Potri.002G140200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G140200.2 pacid=42778937 polypeptide=Potri.002G140200.2.p locus=Potri.002G140200 ID=Potri.002G140200.2.v4.1 annot-version=v4.1
ATGTCCTACAATCAATCAAGGGGTGGCAGTGATAAGAGCGATTTACAATATAGGAAACCAGGACGATCCGTTAGCTCTAGTCAGCAACGGACTTCCTCTG
TTTCTCACGGTAAGGGCGGTGGCCCCCCCGTCCCTTCACCGTCTTCTTCTTCTCTTTCTTCCAATCGCAGTTTTAATAAGAAGCCTAGTAATCTTACGCA
AGGAGGAGGGCAATCTAGTAGGGTAAATTTGCCTTCTGGTGTGAATTCTTCTGATTCCGGTAATAATGCTGCTTCTACAATCCGTAATGTGCAGAATGGT
GTCCTGACGCAACACCAATCACATGGAACATCTGATGCAAGCAGTGTTGCCAAGCCGACAGAAGCATCGGCTGCTCAGAGAAGCACCCGAGATGTTCCAA
AGGCTCCAACTTCTCAACCTGCCGCCATAAGTTCGGAGAGTGGAGCTCACATGACACCTGCCAAGGCCCCCTTAGATTCATCTAAGGCATTTGCTTTTCA
ATTTGGGTCCATTAGTCCTGGATTTATGAATGGGATGCAGGTTCCTGCTCGAACAAGCTCTGCGCCCCCTAATTTGGACGAGCAGAAACGTGACCAGGCA
CATCATGATACATTTAGGCCTGCTCCTTCATTACCGACTCCTGCTCCTAAGCAGCAATTGCCCAGGAAGGAAGTTAGTTCATCAGTCCAAACTAGCACTG
GAGAGGTTCATTTAGTTCCCAAGGCCAGCAAGGAGACACAACTCCCACCTGCACCATCTGTGAGCCAGACACAGAAGCCTTCTGTTCTTCCGATTCCTAT
GAATTCTTTGCAGATGAAATATCAGCAGCCACCAGTCTCGGTTCAGTTTCGTGGTCCAAGCCCACAAATCCAGTCTCAGGGTGTCCCAGCAAATTCACTT
CATGTTCCAATACAATTACCTATGGGGAATGCTCCTCAAGTGCAACAGTCGGTGTTTATTCAAGGTCTCCAACACCATCCTATGCAGCCTCAAGGAATGA
TGCATCAGAGCCAAACCATGAGTTTTACGAATCCAATGGGTCCTCAGATACCTCAGTTAGGCAGCTTAGCATACGGCATGACCTCCCAATATTCCGCACA
GCAGGGTGGTAAATTTGGCAGTCCACATAAAACGCCTGTCAAGATTACTGATCCAAAGACACATGAAGAGCTGAGGCTTGATAAGCGAACAGATGCATAT
CCAGATGCTGGATCATCAGGACTGAGGTCTCATCTCAATGTTCCCCAGACCCAGCCAATTCCATCATTTGCACCTTCTCGTCCAATTAACTATTATCCTA
GTTCCTACAATGCCAGTAATTTATTTTTCCCAGCTCCAAGCTCTCTACCATTAACTGGTAGTCAAATAGCTCCTAATTCTCAGCTACCACCTAGGTTTAA
CTATCCAGTTAGCCAGCCCCCCCAAAATGCACCATATATGAATGCATCTGCTCTTAATTCGCTTCCACTTAGTAAGTCTGGGACAGTATCTCATGGTGTT
GCAGAACCACAGAACTCAGAACATGCTCGTGATGCTCGTAATGCAATCTCCTTGACTCCATCTGGGGCGGTGCAAGTGACTGTTAAACCAGCTGTTGGCT
CTCATGGAGAGAAAGTTGTGGAGCCATCTTTTCCAAAGATCTCCTCTATTGTTGAAAAGGGTGGATTCTTCAAATCCTCAAGGTCATCTGGGGAAGCTAG
