Potri.002G104100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G104100.8 pacid=42778687 polypeptide=Potri.002G104100.8.p locus=Potri.002G104100 ID=Potri.002G104100.8.v4.1 annot-version=v4.1
ATGTTCCGTGAAGGACCTGACAGCCGTTGGGCTCGAGAGAAGCAGAAAGAAGCAACTGCATTCGCACATTCAAGTTTAAGACCTAGAATTGATCCAATTA
GAAACGGCCGAGGGTTTGGCTTACCTCCAGCTTCTAAATTCAGAAGTGGACACTTGCCATCCTCGGCAATTCCGTTACCTAGAACCCTACCGCCTGATGC
CGATGATAGCCGGTCTGTCTCAGACAATGACATGGTCACGGAGTCAGACGAGGATGATGTTTATGGTGGACGGTATTCCTTAGATTCATCTCCACAGGAC
GAAAAAGTCCCCAATTCCACCACTAATCAACGAAGATATGGAAATGCTGCCCGAAGGACCTCGCGATATGCCAGTGATTATGGGTACTCTGATGTTAGTT
CTTCTATGGAGACTGTGGCAGGTAGAGGGGGGAATTTCTCTGAGAGTTTGGTTAGAGGGAATGCGCGGTATGCTTCTGTTGGAAGGAATGGTTATACGGA
GGATGAGGAAGAGGGATCAGATTCGGCAGGTAGTTCAGAGTTTTCTGCCAGTCAAGTTGGGAGTGTTAGTAGTGCTTTGCCGAGGAGTAAATTACATGTT
TCAGAGGGTTATGCTTCAAGTGTACCTTCTCAAGCTAATGTAGAAACTGTTGCTGCCAAGGATTTGCATTCCAGAAACCTGAAGAATAATAAATTTTCTC
ATGATGATGACATTCCCAGTGCGCCTCCATTTTGTGGAGGTCAGGAAATTAAAGAAGGTGCCCAAAAAGCATTTGGAATACATGAAGCAGCTGGCCCTGA
AAATTCACATGGTCTCTATACCAATAATGATCCAAATAAAATAAAAAATGCAACCGGAGTTGAACTGAAGGACAATAGTGGAGACCAGAATCCAGATAAG
TTTGTAAGAGCAACTGCTGGTGCAGAAGCTGGCACATCTGGCTCAAACCCAGCTCGAGTCCCAACATTTCATGCAAGTGCATTGGGGCCATGGCATGCTG
TAATTGCATATGATGGGTGCGTGCGGCTTTGCCTTCATGCTTGGGCTAGGGGTTGCATGGAAGCACCAATGTTCTTGGAAAATGAATGTGCTTTACTACG
GGAGGCATTTAGTGTACATCACGTGCTACTACAATCAGAAGAGGAACTATTGGCAAAGCGCTCTTCTGAGCTTGTTTGTGAGGGAGCTGCCCCAAAGCCT
AAGAAAATAATTGGGAAGATGAAGGTGCAAGTCCGTAAAGTTAAAACAAGTCTCGACCCTCCCAGTGGCTGTAGTATTTCAGCTCTTTCAGCACCAAAGC
TAAAATTGGATGTGGTCCAGTATCGCTTATCCAAGTTTCAGTCATCGCTTTCTTCTGCTTGGAAAACTTTTCGAAAAATCCGTGTCGCCCCTCGTGTGCC
TGCAAATGGTTCTTTCTCACGCCAAAGCTTGGCATATGTGCATGCCAGCACTCAGTACATCAAACAGGTTTCAGGACTTCTGAAGATTGGTGTTACTAGT
CTACGCAACAGCTCATCATCATATGAAGTTGTGCAAGAAACTTACTCCTGCTCATTGAGACTGAAAAGTTCAGCAGAAGAGGATGCCATCAAGTTGCAGC
