Potri.002G131400 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G131400.1 pacid=42779564 polypeptide=Potri.002G131400.1.p locus=Potri.002G131400 ID=Potri.002G131400.1.v4.1 annot-version=v4.1
ATGGCGAAATCGAGCGCCGATGACGAGGAGCTAAGGAGAGCCTGCGAGGCAGCAATTGAAGGCACAAAACAGAAGATCGTATTGTCGATCCGTGTTGCCA
AGAGCCATGGGATCTGGGGCAAATCTGGGAAATTAGGACGTCACATGGCCAAACCTAGGGTTCTCTCTCTCTCCACAAAATCTAAGGGTCAGAGGACGAA
AGCCTTTCTTCGAGTGCTAAAATATTCAACTGGAGGAGTACTTGAGCCAGCAAAGTTATACAAACTCAAGCATCTTTCCAAAGTGGAGGTTATAGCTAAT
GATCCCAGTGGATGCTCGTTTACCCTGGGATTTGATAACCTTAGAAGCCAGAGTGTTACTCCACCGCAATGGACTATGCGCAATATTGATGATAGGAACC
GCCTTTTATTTTGCCTCTTAAACATCTGCAAAGATGTCCTTGGCCGTCTTCCAAAAGTTGTTGGCATAGATGTTGTGGAGATGGCACTTTGGGCTAAGGA
AAATACACCCGCAGTTCCCAAGCAAACGAATCAACAGGATGGAGTGCCTGTTGCAGCTACTGTCACAGAAAGTGACTTGAAAGTGACTGTTGAAAGAGAA
CTTGTCTCCCAAGCGAAGGAGGAGGACATGGAGGCTCTTCTTGGCAATTATTTAATGGGTATTGGTGAAGCAGAGGTGTTTTCAGAAAGGTTGAAGCGTG
AGCTTCTTGCTCTGGAAGCTGCAAATGTGCATGCCATTTTAGAAAATGAACCTTTAATAGAAGAGGTGTTGCAAGGGCTTGAAGCTGCAACATATTGTGT
TGATGACATGGATGAGTGGTTAGGCATTTTCAACGTGAAACTTAGACATATGAGAGAAGACATTGAATCAATAGAAACCCGCAATAACAAATTGGAGATG
CAGTCTGTAAATAATGTATCACTCATTGAAGAACTTGATAAGCTACTTGAACGACTGCGGGTCCCTTCTGAATATGCAGCATGTCTAACAGGAGGTTCAT
TTGATGAGGCTCACATGCTTCAAAACATTGAAGCATGTGAGTGGTTAACTGGTGCTTTACGTGGACTTCAAGTGCCTAATTTGGATCCAAGCTATGCAAA
TACGCGTGCTGTTAAAGAAAAGCGCACAGAACTTGAAAAATTAAAAACTATGTTTGTCAGGAGAGCTTCTGAGTTCTTGAGGAACTACTTTGCAAGTTTG
GTGGATTTCATGATAAGCGACAAGAGTTACTTTTCACAACGTGGACAGTTAAAGAGGCCTGACCATGCTGACTTAAGGTACAAATGCAGGACATACGCTC
GCCTTCTACAGCATCTAAAGAGCCTTGATAAGAACTGCTTGGGTCCTTTGAGGAAAGCATATTGTAGCTCCCTTAATCTGCTTCTTCGTCGGGAGGCTCG
TGAATTTGCTAATGAGCTCCGTGCCAGTACAAAAGCATCAAGAAATCCAACTGTCTGGTTGGAAGCTTCTGCAGGCTCAAGTCATAGTTCACATAATGCA
GATACATCTGCAGTTTCTGAGGCTTATGCCAAAATGCTGACAATATTTATCCCACTCCTTGTAGATGAGAGTTCTTTTTTTGCACACTTCATGTGCTTCG
AGGTTCCTGCACTTGTTCCCCCAGGAGGTGTAGCTAATGGTAATAAAGGTGGTTACAATGATGCCGATGACAATGATGATTTGGGTATCATGGACATTGA
TGAGAATGATGGCAAAGCTGGTAAAAATTCTGCAGATCTGGCAGCACTAAATGAATCTCTTCAGGATTTGCTTAATGGAATCCAAGAAGATTTTTATGCT