CCCATCTCACTCTCAAAGGGATTCAGAGGCTTCTTCAGAAAGCTCTTTACAGCGGATAAAATCTGGTGGTGAATCATTGGTGAAGCCACTGCCAGTGGCA
GCTAAACAGCCTGCAGCAGTTGCTGTTGACGGTGCAGCTTCTGCTTCACTAGCTCAGTGTGAGGAGGCAATACCAAGTGTATCTAATGCTGAGGGCCGAA
AAAAGGAAGCTCTGAGTGGGTCAAACTTTATCAAGGAACATCAGAAGAAACCAGGCAAGAAAGGAAATATCCAGCCTCAGCATCAGATTGGTGGACAGAC
AACCTTGTCTTCTCACACTTTGGAACATGGTGTATCTTCTGGCACTGGGGTTTCTGAAACTGCAGAAAATGAGAAAAGTCCTCCATCATTAGCAAACAGT
GAAGTTTTGACAAAATCAATCAAGGAACCAGTGTCAACGATTGCTGCCTGGAATCCTGATGTTTCTGAAACCAAGGTTGACAATGCTGGAGATGCTTTCG
ACAGTGTTTCATCTCAAGTTCCTGTTGCTGGTATTGCTCATACCACACACATTTCTCCTCATGCTAAGCTGGATGATTCTTCACAGCTGGAAAAACTAAA
ATGTGAAATTCCAGCAACAGAAGACGAAATAGAGAAATCTCTGTCTGAATGTCCTAAACAAGATTACAACATTTCTTCAGCATCAATTAACTCAAAATCT
GCAGACCAGGTCAAGCAAGATAAAGAGGTATCTGATTCAGTGGTCACATCTGTTGGCAATGAGGTTCCAGCCTCAGAAACTGCCCAGGAGGGGCTGGTAG
AACCCGTGACTTGCCACACTGCAAATGACCACATATCAGATAATGCGGGTGCTTCCACATCCAGAAAGTTCAATTCTGCAGATGACATAAAGCCTTTGGA
TGCATCTTTGAGTCACAGTGATAACATAGGTAATAAAGAAGCTTCTGTTACCAAATCTGGTATTTCTGGTCATCAGGGTTCTCCACCAGTGCCTGATCTC
TCCGAGGCAACTGCAAAACATGAAGGGGAAGGTGCAGAAAATGCTGGTAGTGGAACAGTCCCCCTTGAAGTTTCTGGTTACAAAGAAAAGCCTTCTGAAT
TGACTAGATCAAAGAGTACCGCTAATAGAATGAAGAAGAAGAAAAAAGAATTTCTTCTGAAAGCAGATCTTGCTGGGACGACTTCTGATCTTTATGGGGC
ATACAAAGGCCCTGAGGAGAAGAAAGAAAATGTCATCTCTTCAGAAGTCATAGAAAGCACTAGCCCTAATCTGAAGCAGGCACCTGCTGATGCCCTTCAG
GTTCAGACTGTAGCGAGTGAGAAAAGCATGCAGAATAAAGCTGAGCCAGATGATTGGGAAGATGCCACAGACATGTCTACACTGAAATTGGAAAGTTTAA
TTGATGGGGAACTATCACTTGGAGGATTGGGACAACATGATACAGATGGGAATGCCAACAAATTAAAAAAATATTCCAGAGATTTCCTCCTTAAATTTTC
TGAGCAATGTACTGATCTTCCTGGAGGTTTTCAAATTCCATCTGATATAGCAGGCTCCTTGATGGGTGTCGGTGTTTCTCATCTTGCTGATCGTGATCCA
TGCCCCAGTCCTGCAAGGGTTATGGACAGGTCTAATAGTGGATCTCGAATAGACCGCCGTGGGAGTGGTATAGTTGATGATGGCCGATGGAGTAAACAGC
CTGGTCCATCCGGTCCTGGAAGGGATTTACACTTGGATATCAGTTATGGAGCTAATGTAGGTTTTCGACCTGTTGCAGGAGGCAACTATGGTGCTCTGAG
GAATCCACGAGCACAGAGTCCTGTACATTATGGAGGAGGGATTTTATCCGGACCCATGCAATCAATGGGTCCACAGGGAGGGCTGCAAAGAGGTGGCTTA
GATGCTGACAGATGGCAGCGTGCTGCTATTTTTGTCCATAAAGGATCATTTTCTTCTCCTCAGACTCCATTACAGACTATGCACAAAGCTGAGAAAAAGT
