTACAAATCTGCAACAGGATCTGTGCCTGAGTTTTCAGGCAGTCCAACTGAAAAGGCTATTCTTTCATGGGCCGTTTCTGAAC
TGGGCATGGATATGGAAAAATTGAAGGAAAGTTGTACCATTCTTCATGTTGAAACTTTCAACTCAGAGAAGAAACGAAGTGGGGTTTCAATAAGGAAGAA
GGCTGACAATACTGTTCATGTACATTGGAAAGGGGCTGCTGAGATGATATTAGCTTTGTGTTCAAGTTATTATGACAGTCGTGGCAGCATAAAGTCCATG
GACGAAGATGAAAGAAGTAAAATCGAGAAAATAATCCAAGGTATGGCGGCTAGTAGCCTTCGATGCATTGCGTTTGCTCATAAAAGGATCACAGAGGAAG
GAATGAAAGACAACGACGGGGAGCCACACCAGAGGCTACAAGAAGATGGTTTGACCTTGTTAGGGATAGTTGGGCTGAAGGATCCGTGTAGAATTGGAGC
CAAGAAAGCGGTGGAAATTTGCAAAGCTGCCGGAGTGAGCGTGAAAATGATAACTGGTGACAATATTTTCACAGCAAAAGCTATAGCAACAGAATGCGGC
ATATTAGAGCTCAAAAGCCAAGTTGACAGTGAAGAAGTGGTGGAAGGTGTTGTATTTCGTAACTACACGGATGAACAGAGAATGGAGAAGGTAGATAAGA
TCCGTGTGATGGCAAGGTCCTCCCCTTTCGACAAACTTCTAATGGTACAATGCCTGAGACAGAAAGGTCATGTTGTTGCAGTCACAGGAGATGGCACGAA
TGATGCACCTGCATTAAAAGAAGCAGATATAGGACTCTCTATGGGCATCCAAGGTACTGAGGTTGCCAAGGAGAGCTCAGATATTGTGATTTTGGATGAT
AACTTTACTTCTGTGGCTACTGTTTTGAGGTGGGGGCGGTGTGTATACAACAATATTCAGAAATTTATTCAGTTCCAACTAACTGTGAACGTTGCAGCTC
TTGTGATCAACTTTATTGCAGCAGTTTCAGCCGGTGAGGTTCCATTAACAGCCGTTCAGTTGTTGTGGGTAAACCTCATCATGGATACATTGGGAGCTCT
GGCCCTTGCTACAGAGCGACCCACTGATGAGCTAATGGAGATGTCACCAGTGGGGCGAACTGCGCCCCTCATCACTAATATAATGTGGAGAAATCTTCTA
GCTCAAGCTTTTTATCAGATAACAATATTATTGACATTACAGTTTGCAGGCGAGTCTATCTTCAATGTAAGTGCAGAGGTTAACGATACACTAATATTCA
ACACTTTTGTGCTTTGCCAAGTTTTCAACGAATTCAATGCAAGGAATATGGAAAAGCAGAACGTATTCAAAGGCATTCACAGGAACCATTTGTTTCTGGG
AATCATCGCCACTACTATTGTTCTTCAGGTGGTGATGGTGGAATTCCTGAAGAAGTTTGCAAGTACAGAGAGGTTGAACTGGTGGCAGTGGGTGACTTGT
ATTGCATTTGCTGCTGTGTCATGGCCAATTGGTTGGTTTGTGAAACTTATTCCAGTTTCAGGCAAACCATTCCTCAGTCACCTCAAGCGGCCGGTAGCAA
CATATAAAAGAGTCATGCATTCCATATACTTCAGAGGAACGTCTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G047500.1 pacid=42778621 polypeptide=Potri.002G047500.1.p locus=Potri.002G047500 ID=Potri.002G047500.1.v4.1 annot-version=v4.1
MTSSKPTHIDSSILLASTISQAQKRWRIAYLAICSVRAMLSLVREMTSETNSHQYSGILHSVSYTVLDTEPTGSKNQKKGRESTFNISDDDQMKFTKMVK
EKDLASLNNLGGVEGVATAFGINSKTGITGHDEEVRRRREMFGPNTYHKPPPKGFLFFALEAFRDTTILILLVCAALALGFGIKQHGVKEGWYEGGSIFV
AVFLVIVVSASSNFRQETQFDKLSKISNNIKVDVLRNERRQQISIFDIVVGDIVFLNIGDQIPADGLFLDGHSLEVDESSMTGESDHVAVNTQENPFLFS
GSKIADGYARMLVTSVGMNTAWGEMMSSITRDSNERTPLQARLDKLTSSIGKVGLSVAFVVLVVMLVRYFTGNTKDDKGKKEYIGSRTDTDDVLNAVVRI
VAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGSATVICTDKTGTLTLNKMKVTKFWLGQEPIEEDSYKTIAPSILEVFHQGVSL
NTTGSVYKSATGSVPEFSGSPTEKAILSWAVSELGMDMEKLKESCTILHVETFNSEKKRSGVSIRKKADNTVHVHWKGAAEMILALCSSYYDSRGSIKSM
DEDERSKIEKIIQGMAASSLRCIAFAHKRITEEGMKDNDGEPHQRLQEDGLTLLGIVGLKDPCRIGAKKAVEICKAAGVSVKMITGDNIFTAKAIATECG
ILELKSQVDSEEVVEGVVFRNYTDEQRMEKVDKIRVMARSSPFDKLLMVQCLRQKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDD
NFTSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSAGEVPLTAVQLLWVNLIMDTLGALALATERPTDELMEMSPVGRTAPLITNIMWRNLL
AQAFYQITILLTLQFAGESIFNVSAEVNDTLIFNTFVLCQVFNEFNARNMEKQNVFKGIHRNHLFLGIIATTIVLQVVMVEFLKKFASTERLNWWQWVTC
IAFAAVSWPIGWFVKLIPVSGKPFLSHLKRPVATYKRVMHSIYFRGTS
|
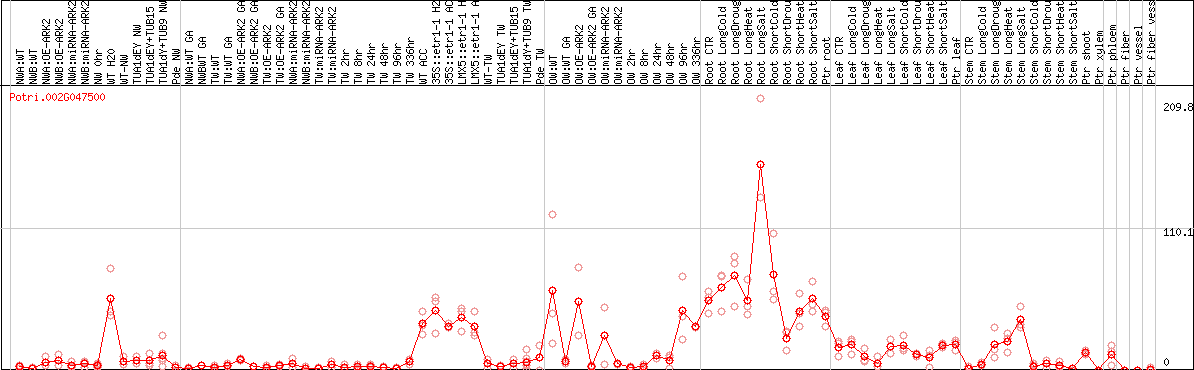
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G047500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
