External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G76760 193 / 1e-63
ATY1, TRX-Y1
thioredoxin Y1 (.1)
AT1G43560 182 / 3e-59
ATY2
thioredoxin Y2 (.1)
AT1G50320 89 / 1e-22
ATHX, ATX
thioredoxin X (.1)
AT1G03680 87 / 1e-21
ATHM1, ATM1, TRX-M1
ARABIDOPSIS THIOREDOXIN M-TYPE 1, thioredoxin M-type 1 (.1)
AT3G15360 81 / 1e-19
ATHM4, ATM4, TRX-M4
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
AT3G51030 78 / 4e-19
ATTRX1, ATTRXH1
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
AT4G03520 78 / 2e-18
ATHM2
Thioredoxin superfamily protein (.1.2)
AT5G39950 73 / 7e-17
ATTRXH2, ATTRX2, ATH2
Arabidopsis thioredoxin h2, thioredoxin 2 (.1)
AT2G15570 74 / 9e-17
TRX-M3, GAT1, ATHM3, ATM3
THIOREDOXIN-M3, GFP ARRESTED TRAFFICKING 1, Arabidopsis thioredoxin M-type 3, Thioredoxin superfamily protein (.1.2)
AT1G45145 71 / 3e-16
LIV1, ATTRX5, ATH5
LOCUS OF INSENSITIVITY TO VICTORIN 1, thioredoxin H-type 5 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018875
190 / 2e-62
AT1G76760 176 / 6e-57
thioredoxin Y1 (.1)
Lus10028569
188 / 2e-61
AT1G76760 174 / 3e-56
thioredoxin Y1 (.1)
Lus10014798
88 / 4e-22
AT3G15360 181 / 2e-58
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10040887
87 / 7e-22
AT3G15360 182 / 4e-59
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10024293
84 / 2e-21
AT3G51030 162 / 4e-53
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
Lus10029755
82 / 4e-20
AT3G15360 162 / 3e-51
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10041799
81 / 4e-20
AT3G51030 182 / 8e-61
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
Lus10042784
82 / 5e-20
AT3G15360 165 / 4e-52
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10029752
82 / 6e-20
AT3G15360 165 / 3e-52
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10000802
81 / 8e-20
AT3G51030 165 / 5e-54
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0172
Thioredoxin
PF13098
Thioredoxin_2
Thioredoxin-like domain
Representative CDS sequence
>Potri.002G066800.1 pacid=42779745 polypeptide=Potri.002G066800.1.p locus=Potri.002G066800 ID=Potri.002G066800.1.v4.1 annot-version=v4.1
ATGGCGATTTCTTCTCTCTCGGCTTCAACGATTCCTTCATTGAATACTCCCAACTCCACTTCTAATTACTCCTCCAAAATATCTTCATTTTCTTCTCTCC
AGTTTCCAGTACAGCCTCATCGACTCCAGTTTCGAAAGAGATGGATTACCTCTCCTTCCCGACCCCCAATTTTGCCACTGGTTGCAGCAAAGAAGCAAAC
TTTTTCCACCTTAGATGAGTTGCTAGAAAAGTCTGACAAACCTGTTTTGGTTGACTTTTATGCAACCTGGTGTGGTCCCTGTCAATTTATGGCTCCAATT
CTAAATGAAGTCAGTGCTGTCCTGGAAGACACGATCCAGGTGGTGAAAATCGACACCGAGAAATATCCTAGCATTGCTGATAAATACAGAATAGAGGCCT
TGCCTACTTTTATCATATTCAAGGATGGAAAGCCTTATGATCGTTTTGAGGGTGCTTTGACTAAAGATCAGCTCATCCAACGCATTGAAAATTCACTGAA
TGTCGAGCAATAG
AA sequence
>Potri.002G066800.1 pacid=42779745 polypeptide=Potri.002G066800.1.p locus=Potri.002G066800 ID=Potri.002G066800.1.v4.1 annot-version=v4.1
MAISSLSASTIPSLNTPNSTSNYSSKISSFSSLQFPVQPHRLQFRKRWITSPSRPPILPLVAAKKQTFSTLDELLEKSDKPVLVDFYATWCGPCQFMAPI
LNEVSAVLEDTIQVVKIDTEKYPSIADKYRIEALPTFIIFKDGKPYDRFEGALTKDQLIQRIENSLNVEQ
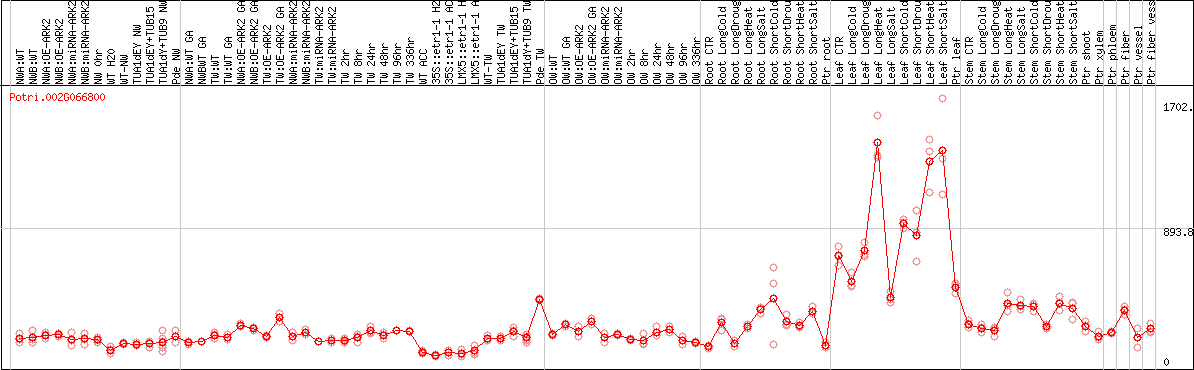
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G066800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.