Potri.002G049600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G049600.1 pacid=42779933 polypeptide=Potri.002G049600.1.p locus=Potri.002G049600 ID=Potri.002G049600.1.v4.1 annot-version=v4.1
ATGGCTGCCAGGTCGCAAGCAGATTCTAAAAGGAAGTATTCGTGGTGGTGGGACAGCCACATTAGCCCAAAAAACTCAAAATGGCTTCAAGAAAATCTTA
CAGATATGGATTCCAAAGTCAAACAGATGATCAAACTCATCGAAGAAGATGCAGATTCCTTTGCAAGGAGGGCAGAGATGTACTACAAGAAACGCCCAGA
GCTTATGAAATTGGTCGAAGAGTTTTATAGAGCGTATCGTGCCTTGGCTGAAAGATATGATCATGCCACCGGGGCCCTTCACCAGGCGCAACGAACAATG
GCAGAAGCATTTCCCAACCAAGCCCCCTTTATTCTGGGTGATGATTCCCCTGCAGGTTCTGCTACTGACTGTGATCCCCGTACACCTGATATGCCACCAA
TACGTGCACCTTTTGACCCTGATGAGCTGCAAAAGGATGCTTTGGGCGTCTCACCATCCCATGCTATCAATAGGAATGGAGCTTTTACTGAGAAATCAGA
TCCTGGAAGAAAGGGTTTGAAACAATTCAATGATCTTTTTGGGCTTGGAGATGGGATGGACAATGCTAAGTTTGCAGAAGGGAGGGTGAGAAAAGGCCTC
AATTTCCATGACCCAGAGGAGAAAGGACGAGGTGTGCAAAACAATGGCATCCATGATCTCAAGGCTCGAGCCCCATCTGAGTCTGAGCAAGTTAGTAAAG
CTGAGCTAGAAATTTTAAATTTAAAGAATGCACTTGCTAAACTAGAAGCCGAAAAAGAAGCTGGCCTGCTTCAGTATGAGCAAAGTTTGGAAAGATTGTC
TAAACTGGAGTCAGAAGTCTCTCGTGCAACGGAGGATTCCAGGGGACTTAATGAACGAGCTAGCAAAGCTGAAGCTGAAGTTCAAACTTTGAAGGAAGTC
CTTGCACAATTAGAGGCTGAAAAGGAATCTAGTTTTCTTCAATACCAGGGGTGTCTGGAAAAGATATCTAATCTGGAGAACAATCTCTCTCTTGTCCAAA
AGGATGCAGGAGAACTAAATGAGCGAGCCAGCAAAGCTGAAACTGAAGCTCGATCCCTAAAGCAAGACCTGTCTAGATTAGAAGCTGAAAAAATTGATGC
ACAGGTTCAGTACAGCCAATGTTTGGAAAAGATATCTGATCTAGAGGACAAATTACATAATGCACAGGAAGATGCCAAGAGGTTCAGTGAGCGTGCAGAT
GATGCTGAAAGGGAAATTGAAGCCTTGAAGCATGCTTTGACCAGGCTAACTGAAGAAAAGGAAGCTGCTGTCACTCAGTACCAGCAGTGCTTGGCAACAA
TTGTCAGTCTGGAGCATAAAATTGCTTGTTTTGAGGAGGAAGCCCGGAGGCTTAACTTGGTGATAGATAATGGGACTGTGAAGTTAAAGAGTTCTGAAGA
GAGGTGTCTTCTGTTAGAAAAATCAAACCAAACCATACACTCAGAGTTAGAGTCAGTGATGCAGAAAGTGGCGGCTCAAAGCAATGAGCTTACGGAAAAG
CAGAAGGAGTTGGGGAGACTTTGGGCTTGCGTCCAAGAAGAGCACTTGAGATTCATGGAGGCCGAAACTGCTTTTCAAACTCTACAGCATCTGCACTCTC
AATCTCAGGAAGAATTGAGATCTGTGGTGGCTCAGCTTCAGAACAGGGCTCAAATTTTGGAGGACCTGGAAGCTCGTAATCAGAGTTTAAAAGATGAAAT
