External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G18520 107 / 5e-28
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT2G03880 77 / 3e-17
REME1
required for efficiency of mitochondrial editing 1, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G50420 75 / 2e-16
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G23330 69 / 1e-14
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G21065 69 / 2e-14
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT1G56690 69 / 2e-14
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G08070 68 / 3e-14
EMB3102, OTP82
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G16835 67 / 6e-14
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G32430 67 / 9e-14
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G22690 66 / 1e-13
unknown protein
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G170300
180 / 1e-54
AT4G18520 717 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.004G208000
82 / 5e-19
AT2G03880 939 / 0.0
required for efficiency of mitochondrial editing 1, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.007G104700
75 / 1e-16
AT2G13600 834 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.016G096400
71 / 3e-15
AT5G03800 993 / 0.0
embryo defective 1899, EMBRYO DEFECTIVE 175, EMBRYO DEFECTIVE 166, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.013G005400
71 / 4e-15
AT1G56690 1018 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.018G085800
70 / 6e-15
AT4G02750 491 / 3e-164
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.012G048800
70 / 7e-15
AT4G02750 1099 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G006400
70 / 7e-15
AT1G56690 993 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.018G067500
70 / 9e-15
AT4G13650 590 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029482
131 / 1e-36
AT4G18520 624 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10039594
129 / 1e-35
AT4G18520 619 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10002054
78 / 1e-17
AT1G74600 930 / 0.0
organelle transcript processing 87, pentatricopeptide (PPR) repeat-containing protein (.1)
Lus10024220
77 / 3e-17
AT1G74600 924 / 0.0
organelle transcript processing 87, pentatricopeptide (PPR) repeat-containing protein (.1)
Lus10001617
75 / 1e-16
AT2G03880 840 / 0.0
required for efficiency of mitochondrial editing 1, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10022974
75 / 2e-16
AT2G03880 542 / 0.0
required for efficiency of mitochondrial editing 1, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10017446
72 / 2e-15
AT1G11290 1152 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10038187
71 / 4e-15
AT3G25060 630 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10008637
70 / 7e-15
AT5G03800 902 / 0.0
embryo defective 1899, EMBRYO DEFECTIVE 175, EMBRYO DEFECTIVE 166, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10019213
67 / 6e-14
AT2G21090 376 / 2e-123
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Potri.002G091101.1 pacid=42779444 polypeptide=Potri.002G091101.1.p locus=Potri.002G091101 ID=Potri.002G091101.1.v4.1 annot-version=v4.1
ATGGGTACTTTTTTAGATGATGAAGCTTTAGGGCTGTTTAGGGAGGCCATAAGGGATGGGGTTGTACCAAATAGCAAGGCTTTTGTTTTTGTATTGAATT
TGTGTAGTGGAAGATTAGATTTTGAGTTAGGAAGACAAGTCCATGCTCGGGTTGTAAAGGGTGATTGGAGGAATTTGATCGCGGATGGTGCCATTGTTTA
TTTATATGTTCAATGTGGTGATTTGAAGAGTGCATTTGGTGTGTTTGATCGGATGGTGGAGCAGGATGTGGTTTCTTGGACAACAAATTTTACTACATCT
GGCGCTTCAAAGGCATTTGGAGAGGAGAAGGCATTGAAGTTTGAGAGACAGATGCACGGGTATAGCGAAGAAGATATACCAAGATGA
AA sequence
>Potri.002G091101.1 pacid=42779444 polypeptide=Potri.002G091101.1.p locus=Potri.002G091101 ID=Potri.002G091101.1.v4.1 annot-version=v4.1
MGTFLDDEALGLFREAIRDGVVPNSKAFVFVLNLCSGRLDFELGRQVHARVVKGDWRNLIADGAIVYLYVQCGDLKSAFGVFDRMVEQDVVSWTTNFTTS
GASKAFGEEKALKFERQMHGYSEEDIPR
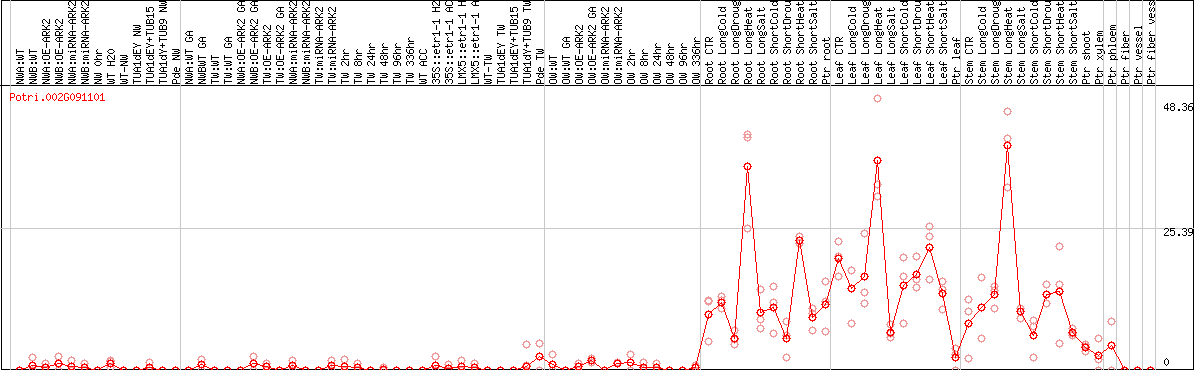
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G091101 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.