Potri.002G079100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G079100.4 pacid=42777238 polypeptide=Potri.002G079100.4.p locus=Potri.002G079100 ID=Potri.002G079100.4.v4.1 annot-version=v4.1
ATGGGCGCGTGTGAGCCTCACGAGCCCTTGAATCTTGCTGCGACTGAGCAGCATTCATGCCTGGAGTTTATCAAGAATTCTGAGCAGCTGCCAGCCTTGG
AGACTGGTCGGAGTGCTATTGAATTTTACGCGGACCCCTCTAGGGCAACAAATGGGTTTGCAGAATTGTCTCAGAAAGATAATGGTGTTTGTGCTAGCTA
CAGCGATGTTATGGAGGCTGTGTTGGATGATAGGGGAATTGGGTTGGCGGGTGAGGGTGAGAACGTGGTTGGTTCGATAGCAGGAAGATTGTTGGAGGAT
GAATCTGGGGTTGGTGAGGAATGTTTGGATGAGGGTCAAAGTGGAAGGGATGATTGTATCAGGGAGACAGATAGATTCTGGGAGGAAAAAGTAGGGCTTG
GAGGTGAGAATGGTGGAGCTGTAGATTGTGAAGGGTCTTTGGAACTGCTGGTTGTTCCTGATTCACTGAAAAATTGCAATCAACAGGATGACCAAATGGA
TGATAAAAATGCTGGTGTTCAAGGCACTATGGAAGAGGGCAGTGATGGTCTAGCTACAGCAGAGACTGACGCTTCTGATGAAACAGTGCCACCAACCGCT
TGTGGAACATCTGTAGAATTAAACCCTGTGAATGATATGCCTAGAAATTGTGATCAACAAGATGGCCAGAATGAAGATGAGAGTGGAAATGTTCAAGGGG
TTATTGAAGATGACGGTTTAGATGCAATAGAGACTGTTAAACCCAACAAAATAGTTCTTTCACTGGGCTGTAAGATGCCAGCTGAATTAGTGCAAGCAAA
TGATTGGTGCAGAAATGGCGGTCAACAAGATGACCAGAGGGATGATAAGAACGTTAGTGTTCAAGGGGTTATGGAAGAAAACAGTGGTGACTTGGCTCCA
ATAGAGGTTGATACACATGATGAAATAATGCTTTTATCGGGTAGAGAAATGCCTGCAGAACTAATACCAGTAAAAAGTTTGCCTGGTGATGTCAGTGAAC
AGTACAACCGTGATTGTGGTGCCTCTCAAGAAGTGATTGTGGAAGAGAAAATTGGCAATTTTACTGCCCTAGGGAAGAGCAACGTTCAAGGGGTGATGGA
ACAGAAAAGTAATGGTTTAGTGGCGACGGAGACTGTTGATACTTGTGAAAACATATTGCCTTCACTGGGCTATGAAATGCTTGCAAAATGTTTGCCTAGA
AATGGTGTTGAACAGGACATGCAAGACAGTGGTACCTCTATGGTGGTGACCATGGAAGAGAAAAACAATGATTTAGCTGGGATAGAGAGTATTAGTATTC
AAGGGGTCATGGAAGACAAAACTGATGGTTTAGCTGCAACAGAGACTGCTACCTGTAATGAAATAGGTCCTTCACCAAGCTGTGAAATGCCTGTTGGATT
GATTTCAGTGAATGTTTCACCTAGAAACGGTGTTGAACAGGACAAGCAAGATGATGGGACATCACGTACTATGGTTTCAGAAGAGAAAAGTGTTATCTTA
ACTAGGTTAGAGACTGATAATCAAGACCAAAAGTTGCTTCCACTGGACCATGAAATACTCTTGGAATTGACACCTGTGACATGTCCACCAAGCAAGTGTC
TTCAGCAGGATGACCAGAAGGGTGATCAGATCATTAGCCGTCCCTTTGCAGGAGGGGTTATGGAGGAGCCAACTTTTGTTTTAGATGCGGCTGAGACTAC