GGAGCATGTCAATGTGGAGAACAAGAGCCTTAGTGACGTCAACTTATCTTCAGCTCTGACTATACAGAATCTGCAAGATGAGATTTCAAGCTTAAGGGAG
ACCATAAAGAAACTTGAAGCAGAGGTGGAGCTTCGAGTGGACCAAAGAAATGCTCTTCAGCAAGAGATTTACTGTCTAAAGGAGGAGCTAAATGAACTTA
ACCAGAAGCATCAGGCAATCATGAGGCAGGTGGAATCAGTTGGCTTCAGTCCAGAGTCCTTTGGATCATCTGTGAAGGATTTGAAGGATGTCAACATCAA
GCTGAAGGAGGTTTGTGAGAGAGACAGAACTGAAAAGGTAGCTCTTTTGGAAAAGTTGGAGAACATGGAGAAACTTATTGATAAAAATGCTCTTCTGGAG
AATTCTTTATCAGATTTGAATGTTGAGCTAGAAGGGGTCGGAGAGAAGTTAAAGGCACTAGAAGAATCCTGTCAATATCTTGTGGAAGAGAAATCCGTTC
TTGTTTCTGAGAAGGATTTGATGGCTTCTGAGTTGCAATTTGCTACTGATGATTTGGAGAAACTTACAGAGAAGAACCACATTCTGGAGAATTTCCTCCT
TGATGCAAATGCTGAACTTGAAGGGTTGAGGGAAAAATCAAAGAGCTTAGAAGATTTTTGCCTGCTGCTTGTGAATGAGAAGTCTGAGCTGGCCTCCATG
AAAGGAAGTTTATCTTCTCAGTTGGACATCAGTGAGAAAAGCTTGCAGGATTTGGAGAAGAATTATACAGAATTAGAAGAAAAATATTCCCATCTGGAGA
AGGAGAGACAATCCTCACTTCATGAAGTACAAGAATTACAGGTTCGTTTAGATGCTGAAAAGCAAGAACATGCTAATCTAGCCCAGTTGAGTGAGTCCCA
ATTGGCTGGTATGGCATCACAGATTTGTTTGCTACAAGAAGAAAGTCTGTGCAGAAAGAAAGAATATGAAAAGGAGCTGGACAAAGCTGTGAATGCAGAG
ATTGAGATCTTTATCTTGCAGAAATGTGCGCAAGAATTGGAAGAGAAGAATTCTTCCCTCTTGCTTGATCATCAGAAGCTCGTGGAGGCATCTAAACTGT
CAGAGAAACTGATCTCTGATATGAGACATGAAAATTGTGAACAACAAGAAGAGGTGAAATGCTTATCTGATAAAATTAAAACACTGAGGATGGGACTTTA
TCAGGTGTTGATGACTCTTGAACTTGATGCGAACCAGTGTGAAAATAAACCCAAGCAAGACCAAAAACTGCTGAACCATGTACTCAACAGGCTTCAAGAA
TCACAGGAGTTTCTCTTCAAGACACAAGATGAAAATCAGCGGTTGTTCACTGAGAACTCTGTTCTAGTTACTCTGCTTAGGCAACTGCAATTAGAGGTGG
AAAATCTTGTGAAGACTAAAAATATCCTCGATCAAGAATTGACAACCAGGTCTGAGCAATTCTTGGTTTTGCAGAATGAGAGCCAAGAGCTTTCAGGGAT
AAACGAAGAGATGAAATTGAAGTTAATAGAGGGAGATCGCAAAGAGGAGGCTTTGAAGGTTGAACTAAATAATCTCCATGTACAGCTGTCAGACTTGCAA
GGTGCTTTCCAGAACTTACAAGAGGAGAACTGCAAGGTTCTTGATGATCAGAGGTCCTTGATGAAATCATTTTCAGACCTGCAGATGGAGAAATGTGAAC
TGGAAGAGGAAAACTTCTGCATTCTTGTTGAAACTGTATCTCAAAGTACCCTTTCTCTAATTTTCAGGGATATCATCTGTGAAAAATCTGTGGAAATAAA
AAGCCTTGGTGTAAGCCTAGATAAACAGTGTCATGACAATAATGGTCTCAACGAGAAGGTGAAGACACTGGAGAAAGAGTTAGATAACTTCAGTGGCTTA
