External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G21360 253 / 9e-86
GLTP2
glycolipid transfer protein 2 (.1)
AT3G21260 251 / 8e-85
GLTP3
GLYCOLIPID TRANSFER PROTEIN 3, Glycolipid transfer protein (GLTP) family protein (.1), Glycolipid transfer protein (GLTP) family protein (.2), Glycolipid transfer protein (GLTP) family protein (.3)
AT2G33470 130 / 1e-37
ATGLTP1, GLTP1
ARABIDOPSIS GLYCOLIPID TRANSFER PROTEIN 1, glycolipid transfer protein 1 (.1.2)
AT2G34690 49 / 7e-07
ACD11
ACCELERATED CELL DEATH 11, Glycolipid transfer protein (GLTP) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G188600
400 / 6e-144
AT1G21360 259 / 3e-88
glycolipid transfer protein 2 (.1)
Potri.010G069600
140 / 2e-41
AT2G33470 332 / 3e-117
ARABIDOPSIS GLYCOLIPID TRANSFER PROTEIN 1, glycolipid transfer protein 1 (.1.2)
Potri.008G168800
129 / 3e-37
AT2G33470 315 / 1e-110
ARABIDOPSIS GLYCOLIPID TRANSFER PROTEIN 1, glycolipid transfer protein 1 (.1.2)
Potri.002G035600
122 / 8e-35
AT2G33470 273 / 2e-94
ARABIDOPSIS GLYCOLIPID TRANSFER PROTEIN 1, glycolipid transfer protein 1 (.1.2)
Potri.006G051800
54 / 8e-09
AT2G34690 305 / 9e-107
ACCELERATED CELL DEATH 11, Glycolipid transfer protein (GLTP) family protein (.1)
Potri.016G055300
46 / 6e-06
AT2G34690 295 / 1e-102
ACCELERATED CELL DEATH 11, Glycolipid transfer protein (GLTP) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018917
290 / 3e-100
AT3G21260 229 / 3e-76
GLYCOLIPID TRANSFER PROTEIN 3, Glycolipid transfer protein (GLTP) family protein (.1), Glycolipid transfer protein (GLTP) family protein (.2), Glycolipid transfer protein (GLTP) family protein (.3)
Lus10028618
268 / 3e-91
AT1G21360 224 / 6e-74
glycolipid transfer protein 2 (.1)
Lus10042793
273 / 5e-89
AT1G21370 486 / 2e-170
unknown protein
Lus10029765
251 / 2e-85
AT3G21260 212 / 9e-70
GLYCOLIPID TRANSFER PROTEIN 3, Glycolipid transfer protein (GLTP) family protein (.1), Glycolipid transfer protein (GLTP) family protein (.2), Glycolipid transfer protein (GLTP) family protein (.3)
Lus10003626
130 / 3e-37
AT2G33470 242 / 3e-81
ARABIDOPSIS GLYCOLIPID TRANSFER PROTEIN 1, glycolipid transfer protein 1 (.1.2)
