External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G34150 185 / 3e-58
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT1G63220 57 / 5e-10
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
AT3G55470 51 / 7e-08
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
AT5G47710 45 / 7e-06
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
AT1G03370 46 / 2e-05
C2 calcium/lipid-binding and GRAM domain containing protein (.1)
AT1G05500 45 / 2e-05
SYT5, NTMCTYPE2.1, ATSYTE ,NTMC2T2.1 ,NTMC2TYPE2.1
synaptotagmin 5, ARABIDOPSIS THALIANA SYNAPTOTAGMIN HOMOLOG E, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT4G05330 44 / 8e-05
AGD13
ARF-GAP domain 13 (.1)
AT5G11100 43 / 0.0001
SYT4, NTMCTYPE2.2, ATSYTD ,NTMC2T2.2 ,NTMC2TYPE2.2
synaptotagmin 4, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT3G61050 43 / 0.0002
AtCLB, NTMCTYPE4 ,NTMC2T4 ,NTMC2TYPE4
calcium-dependent lipid-binding protein, Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G301900
213 / 2e-69
AT4G34150 204 / 8e-66
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.009G097900
204 / 2e-65
AT4G34150 213 / 5e-69
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.002G155600
57 / 3e-10
AT1G63220 202 / 1e-67
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Potri.010G203000
56 / 2e-09
AT3G55470 167 / 6e-54
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Potri.012G056000
52 / 1e-07
AT3G18370 971 / 0.0
C2 domain-containing protein (.1)
Potri.015G046700
51 / 3e-07
AT3G18370 968 / 0.0
C2 domain-containing protein (.1)
Potri.006G063900
50 / 9e-07
AT1G05500 847 / 0.0
synaptotagmin 5, ARABIDOPSIS THALIANA SYNAPTOTAGMIN HOMOLOG E, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.018G124000
47 / 8e-06
AT1G05500 877 / 0.0
synaptotagmin 5, ARABIDOPSIS THALIANA SYNAPTOTAGMIN HOMOLOG E, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.018G025000
46 / 1e-05
AT5G11100 824 / 0.0
synaptotagmin 4, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10027875
201 / 2e-64
AT4G34150 246 / 3e-82
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10002825
190 / 6e-60
AT4G34150 243 / 5e-81
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10014088
173 / 7e-53
AT4G34150 202 / 4e-64
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10019826
174 / 1e-52
AT4G34150 191 / 2e-59
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10029565
63 / 2e-12
AT1G63220 221 / 1e-75
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Lus10002235
62 / 5e-12
AT1G63220 216 / 2e-73
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Lus10040291
54 / 8e-09
AT3G55470 191 / 4e-63
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Lus10009032
51 / 5e-07
AT3G18370 907 / 0.0
C2 domain-containing protein (.1)
Lus10039113
46 / 5e-06
AT5G47710 239 / 6e-82
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Lus10006193
47 / 6e-06
AT5G11100 862 / 0.0
synaptotagmin 4, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0154
C2
PF00168
C2
C2 domain
Representative CDS sequence
>Potri.002G123200.1 pacid=42778470 polypeptide=Potri.002G123200.1.p locus=Potri.002G123200 ID=Potri.002G123200.1.v4.1 annot-version=v4.1
ATGGCGATCTGTGGAATCCAAGGCTTTCCTCTTGAGGTCACTGTCGTTGCTTGTTACAATTTGGAGGACAAGGAATGGATATCAAGACAAGACCCATATG
TTAGTGTTGAGTATGGAAACACCAAGTACCGAACCAAAACGTGCACAGATGGAGGCAGGAACCCCGTTTTTCAAGAAAAGTTCATTTTCACACTTGTTGA
AGGCCTCCAGGAATTGAGCGTTGTGGTTTGGAACAGCCACACCCTCTCTGCTGATGAGCATATTGGCACCGGAAGAATCCAGCTCCATAAAGCCCTTTCC
CAAGGTTTTGATGATGCTTCCTGGCCCATACAGAGCAAAACTGGCAGGCATTCAGGGGAAGTGCGGCTTATGTTACACTACTCAAATCCCAATCAACATA
AAGGGAAAATGACTCCATCAGATTTAGGATCAAAATATGCATCGCCACCACTGAACCAGGTTCTGCCATACCCTTCCATGCCACCAGCACCCGCTGCTCT
TTACCCTGCTACAACTACCTACTCCTCCCCTTCTCCGTACATGGGGTACTCTCCTAACCCTGCGAGCTATCCGCCACCCCCTCACGTTCCTCCTCCAGCT
GGCCGCTACCCTCCGCAAGCGTGTTTTCCTGCAGCCTACCCTCCCCAAGCATATCCACCTGCTCCTCAGCCCTCAACGTACTATCCTACAGGTGCATCCG
GGGTTTATCCACCACCACCACCACCAGGAACATATCCACCGCCGCCGTACTGA
AA sequence
>Potri.002G123200.1 pacid=42778470 polypeptide=Potri.002G123200.1.p locus=Potri.002G123200 ID=Potri.002G123200.1.v4.1 annot-version=v4.1
MAICGIQGFPLEVTVVACYNLEDKEWISRQDPYVSVEYGNTKYRTKTCTDGGRNPVFQEKFIFTLVEGLQELSVVVWNSHTLSADEHIGTGRIQLHKALS
QGFDDASWPIQSKTGRHSGEVRLMLHYSNPNQHKGKMTPSDLGSKYASPPLNQVLPYPSMPPAPAALYPATTTYSSPSPYMGYSPNPASYPPPPHVPPPA
GRYPPQACFPAAYPPQAYPPAPQPSTYYPTGASGVYPPPPPPGTYPPPPY
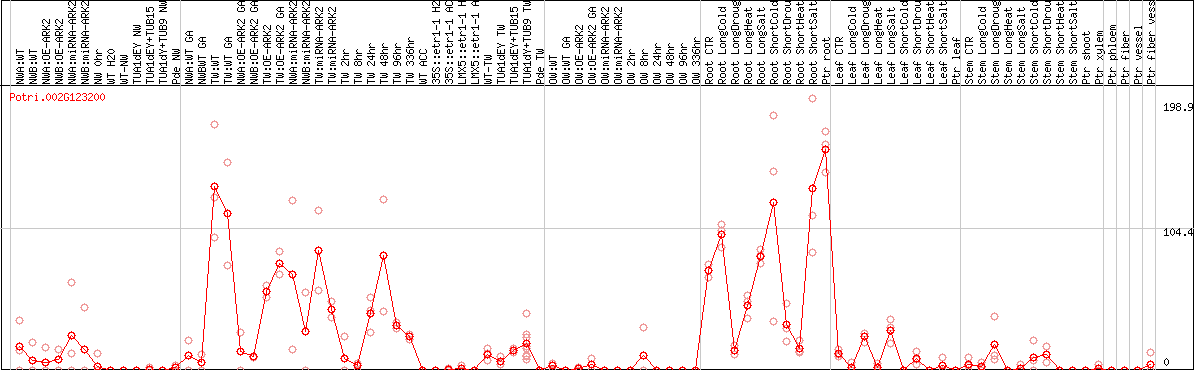
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G123200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.