CTGGATCTGGTGAAACTCATGTATTCTTTCCAGATAGTCTAGGGGATGATCTAATTGTTGAAGTACTAGATTCCAAAGGGAAATATTATGGCCGTGTCCT
TGCTCAAGTCGCTAGTATTGCTGAAGACTCGGTTGATAAGCTCCGCTGGTGGTCTATATACCGAGAACCAGAACATGAGCTTGTGGGCAAACTGCAGCTC
TACATAAATTACTCAACAAGTTCAGATGACAGTAATCTCAAGTGTGGCTCTGTAGCCGAAACTGTTGCGTATGACCTCGTTCTAGAAGTTGCCATGAAGG
TTCAACATTTTCAACAAAGGAATCTATTGCTTTATGGTTCATGGAAATGGCTCCTAGCTGAATTTGCCACATATTATGGAGTATCCGATGTCTATACCAA
GCTGAGGTACCTTTCATATATAATGGATGTAGCTACACCAACAGCTGACTGCCTGACTTTGGTGTATGATTTGTTAAAGCCTGTTATTATGAAGGGCCAC
AACAAGAGCATGCTGAGTCATCAAGAGAACCGAATCCTGGGAGAAATCAAGGATCAGATTGAACAGGTTCTTTCTGTTGGTTTTGAGAACTACAAGTCTC
TTGATGAATCATCACTGTCTGGTATTATGGATGTGTTCAAGCCTGCAACTGGGCTTGCAGCACCTGCCTTGGAGCCTGCTGTCAAGCTTTATACTCTTCT
TCACGACATACTGTCTCCGGAGGCACAAACAAATTTAACTCATTATTTCCAGGCTGCAGCCAAGAAAAGATCAAGAAGGCACTTGACCGAGACAGATGAA
TTTGTTAACAACAATAATGAAGCAACATTGATGGATTCTGTTGCTATGTCTACTGCTTATCAGAAGATGAGTTCTCTTTGTATGAACATAAAAAATGAAA
TTCAGACTGACATCGAAATACACAATCAACATATACTTCCCAGTTTCATAGACCTTCCAATTTTATCTTCGTCCATATACAGCACAGAGCTTTGTAGTAG
ACTGCGTGCTTTCTTGCTTGCATGCCCCCCATCCGGTCCTTCACCCCCTGTAGCAGAACTTGTTATTGCAACAGCTGATTTTCAGAGGGACCTTGCAAGC
TGGAACATTAGTCCTGTGAAAGGAGGAGTGGATGCAAAAGAACTGTTTCATCTGTATATCATGATATGGATACAAGACAAGCGCCTTTCTTTGCTGGAAT
CGTGCAAATTAGACAAGGTAAAATGGTCAGGTGTCAGGACGCAACATTCTACAACCCCCTTTGTGGATGACATGTATGACCGCCTTAGAGATACACTGGA
ACAGTATGAAGTAATTATATGCAGATGGCCAGAGTATATCTTTGTTCTAGAGAATGCTATTGCTGATGTCGAAAAGGCTATAGTGGAAGCTTTGGACAAG
CAATATACAGACGTTCTAGCACCATTGAAGGAGAACTTGGAACCTTCGAAATTCGGCCTCAAGTATGTGAAGAAGCTTACAAAACGATCTGTTTGCTCAT
ATATTGTCCCTGATGAGCTTGGAATTCTGCTTAATTCCATGAAGAGGATGCTTGATGTCCTGCGTCCCAAGATAGAAACTCAATTTAAGGCATGGGGTTC
TTGCATGCCTAATGGTGGACACACTGCTCCTGGAGAGCGTCTTAGTGAAGTGACTGTCATGCTAAGAGCCAAATTTAGAAGTTATCTGCAGGCTGTTGTA
GAAAAGCTAGCAGAGAATACCAAATTACAGAATCCTACAAAATTGAAAAAGATTCTCCAAGAATCAAAGGAATCAATGGTTGAATCTGATATTCAAAGTA
GGATGCAGCCACTGAAAGACCAGCTGACAAACACAATTACCCACCTGCAATCTGTTTTTGAGACTCATGTCTTTGTTGCTATCTGCCGAGGTTACTGGGA