TACTTCCAACTTGTCGTTGCCATCACAGGAGAACTTGAAATTAATGCCCACGACTGGTTTGCCAGAAGAAAATGTTCATCATGATGAGCAGAAGTTGATA
CCATGTAAACTGGATTCCAAGGCTGTCAATGGTCTAGCTATAGAGTGGGTTCCAGAACAGGAAAGCAATGCCTTGGCTAGGACAGAAGCTGGCATTTGTT
CCCAGGCATCAGCACATGGCACTATTGATTCTAGCAGTGCAGTTGACTGTTCTGGAGAGACAGACTATGAAGCAAAGAATAATGTCAGCATTGATAGTGT
TTCTGAAACCAAGTGTCATGTCATTGTATCACCATCTTCACGAAGGAGCAATGGAACACGGAAATCAAGCCAGAAGACCCAGACAAAAAGGGGTGCAAGG
AAATGTAGGAACACAACCAAAGTCCCAAACCTGCATCGAGGTATTGAGATCGTCTTCAAGAGTGTGACAAGGAGGCGGAGCTGCTTCTCCAAACCAGCTC
GCTCTTCTGCTTGGGGATTATTGGGGAATATTACACAAACTTTTATGCTGATTAATGGTCTCAGACCTGATGAAATTGAGAATCTTGGATCACAGAAGGC
AAGGGGTGACCAAGGAAGTGGAAAGCGGAATAAACTTGCTGGTGGAACCTCACGAAGGTCCAGCAAGAAAGGTCATGCTTCAGCTCATTGCATTCGTTTG
AAAGTTAAAGTGGGAAAAGATGCTTGTCAAACTGAAAGTAATCCAAAGATGATAATCCCCGAGGTAATTAATACTAAGGCATCTGGTGATCTTGTTAGTG
ATTATGGGGCTGAGTCATGCCAGGAAACCAGTTTTGAAATTTCAAAGTTAGCCTATTGTGTGGGAGATAACATGGTTGAAGAGGGAACTCAAAAACAGCT
TCAGTCTTTTTACATTAAGTTGGGGAAGGCAAAAGCACACTGTGATGCTTCTGCTATGGATGTAAAACTTGCAAACAAGGACATGGAGGGCACTGTAATT
TCTGAGAAGTCATCTAGAGATATTATGGAAGATTATCTTGGGGTTCCTTCCCATACAGAGGTTGAAGCACTGGGAGTTGCAACTGAGAAGAGGTATACAG
ATGCTGGAACTTCACCAGATTCGGAAGTTATAAATTCAGTACCCGAGGTTCAAGTCAATGCAAGATGTCAAGAAGATTATCCTGATGCTGTTCTGTCTCC
TTCTAAGGCTTTTGCTGCAGATGAAGAGGGTACTGGTGGAAAGAGGGGAAAAAAGAAAGAAAGCCTCCCTCAAGCAGGAAACTGTTCACCTGCTGTGGCA
AGCTTAAAGAAAGTTAAATTGGCAAAGAAACGTGGGGGCAGGCAGAGAAAGGGTGATAGTCTTTCCTCTAGTGAGATTCTTACATCATGCACCAGTGCTA
ATGGTTCTGTCAACACAACAAGCACTAAAGAATATTCAGCAGAACTGGTACTTTCATCAGGAAAGACGGAACTTGGAGACCCTGAAGGGGCTTTGAGAGG
AGAAATCATTATGGAAACTAAAATATGTGGTGAACTGGATGCTGACGTTAGGTCGTCTGAATCACAGATTTCAAAGAATCCTCTTCCTTCAACTAAATCC
AGGGGACGCCGACTTCCAAGGAAATCTGATGGAGTAAACAAGAGAAGGTCCAAAGTCTCTGACTCTGCAAAAAGCAGGAGGGCAAATGGTTGCAAGGAGA
GAGGAAATGATAGGAAGTCAGTTAAAAAGAACAAAGCTGAGGAGAAGAGTGTTTGTGATCATGTTGTTTACAAAGATGACAATGGAAAAACTGATGCTGG
CAATGATACTACTGCAGAAGAGGTAACTAATCTGGACATGCCGTCAAGTGGGGTAATGGAACAAAATTTATTCCCAGATAATGCTTGGGTTCGCTGTGAT