GAAGATGATAAGAGAGAGTTGCATAAAATGGTGGAGGATCTGAAGTGCAAGTATGATGAGGTTGAGGTGATACGATCAGATCAAGAAATGCAGATCATTA
AACTATTAGGAGATTATGATCAGAAAATTAAGGAGGCTGAAAACATTCGTGAAGTAAATCAGAAATTGGAGTCTGAAATTAGGAAATTGCATGAAGAATT
TCAAGAAGTTAAAGACAGGAAGGAAAATTTGAGTCATGAACTGGTCAAGGAAAGAAATGAGGTTGAACTACAAGAGTCTCAGGCTGTTGCATTGTTTGGT
GAGCTGCAGATATCTGCTGTCCGAGAAGCCTTGTTTGAAGGAAAGCTTCGCGAGCTCCTTAAAATATGTGAGAGCCTTGAAGATGGAAATTGCTCAAAAG
ACATGGAGATTGACCAGCTGAAAGAAAGAGTTAGCACTTTGGAAGGCGGAAATGCAGAACTAAAAGCTCTTGTGGCTGCATATTTGCCAGCTTTCATGTC
GTTGAGGGATTGTGTAACATCACTGGAGAAGCACACTCTTCCAGATGCGACACTTCATGAAGGTGACAGCAAGGAATCAAAGGATGCTGCATTGGTGGTG
CATGCCAAAGGCTTTCACCAAATGAGTGAAGGACAGAGTGGCATGGTGCCAGGTGGAACCTTGGACTTCCAGGATTTGCAAATGAGGATTAGAGCTATTG
AGAAAGAAATCATTGAGAAGGAGAGACTTGTTATGCTAGAAAATTTGAGTTATCATTCCAAACTCGATGCTGCAATAAGGCAGATTGAGGACTTGAAATC
TGGAAGCAGTGCACGTCAAAAAGGTGTCGAGACAAGGAGGTATGTTAAGCCAAAACCGGAGGATGGGGAACTGGGGGCCACACCAAGTGATGATCTGAGG
CGGCAGAAACGAACACATGAAATCTCAGAGGATGGGAATGAAGTGATGACAAAAGACATAATACTTGATCAGATATCAGAATGTTCATCCCATGGAATCA
GCAGGAGAGAAACTATGCAGGCTGATGAACAGATGCTTGAAATATGGGAAACTGCTGATCGGGATGACAGCATTGACCTGACTGTTGGAAAAACGCAGAA
GGTGACTGCTTCACAGAAGAAGAAAAAACACATTAGGCAGCATCCTTCTGCAGAATCAATGGTTGAGAAGGAGGTGGGTGTGGACAAATTAGAGATCTCA
AAAAGATTATCAGGGTCTCGTCAAGAAGGGAATGAGAGGAAAATTCTAGAAAGACTTGATTCTGATGCTCAAAAGTTGACAAATCTTCAAATAACAGTTC
AAGATTTGATGAGCAAGGTGGAGATTACAGAGAAGAGCGAGAAGGGAAAAGGTATTGAATATGATAATGTGAAGGAGCAGCTGGAAGAATCCGAGGAGGC
CATCATGAAGTTGTTTGAAGTGAATCGCAAACTAATGAAGACTGTTGAAGATGAGCCACTGTACTTTGACGAGAAACCTGAATTAGCACCAGATGAGAGT
GGCAGTGTCAGGAGAAGAAAGATTACAGAGCAGGCAAGAAGAGTGTCAGAGAAAATCGGGCGTTTGCAGTTGGAGGTGCAGAAATTGCAGTTCGTTTTGC
TGAAACTTGATGATGAAAACAGAAGTAGAGGGAAAACAAAAATCACAGAGCAAAAAACAAAAGTTCTACTGCAGGACTATCTGTATGGCAGCACAAGAAC
TCGCCAGAAGAGGAAGAAAGGGCACTTCTGTTCATGCGTGCAACCTCCAACTAAAGGAGATTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G049600.1 pacid=42779933 polypeptide=Potri.002G049600.1.p locus=Potri.002G049600 ID=Potri.002G049600.1.v4.1 annot-version=v4.1