ATGAAGTGGGTAAAGTGACAGATGAAGAAGCAGCCAAGCAAAGACAACTGAAAGGCATATTGAACAAGCTAACACCTCAGAATTTTGAGAAGCTTTTTGA
GCAAGTAAAAGCTGTTAACATTGACAATGTTGTCACACTTAATGGCGTTATCTCACAGATCTTTGACAAAGCCTTAATGGAGCCTACTTTCTGTGAAATG
TATGCTAATTTTTGCTTTCATCTAGCAGCAGAGTTGCCCGAACTCACTGAAGATAATGAAAAGGTAACTTTCAAGAGGATACTTCTGAACAAGTGTCAGG
AAGAATTTGAGAGAGGGGAGCGAGAGCAAGAAGAAGCTAATAAAGCTGATGAAGAGGGTGAGATTAAACAGTCTGAAGAAGAAAGAGAAGAAAAGAGAAT
CAAGGCTCGTAGACGAATGCTTGGTAACATAAGACTTATTGGAGAGCTGTATAAAAAAAGAATGTTGACTGAAAGAATAATGCACGAGTGCATCAAAAAG
TTGCTGGGTCAGTATCAGAATCCCGATGAGGAAGATCTTGAGGCTCTTTGCAAACTAATGAGCACGATTGGAGAGATGATTGACCATCCCAAAGCCAAGG
AGCATATGGATGTATATTTTGACATGATGGCAAAATTATCGAACAACATGAAACTCTCATCTAGGGTTAGGTTTATGTTGAAGGATTCTATTGATCTGAG
GAAAAACAAATGGCAGCAGAGGAGGAAGGTTGAGGGGCCGAAAAAGATTGAGGAAGTGCATAGAGATGCTGCTCAAGAACGACAGCTGCAAACTAGTAGG
CTGGCTCGCAATCCTGGCATCAATCCTTCTCCAAGAAGGGGGCCTATGGATTTTGGTCCAAGAGGGTCAACAATGTTGCCTTCTCTGAATGCACAGATGG
GGGGTTTTCGAGGGTTTCCCACTCAAGTTCGTGGGCATGGCACTCAGGATGTTCGGTTTGAGGAAAAACAGTCTTACGAGGCTAGGACTATGTCTGTTCC
CTTACCTCAAAGACCCCTTGGTGATGATTCTATTACTTTGGGTCCCCAAGGTGGACTTGCAAGAGGAATGTCCATTAGAGGACAACCTGCCAGTATGGGT
ACTCTTGTAGCTGATATTTCTCCAAGCCCAGGTGACCCAAGAAGAATGGCAGCTGGTTTGAATGGTTCTAGTGCCATTTCAGGACGGTCAAATTACAGCC
CAAGGGAAGATATCATTCCAAGATATACTCCAGACAGATTTGCAGTTCCACCTGCTTGTGATCAAATGAATGGTCAGGAACGAAATATGAATTATGTTAA
CAGGGATCTGAGGAATCTAGATCATGGTTTTGATAGGCCTCTTGGATCATCACCACCAACAAGAGCACAAGGACCACCTTTTTCCCAGACCACTCCAACA
GGAAAGCTGTGGCCTGAAGAACGACTGCGAGATATGTCCACGGCTGCAATAAAAGAGTTTTACAGTGCCAGAGACGAGAAAGAAGTTTCTCTATGCATTA
AAGAATTGAATTCCCCAAGTTTCCATCCTTCAATGATCTCTATCTGGGTTACAGACTCATTTGAGAGGAAGGACTTGGAGAGAGATCTTTTGGCCAAGCT
TCTTGTCAGCCTTGCAAGATCTCAAAATGGCATATTGGATTCAAACCAGCTCATCAAAGGGTTTGAATCCATCCTGACTACTTTGGAGGATGCTGTAAAT
GATGCCCCCAAAGCACCAGAATTTCTCGGACGCATCATTGGCAGGGTTGTTGTAGAAAATGTTGTCCCTCTATCAGAGATTGGGCCGTTATTACATGAAG
GCGGAGAGGAACCGGGCAGCCTTCTCAAGTTGGGGCTTGCAGGGGATGTTCTTGGCAGTATTTTGGAGATGATCAAAGTGGAGAAAGGAGAAGCTGTCTT