GATTGTCTTAAATGGCGGCGTATTCCAGTTAGACTTGTAGAATCTATCAGTCAAACACATCGCCAATGGATCTGCGAGGACAACATGGATAAAGCCTTTG
CTGATTGCTCCTTCCCCCAAGAGAAGTCAGATGCAGAAATTAATGCGGAGTTGGGCATATCAGATGCTGATGAGGATGTTTGCGATGCTCCTTCAAATTA
TATGGAATTAGAATGTGGACCGACATCAGTTTCCAAGGAGTATGAATTTACGCGCATTACTACCAATCAGTTTCTGCACCGCACCCGTAAAACTCAAACT
ATTGATGAGATAATGGTTTGCTACTGCAAAGCACCTGTGGGCGGCAGGTTAGGTGGTTGTGGAGATGAATGCTTGAATCGAATGCTCAATATTGAGTGTG
TTCAAGGCACCTGTCCATGTGGGGACCTTTGTTCAAATCAACAGTTCCAAAAGCACAATTATGCCAAAATGACATGGGACCGCTGTGGGAAGAAAGGTTT
TGGGCTGCGGTTGGAAGAGGATATAACCAGAGGGCAATTCCTTATTGAATATGTTGGAGAGGTGCTTGATGTGCATGCTTATGAGGCAAGGCAAAAAGAG
TATGCTTCCAAGGGTCACAAACATTTCTACTTCATGACATTGGATGGCAGTGAGGTAATTGATGCATGTGTGAAGGGAAATTTGGGGCGTTTCATTAATC
ATAGCTGTGATCCTAATTGTCGTACCGAAAAGTGGGTGGTCAATGGAGAAATCTGTATTGGATTGTTTGCATTAAGGGATATCAAGAAGGGTGAAGAGGT
GACATTTGACTACAACTATGTAAGAGTGGTTGGGGCTGCTGCTAAAAGATGCTATTGTGGTTCACCTCAATGCCAAGGCTATATTGGTGGTGATCCAACT
AGCTCTGAAGTAACTGATCAAGTTGATTCGGATGAAGAATTTCCTGAACCTGTAATGCTTGAAGATGGGGAAGTTGGAGATGGCTTAAAAAATAAAATAT
CCAAAACCAGTTTCTTCGGTTTATCAAAAGGTAGGGAAATGGAATCCAAAACAGCAGTTGGGAACTTGGAGGTTGCCACTGAAATTAAGGATTCAATGAA
CCAATCAACTCCTGCCATTTCTCAATCACCCAGTGAATCAGAAATGAATGGCTTACCTGGAGATTTTTCTTCCTCAAGTAAACGAGTAGAAATCTCCCCA
CAGACTGAAGACATGACAACCCAACCCACACCTGCTGTTCAGCAGGAGATTTCCATGGAAGAGATGATGGATAAATCCTTGTATTCCAGTCAAAAGTTGA
AGACTTCTCTGACCTCAGTGCTCATCAAACCATTGCCTGATGACATTATGATTAACAGGAAGTCTAAATCTGCCACAGCTGAAAACAAGCGGGTTTTTGT
TAAATCTCGTTTTATTATTAAAACTCCACCCCAATCTGGTTTGATTAAGAAAGGAAAGTCCGCCAGCAATTTTATAAACATAAATAAGGTTCAGACAATA
ACCAACAAACCTCATATGCCGCCTATCAAACCCAAAAAATTATCGGAAAGTACCTCAGATGGCCATTTTGAAGCAGTTCAGGAGAAACTTAACGAGTTGT
TGGATTCTGAGGGTGGCATAAGCAAGCGGAAAGATGCCCCCAAAGGCTACCTGAAACTTCTCCTCCTCACTGCAGCTTCTGGTGCTATTAGAAATGGTGA
AGCAATTCAGAGTAATCGAGAGCTTTCAATGATACTTGATGCACTCTTGAAAACTAGATCCCGAATGGTATTGATGGATATAATTGAAAAAAATGGTTTA
CGGATGCTTCACAACATAATGAAGCAGTACAGAAGGGATTTTAAAAAGATTCCGATTCTTCGAAAGCTTCTAAAGGTTTTAGAGTATTTGGCCGTTAGGG