Lus10021169
128 / 1e-36
AT2G33470 334 / 3e-118
ARABIDOPSIS GLYCOLIPID TRANSFER PROTEIN 1, glycolipid transfer protein 1 (.1.2)
Lus10011806
124 / 3e-35
AT2G33470 324 / 4e-114
ARABIDOPSIS GLYCOLIPID TRANSFER PROTEIN 1, glycolipid transfer protein 1 (.1.2)
Lus10008244
98 / 4e-25
AT2G33470 216 / 1e-71
ARABIDOPSIS GLYCOLIPID TRANSFER PROTEIN 1, glycolipid transfer protein 1 (.1.2)
Lus10023267
42 / 0.0002
AT2G34690 293 / 5e-102
ACCELERATED CELL DEATH 11, Glycolipid transfer protein (GLTP) family protein (.1)
Lus10038537
42 / 0.0002
AT2G34690 279 / 5e-94
ACCELERATED CELL DEATH 11, Glycolipid transfer protein (GLTP) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF08718
GLTP
Glycolipid transfer protein (GLTP)
Representative CDS sequence
>Potri.002G071100.1 pacid=42778017 polypeptide=Potri.002G071100.1.p locus=Potri.002G071100 ID=Potri.002G071100.1.v4.1 annot-version=v4.1
ATGAAAAGGACCAGAGAAATAGAGAAGGGATCAGAGATCAAGTCTGCAATTGAAGAACTTTCGATGCTGATCAAACTCAAGCCTACAGGTGACAATCACG
ATAGGACTACTGTTCATATACCCACAAGGCCTTTTATGTATGTCTGCAACCTTGTTATTCAGGTTCTTGATAAGATAGGCCCAACAATGACAGTTTTGAG
ACAAGATATTGATCAGAATATTCAGAGATTGAAGATGCTATGTGATTCTGATCCTTCAATGTACTCCAATTTGGTTGAGATTTTAAAGAAAGAGGCCGAT
GAAGGCGGTGCAAGAAAGGGTGCCAGCTGCAGTAAAGCCTCTGTTTGGCTTGCCAGATCCTTGGATTTTACGGTTGCTTTGTTAGAAAGGTTAGTGGCGG
ATCCTGGTCAGGAAATGGAGAAACTCGTGGAAGAATCTTACAACATCACTTTAAAGCCATGGCATGGTTGGATTTCATCAGCCGCTTATAAAGTAGCTCT
TAAATTAGTACCTGATAATAAAACCTTGATTGATCTGCTCATGCCAAAAGACGAAACCTACGACACCCTTAAAGAGGATGTGCAAACATTGATCTCACTA
CTTGTGCCTTTTTTAGAAGAGATCCACTCTGTTCTGATATTGTACGGGTTGGATAGGTTGAAGTCCACTTAA
AA sequence
>Potri.002G071100.1 pacid=42778017 polypeptide=Potri.002G071100.1.p locus=Potri.002G071100 ID=Potri.002G071100.1.v4.1 annot-version=v4.1
MKRTREIEKGSEIKSAIEELSMLIKLKPTGDNHDRTTVHIPTRPFMYVCNLVIQVLDKIGPTMTVLRQDIDQNIQRLKMLCDSDPSMYSNLVEILKKEAD
EGGARKGASCSKASVWLARSLDFTVALLERLVADPGQEMEKLVEESYNITLKPWHGWISSAAYKVALKLVPDNKTLIDLLMPKDETYDTLKEDVQTLISL
LVPFLEEIHSVLILYGLDRLKST
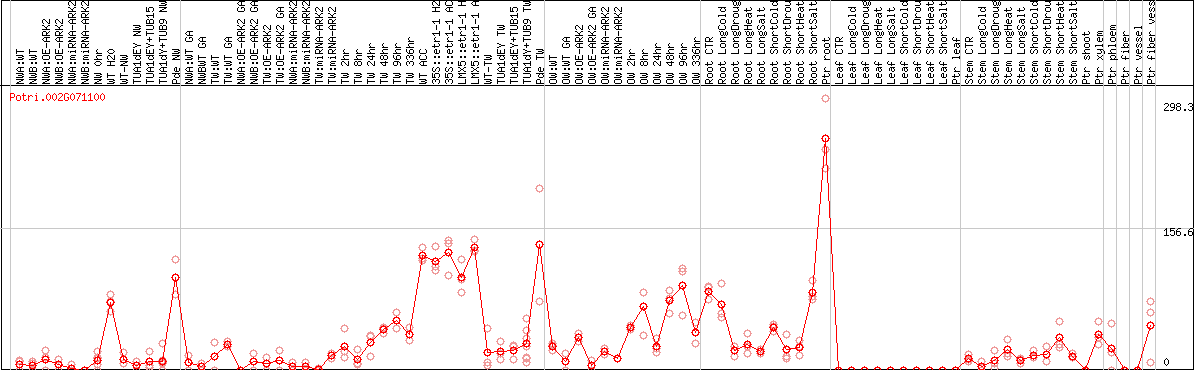
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G071100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.