GTGGTTGATTGGGCATACAAGATAGATCCTCTACGCTGCATATCAATGCATGGGATTACAGAGCGCTATCTCTCTGGTCAGAAAGCTGATGCTGCAGGAT
TTGTGCGTCTCTTGCTTGGTGACCTAGAGTCCAGAATTTCTGTGCAATTCAATCGTTTTGTTGATGAAGCTTGCCACCAAATTGAAAGAAATGAGCGTAA
TGTGCGGCAGATGGGGGTTTTATCCTACATCCCAAGATTTGCAACTCTTGCAACTCGTATGGAGCAGTATATCCAGGGACAATCTAGGGATTTGGCTGAT
CAAGCGCACACAAAATTTGTTAGCATAATGTTTGTGACTTTGGAGAAAATTGCACAAACAGATCCAAAATATGCAGATGTTTTCCTTTTAGAAAACTATG
CTGCATTTCAAAATAGCTTGTATGACCTTGCCAATGTTGTGCCAACTTTAGCCAAATTTTATCACCAGGCAAGCGAAGCTTATGAACAAGCTTGTACACG
TCATATCAGCATCATTATTCTCTATCAATTTGAAAAACTTTTCCAGTTTACTCGCAAGATTGAGGATTTGATGTTCACCATAACACCAGAAGAGATCCCT
TTCCAGCTTGGATTGTCAAAAATGGATCTCCGGAAGATGTTAAAATCTAGTTTATCTGGGGTTGACAAGTCTATTAGTGCAATGTATAAGAGGTTACAGA
AGAACTTAACTTCAGAAGAACTGTTGCCTTCTTTGTGGGATAAGTGCAAGAAGGATTTCTTGGACAAGTATGAGAGTTTTGCACAACTAGTTGCAAAGAT
CTATCCCAATGAAAGCATCCCTTCCGTGTCAGAAATGAGAGAGCTTCTGGCGTCCATGTGA
|
||||||||||||||||||||
|
AA sequence
|
>Potri.002G131400.1 pacid=42779564 polypeptide=Potri.002G131400.1.p locus=Potri.002G131400 ID=Potri.002G131400.1.v4.1 annot-version=v4.1
MAKSSADDEELRRACEAAIEGTKQKIVLSIRVAKSHGIWGKSGKLGRHMAKPRVLSLSTKSKGQRTKAFLRVLKYSTGGVLEPAKLYKLKHLSKVEVIAN
DPSGCSFTLGFDNLRSQSVTPPQWTMRNIDDRNRLLFCLLNICKDVLGRLPKVVGIDVVEMALWAKENTPAVPKQTNQQDGVPVAATVTESDLKVTVERE
LVSQAKEEDMEALLGNYLMGIGEAEVFSERLKRELLALEAANVHAILENEPLIEEVLQGLEAATYCVDDMDEWLGIFNVKLRHMREDIESIETRNNKLEM
QSVNNVSLIEELDKLLERLRVPSEYAACLTGGSFDEAHMLQNIEACEWLTGALRGLQVPNLDPSYANTRAVKEKRTELEKLKTMFVRRASEFLRNYFASL
VDFMISDKSYFSQRGQLKRPDHADLRYKCRTYARLLQHLKSLDKNCLGPLRKAYCSSLNLLLRREAREFANELRASTKASRNPTVWLEASAGSSHSSHNA
DTSAVSEAYAKMLTIFIPLLVDESSFFAHFMCFEVPALVPPGGVANGNKGGYNDADDNDDLGIMDIDENDGKAGKNSADLAALNESLQDLLNGIQEDFYA
VVDWAYKIDPLRCISMHGITERYLSGQKADAAGFVRLLLGDLESRISVQFNRFVDEACHQIERNERNVRQMGVLSYIPRFATLATRMEQYIQGQSRDLAD
QAHTKFVSIMFVTLEKIAQTDPKYADVFLLENYAAFQNSLYDLANVVPTLAKFYHQASEAYEQACTRHISIIILYQFEKLFQFTRKIEDLMFTITPEEIP
FQLGLSKMDLRKMLKSSLSGVDKSISAMYKRLQKNLTSEELLPSLWDKCKKDFLDKYESFAQLVAKIYPNESIPSVSEMRELLASM
|
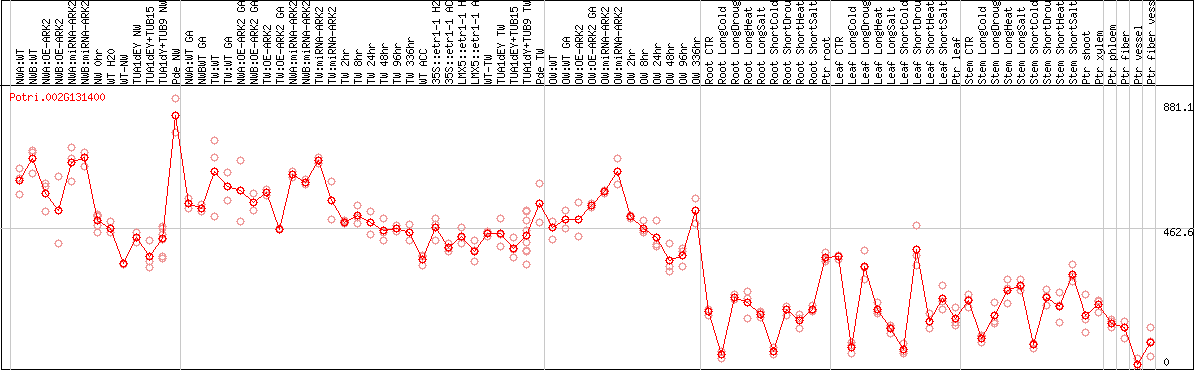
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G131400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