MAARSQADSKRKYSWWWDSHISPKNSKWLQENLTDMDSKVKQMIKLIEEDADSFARRAEMYYKKRPELMKLVEEFYRAYRALAERYDHATGALHQAQRTM
AEAFPNQAPFILGDDSPAGSATDCDPRTPDMPPIRAPFDPDELQKDALGVSPSHAINRNGAFTEKSDPGRKGLKQFNDLFGLGDGMDNAKFAEGRVRKGL
NFHDPEEKGRGVQNNGIHDLKARAPSESEQVSKAELEILNLKNALAKLEAEKEAGLLQYEQSLERLSKLESEVSRATEDSRGLNERASKAEAEVQTLKEV
LAQLEAEKESSFLQYQGCLEKISNLENNLSLVQKDAGELNERASKAETEARSLKQDLSRLEAEKIDAQVQYSQCLEKISDLEDKLHNAQEDAKRFSERAD
DAEREIEALKHALTRLTEEKEAAVTQYQQCLATIVSLEHKIACFEEEARRLNLVIDNGTVKLKSSEERCLLLEKSNQTIHSELESVMQKVAAQSNELTEK
QKELGRLWACVQEEHLRFMEAETAFQTLQHLHSQSQEELRSVVAQLQNRAQILEDLEARNQSLKDEMEHVNVENKSLSDVNLSSALTIQNLQDEISSLRE
TIKKLEAEVELRVDQRNALQQEIYCLKEELNELNQKHQAIMRQVESVGFSPESFGSSVKDLKDVNIKLKEVCERDRTEKVALLEKLENMEKLIDKNALLE
NSLSDLNVELEGVGEKLKALEESCQYLVEEKSVLVSEKDLMASELQFATDDLEKLTEKNHILENFLLDANAELEGLREKSKSLEDFCLLLVNEKSELASM
KGSLSSQLDISEKSLQDLEKNYTELEEKYSHLEKERQSSLHEVQELQVRLDAEKQEHANLAQLSESQLAGMASQICLLQEESLCRKKEYEKELDKAVNAE
IEIFILQKCAQELEEKNSSLLLDHQKLVEASKLSEKLISDMRHENCEQQEEVKCLSDKIKTLRMGLYQVLMTLELDANQCENKPKQDQKLLNHVLNRLQE
SQEFLFKTQDENQRLFTENSVLVTLLRQLQLEVENLVKTKNILDQELTTRSEQFLVLQNESQELSGINEEMKLKLIEGDRKEEALKVELNNLHVQLSDLQ
GAFQNLQEENCKVLDDQRSLMKSFSDLQMEKCELEEENFCILVETVSQSTLSLIFRDIICEKSVEIKSLGVSLDKQCHDNNGLNEKVKTLEKELDNFSGL
EDDKRELHKMVEDLKCKYDEVEVIRSDQEMQIIKLLGDYDQKIKEAENIREVNQKLESEIRKLHEEFQEVKDRKENLSHELVKERNEVELQESQAVALFG
ELQISAVREALFEGKLRELLKICESLEDGNCSKDMEIDQLKERVSTLEGGNAELKALVAAYLPAFMSLRDCVTSLEKHTLPDATLHEGDSKESKDAALVV
HAKGFHQMSEGQSGMVPGGTLDFQDLQMRIRAIEKEIIEKERLVMLENLSYHSKLDAAIRQIEDLKSGSSARQKGVETRRYVKPKPEDGELGATPSDDLR
RQKRTHEISEDGNEVMTKDIILDQISECSSHGISRRETMQADEQMLEIWETADRDDSIDLTVGKTQKVTASQKKKKHIRQHPSAESMVEKEVGVDKLEIS
KRLSGSRQEGNERKILERLDSDAQKLTNLQITVQDLMSKVEITEKSEKGKGIEYDNVKEQLEESEEAIMKLFEVNRKLMKTVEDEPLYFDEKPELAPDES
GSVRRRKITEQARRVSEKIGRLQLEVQKLQFVLLKLDDENRSRGKTKITEQKTKVLLQDYLYGSTRTRQKRKKGHFCSCVQPPTKGD
|
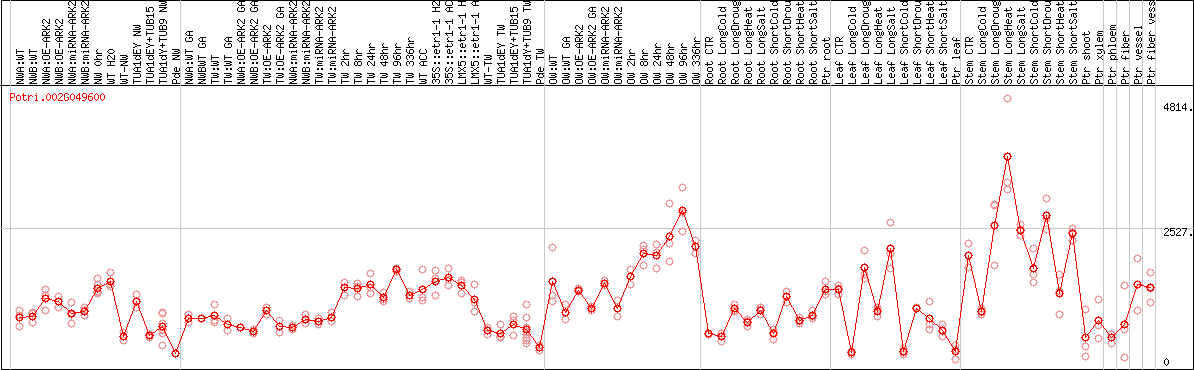
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G049600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