CCGAATGGGGCAGGATGTTCTGAGCTTCTTGGAGAACAGGAAAGAAAATCGATCATGGTATAAAGGTTCACGAATTGCAGTGTCTGTCTTGGATGATACC
TTTGCCTCGCACATGCAGCAGCTGCTCGGAAATGCACTGCAGGAGAAAGACCTCGAGCCTCCTAGATCGATCATGGAAGTTCGCTCAATGCTCTGCAAGG
ATGCTCCAAATCACAAAGACAGCACCTATTACTATTAA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G104100.8 pacid=42778687 polypeptide=Potri.002G104100.8.p locus=Potri.002G104100 ID=Potri.002G104100.8.v4.1 annot-version=v4.1
MFREGPDSRWAREKQKEATAFAHSSLRPRIDPIRNGRGFGLPPASKFRSGHLPSSAIPLPRTLPPDADDSRSVSDNDMVTESDEDDVYGGRYSLDSSPQD
EKVPNSTTNQRRYGNAARRTSRYASDYGYSDVSSSMETVAGRGGNFSESLVRGNARYASVGRNGYTEDEEEGSDSAGSSEFSASQVGSVSSALPRSKLHV
SEGYASSVPSQANVETVAAKDLHSRNLKNNKFSHDDDIPSAPPFCGGQEIKEGAQKAFGIHEAAGPENSHGLYTNNDPNKIKNATGVELKDNSGDQNPDK
FVRATAGAEAGTSGSNPARVPTFHASALGPWHAVIAYDGCVRLCLHAWARGCMEAPMFLENECALLREAFSVHHVLLQSEEELLAKRSSELVCEGAAPKP
KKIIGKMKVQVRKVKTSLDPPSGCSISALSAPKLKLDVVQYRLSKFQSSLSSAWKTFRKIRVAPRVPANGSFSRQSLAYVHASTQYIKQVSGLLKIGVTS
LRNSSSSYEVVQETYSCSLRLKSSAEEDAIKLQPGSGETHVFFPDSLGDDLIVEVLDSKGKYYGRVLAQVASIAEDSVDKLRWWSIYREPEHELVGKLQL
YINYSTSSDDSNLKCGSVAETVAYDLVLEVAMKVQHFQQRNLLLYGSWKWLLAEFATYYGVSDVYTKLRYLSYIMDVATPTADCLTLVYDLLKPVIMKGH
NKSMLSHQENRILGEIKDQIEQVLSVGFENYKSLDESSLSGIMDVFKPATGLAAPALEPAVKLYTLLHDILSPEAQTNLTHYFQAAAKKRSRRHLTETDE
FVNNNNEATLMDSVAMSTAYQKMSSLCMNIKNEIQTDIEIHNQHILPSFIDLPILSSSIYSTELCSRLRAFLLACPPSGPSPPVAELVIATADFQRDLAS
WNISPVKGGVDAKELFHLYIMIWIQDKRLSLLESCKLDKVKWSGVRTQHSTTPFVDDMYDRLRDTLEQYEVIICRWPEYIFVLENAIADVEKAIVEALDK
QYTDVLAPLKENLEPSKFGLKYVKKLTKRSVCSYIVPDELGILLNSMKRMLDVLRPKIETQFKAWGSCMPNGGHTAPGERLSEVTVMLRAKFRSYLQAVV
EKLAENTKLQNPTKLKKILQESKESMVESDIQSRMQPLKDQLTNTITHLQSVFETHVFVAICRGYWDRMGQDVLSFLENRKENRSWYKGSRIAVSVLDDT
FASHMQQLLGNALQEKDLEPPRSIMEVRSMLCKDAPNHKDSTYYY
|
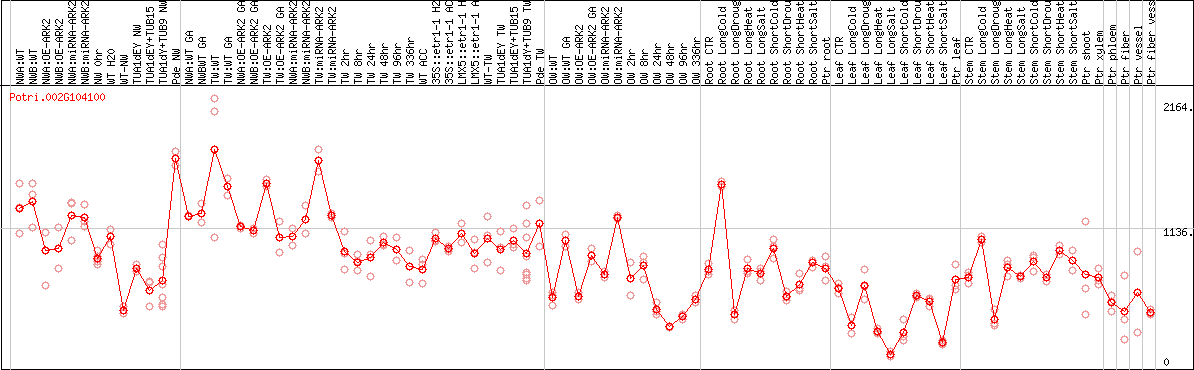
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G104100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