AGATACTTACATTGGAACATATAAACGGAGGTCCTCCTTGTCCTGGAATGGAAAGCTTTAGAGAGTCCATGCTGTCCCTGACAGAGCACAATGACAAACA
GGTTCATCAAATTGCACGAAGTTTTCGAGATAGATGGATCCCTAGACAAGTCAGAAAACTTGGCTATATGGATAGGGATGGTGGTAGGATGGAGATTCAG
AGAGGTTCAAACTGCAACAAGGTTTTAGCTTCTCATAGTCAATGGCATGATCAGGGTGTAAGACACTTGGAAGCACTGAATGGTACGGTGGAATCAAACC
TAGCAACAACCTCAGTGGGCACTGCTGTCCACGAGGACAGCTCTGCGAACCGTGTAGGTAGTGGTACAAGAACTCGGAAGCGTAAAAGCCGTTGGGATCA
ACCAGCTGAGGAAAACATAGCTTCCAGATCTCTTCAGCATGTTGAACAGAACGAGTCTGGGTTATTACAACAATCTGAATCCAACTCATTACCTGAGTTA
AGTAAAGAAGTGCCAGATCATGTAGACAAGGCAGGTGGAGAGTACAGTTATTGCCCTCATTGTGTTCACAGTTATTGTTGGCAAGATGAAGCTAGTGGTG
CCGATAATGGAAGGCAGAATATTCATGAGGATGTTCCACCAGGATTCTCATCTCCTATTGACCCCGCTCTTGTGTCCAATGCTTCTTCTACTGTTGATGA
TCTCCCACACCAAAATGTTTTCCATTTGAAGTTTCCTGTTGGTGTGGTTGTTGGCCTTCCACAGAGGAAATTCAATTCCCGCTTTCCTGTCTCATATGGA
ATTCCGTTGCCTGTTGTGCAGCAATTAGGGTCACCTCTGGCTGAAACTGTAGAGGGTTGGATTGTTGCTCCTGGCATGCCTTTCCATCCTTTTCCGCCAT
TACCACCACTTCCCTCCTGTAAGAAAGGGACTCTGCCATCTGCTATGAACTCTATGGAGATTGATGACACTGCAGATCGAGGGAAACAGGACTGCTATGA
TCGCACCACTTGTCTAGATGAAAATAGTCCAAGCACGACTGGTGCTAACCAACCAGACCTGAACAGTCCAGGTCCGAAAGACCATCAAACATTTAAACGT
GCAAGGGGGTCTTATGATTTGGGTAGGAGGTACTTTAGGCAACAGAAGTGGACTAAAATGCTTCCTCCATGGGTTCGGAGTAGGAATGGGTGGGGATGCA
TTGGAGGCAACTCGAGAGGTGGAATGTGCAGCACTGACTTGGGAAGTCTAACAAATGAACAGAGAAACTCGTATTATTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G079100.4 pacid=42777238 polypeptide=Potri.002G079100.4.p locus=Potri.002G079100 ID=Potri.002G079100.4.v4.1 annot-version=v4.1
MGACEPHEPLNLAATEQHSCLEFIKNSEQLPALETGRSAIEFYADPSRATNGFAELSQKDNGVCASYSDVMEAVLDDRGIGLAGEGENVVGSIAGRLLED
ESGVGEECLDEGQSGRDDCIRETDRFWEEKVGLGGENGGAVDCEGSLELLVVPDSLKNCNQQDDQMDDKNAGVQGTMEEGSDGLATAETDASDETVPPTA
CGTSVELNPVNDMPRNCDQQDGQNEDESGNVQGVIEDDGLDAIETVKPNKIVLSLGCKMPAELVQANDWCRNGGQQDDQRDDKNVSVQGVMEENSGDLAP
IEVDTHDEIMLLSGREMPAELIPVKSLPGDVSEQYNRDCGASQEVIVEEKIGNFTALGKSNVQGVMEQKSNGLVATETVDTCENILPSLGYEMLAKCLPR
NGVEQDMQDSGTSMVVTMEEKNNDLAGIESISIQGVMEDKTDGLAATETATCNEIGPSPSCEMPVGLISVNVSPRNGVEQDKQDDGTSRTMVSEEKSVIL
