External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G43000 281 / 6e-95
NAC
ANAC042, JUB1, JUNGBRUNNEN1
NAC domain containing protein 42 (.1)
AT3G12910 231 / 1e-74
NAC
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT5G39820 220 / 6e-70
NAC
ANAC094
NAC domain containing protein 94 (.1)
AT1G26870 216 / 2e-67
NAC
FEZ, ANAC009
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT2G02450 176 / 2e-52
NAC
LOV1, ANAC034, ANAC035
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
AT1G69490 170 / 3e-51
NAC
NAP, ANAC029, ATNAP
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
AT3G04070 170 / 2e-50
NAC
ANAC047
NAC domain containing protein 47 (.1.2)
AT1G61110 166 / 2e-49
NAC
ANAC025
NAC domain containing protein 25 (.1)
AT2G17040 165 / 3e-49
NAC
ANAC036
NAC domain containing protein 36 (.1)
AT1G01720 163 / 2e-48
NAC
ATAF1, ANAC002
Arabidopsis NAC domain containing protein 2, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G205400
470 / 8e-169
AT2G43000 269 / 6e-90
NAC domain containing protein 42 (.1)
Potri.001G080900
301 / 2e-102
AT2G43000 257 / 3e-85
NAC domain containing protein 42 (.1)
Potri.007G066300
291 / 3e-98
AT3G12910 269 / 2e-89
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.005G098000
291 / 4e-98
AT3G12910 271 / 2e-90
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.003G149700
281 / 2e-94
AT2G43000 245 / 2e-80
NAC domain containing protein 42 (.1)
Potri.017G082000
231 / 1e-73
AT1G26870 299 / 3e-98
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.004G126901
231 / 2e-73
AT5G39820 305 / 1e-101
NAC domain containing protein 94 (.1)
Potri.003G046700
173 / 4e-51
AT2G02450 318 / 2e-106
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Potri.011G046700
172 / 4e-51
AT1G61110 331 / 8e-113
NAC domain containing protein 25 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031189
249 / 4e-81
AT3G12910 264 / 9e-87
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10038332
233 / 1e-75
AT2G43000 231 / 1e-75
NAC domain containing protein 42 (.1)
Lus10031767
230 / 2e-75
AT2G43000 236 / 3e-78
NAC domain containing protein 42 (.1)
Lus10030478
236 / 3e-75
AT5G39820 297 / 4e-98
NAC domain containing protein 94 (.1)
Lus10036194
227 / 1e-73
AT3G12910 224 / 1e-72
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10037178
231 / 2e-72
AT5G39820 306 / 3e-101
NAC domain containing protein 94 (.1)
Lus10036749
230 / 2e-72
AT1G26870 306 / 4e-100
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10015367
222 / 9e-71
AT1G26870 298 / 8e-99
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10007263
219 / 2e-69
AT5G39820 300 / 3e-100
NAC domain containing protein 94 (.1)
Lus10037156
177 / 1e-53
AT1G69490 318 / 2e-109
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Potri.002G057200.3 pacid=42777055 polypeptide=Potri.002G057200.3.p locus=Potri.002G057200 ID=Potri.002G057200.3.v4.1 annot-version=v4.1
ATGGAGGCCTTGTCAAAGATGAGTAGTTCTGATATTGATCATAACGAGGATGAGGAAGTGATGCTTCCAGGTTTCCGATTTCACCCCACTGATGAAGAGC
TTGTTGGGTTCTATCTGCGGCGGAAGGTGGAAAAGAAACCACTTAGGTTTGAACTCATCAAACAGATCGATATCTACAAGTACGATCCATGGGATCTTCC
AATTAGCAATGCGGGGGACAGGGAGTGGTACTTCTTCTGCATACGAGGGAGAAAGTATAGGAACAGCATAAGGCCTAACAGAGTCACAGGATCTGGATTT
TGGAAGGCCACCGGCATCGACAAGCCCATATATTCAGTCAAAGAACCACATGAGTGCATTGGCCTCAAGAAATCCTTGGTCTACTACCGGGGAAGTGCTG
GAAAAGGCACTAAAACTGATTGGATGATGCACGAGTTTCGCCTTCCACCAAACGGGAAGACCACAAACCCTAAGAACATTGCACAAGAAGCTGAGGTTTG
GACGCTCTGTCGAATTCTCAAACGAGTCCCGTCCCACAAAAAGTACGCACAGGCAGCAGCAGCAAAGCAGCACCCTATTATTGACTCGAATTCAAAAACA
TGCAGCTTTGAATCTGAAATTAGTGCCGAGCAGAATGTGAGTTTTGGAGACTCGTTCGGTCAACGAAATGAAAGGAAGCCTGTCATCATAGATAATCATG
TTAACCATGAAAGGAATCACTTTTTTACGGGTGGTCAGTATGATTCGATAACTGAAGCTCCATTTACAGCAGCTTTCCAGAACTTCTGGAATCCAGATGT
TGGAGATGATTTCTTTGCTAATGGAAACTGGGATGAACTAAGAACAGTGGTCGAGCTAGGCGCCATTGATCCAGCTCAAGCTTACGACTATTGTAGATAA
AA sequence
>Potri.002G057200.3 pacid=42777055 polypeptide=Potri.002G057200.3.p locus=Potri.002G057200 ID=Potri.002G057200.3.v4.1 annot-version=v4.1
MEALSKMSSSDIDHNEDEEVMLPGFRFHPTDEELVGFYLRRKVEKKPLRFELIKQIDIYKYDPWDLPISNAGDREWYFFCIRGRKYRNSIRPNRVTGSGF
WKATGIDKPIYSVKEPHECIGLKKSLVYYRGSAGKGTKTDWMMHEFRLPPNGKTTNPKNIAQEAEVWTLCRILKRVPSHKKYAQAAAAKQHPIIDSNSKT
CSFESEISAEQNVSFGDSFGQRNERKPVIIDNHVNHERNHFFTGGQYDSITEAPFTAAFQNFWNPDVGDDFFANGNWDELRTVVELGAIDPAQAYDYCR
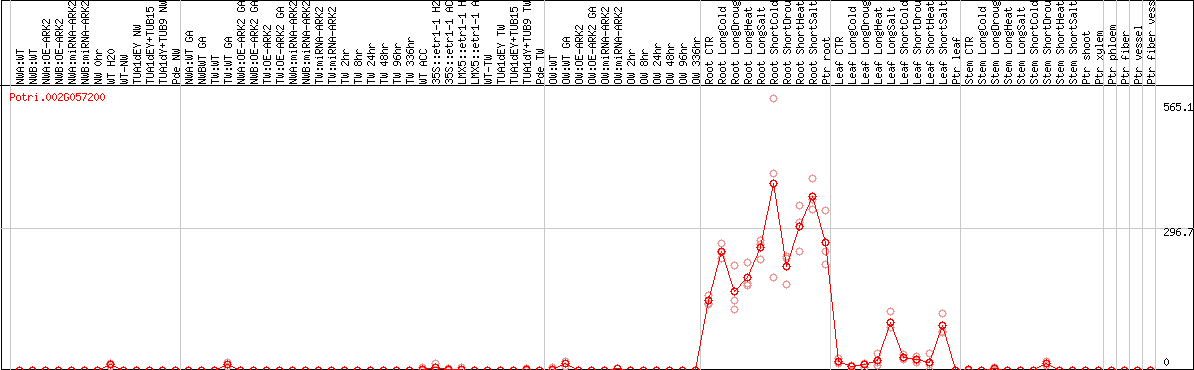
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G057200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.