TRLETDNQDQKLLPLDHEILLELTPVTCPPSKCLQQDDQKGDQIISRPFAGGVMEEPTFVLDAAETTTSNLSLPSQENLKLMPTTGLPEENVHHDEQKLI
PCKLDSKAVNGLAIEWVPEQESNALARTEAGICSQASAHGTIDSSSAVDCSGETDYEAKNNVSIDSVSETKCHVIVSPSSRRSNGTRKSSQKTQTKRGAR
KCRNTTKVPNLHRGIEIVFKSVTRRRSCFSKPARSSAWGLLGNITQTFMLINGLRPDEIENLGSQKARGDQGSGKRNKLAGGTSRRSSKKGHASAHCIRL
KVKVGKDACQTESNPKMIIPEVINTKASGDLVSDYGAESCQETSFEISKLAYCVGDNMVEEGTQKQLQSFYIKLGKAKAHCDASAMDVKLANKDMEGTVI
SEKSSRDIMEDYLGVPSHTEVEALGVATEKRYTDAGTSPDSEVINSVPEVQVNARCQEDYPDAVLSPSKAFAADEEGTGGKRGKKKESLPQAGNCSPAVA
SLKKVKLAKKRGGRQRKGDSLSSSEILTSCTSANGSVNTTSTKEYSAELVLSSGKTELGDPEGALRGEIIMETKICGELDADVRSSESQISKNPLPSTKS
RGRRLPRKSDGVNKRRSKVSDSAKSRRANGCKERGNDRKSVKKNKAEEKSVCDHVVYKDDNGKTDAGNDTTAEEVTNLDMPSSGVMEQNLFPDNAWVRCD
DCLKWRRIPVRLVESISQTHRQWICEDNMDKAFADCSFPQEKSDAEINAELGISDADEDVCDAPSNYMELECGPTSVSKEYEFTRITTNQFLHRTRKTQT
IDEIMVCYCKAPVGGRLGGCGDECLNRMLNIECVQGTCPCGDLCSNQQFQKHNYAKMTWDRCGKKGFGLRLEEDITRGQFLIEYVGEVLDVHAYEARQKE
YASKGHKHFYFMTLDGSEVIDACVKGNLGRFINHSCDPNCRTEKWVVNGEICIGLFALRDIKKGEEVTFDYNYVRVVGAAAKRCYCGSPQCQGYIGGDPT
SSEVTDQVDSDEEFPEPVMLEDGEVGDGLKNKISKTSFFGLSKGREMESKTAVGNLEVATEIKDSMNQSTPAISQSPSESEMNGLPGDFSSSSKRVEISP
QTEDMTTQPTPAVQQEISMEEMMDKSLYSSQKLKTSLTSVLIKPLPDDIMINRKSKSATAENKRVFVKSRFIIKTPPQSGLIKKGKSASNFININKVQTI
TNKPHMPPIKPKKLSESTSDGHFEAVQEKLNELLDSEGGISKRKDAPKGYLKLLLLTAASGAIRNGEAIQSNRELSMILDALLKTRSRMVLMDIIEKNGL
RMLHNIMKQYRRDFKKIPILRKLLKVLEYLAVREILTLEHINGGPPCPGMESFRESMLSLTEHNDKQVHQIARSFRDRWIPRQVRKLGYMDRDGGRMEIQ
RGSNCNKVLASHSQWHDQGVRHLEALNGTVESNLATTSVGTAVHEDSSANRVGSGTRTRKRKSRWDQPAEENIASRSLQHVEQNESGLLQQSESNSLPEL
SKEVPDHVDKAGGEYSYCPHCVHSYCWQDEASGADNGRQNIHEDVPPGFSSPIDPALVSNASSTVDDLPHQNVFHLKFPVGVVVGLPQRKFNSRFPVSYG
IPLPVVQQLGSPLAETVEGWIVAPGMPFHPFPPLPPLPSCKKGTLPSAMNSMEIDDTADRGKQDCYDRTTCLDENSPSTTGANQPDLNSPGPKDHQTFKR
ARGSYDLGRRYFRQQKWTKMLPPWVRSRNGWGCIGGNSRGGMCSTDLGSLTNEQRNSYY
|
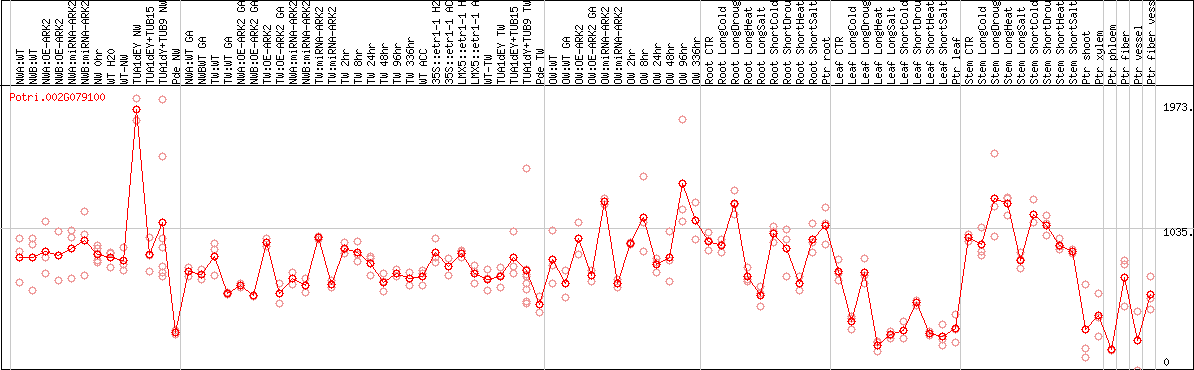
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G079100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