AAATGAGATTCGTGGTGCCTCCAATTTGCGTTTGGAAGATTTCCGGCCTCCAGACCCTAACAGATCAAGGATACTAGAAAAGTTTATTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G140200.2 pacid=42778937 polypeptide=Potri.002G140200.2.p locus=Potri.002G140200 ID=Potri.002G140200.2.v4.1 annot-version=v4.1
MSYNQSRGGSDKSDLQYRKPGRSVSSSQQRTSSVSHGKGGGPPVPSPSSSSLSSNRSFNKKPSNLTQGGGQSSRVNLPSGVNSSDSGNNAASTIRNVQNG
VLTQHQSHGTSDASSVAKPTEASAAQRSTRDVPKAPTSQPAAISSESGAHMTPAKAPLDSSKAFAFQFGSISPGFMNGMQVPARTSSAPPNLDEQKRDQA
HHDTFRPAPSLPTPAPKQQLPRKEVSSSVQTSTGEVHLVPKASKETQLPPAPSVSQTQKPSVLPIPMNSLQMKYQQPPVSVQFRGPSPQIQSQGVPANSL
HVPIQLPMGNAPQVQQSVFIQGLQHHPMQPQGMMHQSQTMSFTNPMGPQIPQLGSLAYGMTSQYSAQQGGKFGSPHKTPVKITDPKTHEELRLDKRTDAY
PDAGSSGLRSHLNVPQTQPIPSFAPSRPINYYPSSYNASNLFFPAPSSLPLTGSQIAPNSQLPPRFNYPVSQPPQNAPYMNASALNSLPLSKSGTVSHGV
AEPQNSEHARDARNAISLTPSGAVQVTVKPAVGSHGEKVVEPSFPKISSIVEKGGFFKSSRSSGEASPSHSQRDSEASSESSLQRIKSGGESLVKPLPVA
AKQPAAVAVDGAASASLAQCEEAIPSVSNAEGRKKEALSGSNFIKEHQKKPGKKGNIQPQHQIGGQTTLSSHTLEHGVSSGTGVSETAENEKSPPSLANS
EVLTKSIKEPVSTIAAWNPDVSETKVDNAGDAFDSVSSQVPVAGIAHTTHISPHAKLDDSSQLEKLKCEIPATEDEIEKSLSECPKQDYNISSASINSKS
ADQVKQDKEVSDSVVTSVGNEVPASETAQEGLVEPVTCHTANDHISDNAGASTSRKFNSADDIKPLDASLSHSDNIGNKEASVTKSGISGHQGSPPVPDL
SEATAKHEGEGAENAGSGTVPLEVSGYKEKPSELTRSKSTANRMKKKKKEFLLKADLAGTTSDLYGAYKGPEEKKENVISSEVIESTSPNLKQAPADALQ
VQTVASEKSMQNKAEPDDWEDATDMSTLKLESLIDGELSLGGLGQHDTDGNANKLKKYSRDFLLKFSEQCTDLPGGFQIPSDIAGSLMGVGVSHLADRDP
CPSPARVMDRSNSGSRIDRRGSGIVDDGRWSKQPGPSGPGRDLHLDISYGANVGFRPVAGGNYGALRNPRAQSPVHYGGGILSGPMQSMGPQGGLQRGGL
DADRWQRAAIFVHKGSFSSPQTPLQTMHKAEKKYEVGKVTDEEAAKQRQLKGILNKLTPQNFEKLFEQVKAVNIDNVVTLNGVISQIFDKALMEPTFCEM
YANFCFHLAAELPELTEDNEKVTFKRILLNKCQEEFERGEREQEEANKADEEGEIKQSEEEREEKRIKARRRMLGNIRLIGELYKKRMLTERIMHECIKK
LLGQYQNPDEEDLEALCKLMSTIGEMIDHPKAKEHMDVYFDMMAKLSNNMKLSSRVRFMLKDSIDLRKNKWQQRRKVEGPKKIEEVHRDAAQERQLQTSR
LARNPGINPSPRRGPMDFGPRGSTMLPSLNAQMGGFRGFPTQVRGHGTQDVRFEEKQSYEARTMSVPLPQRPLGDDSITLGPQGGLARGMSIRGQPASMG
TLVADISPSPGDPRRMAAGLNGSSAISGRSNYSPREDIIPRYTPDRFAVPPACDQMNGQERNMNYVNRDLRNLDHGFDRPLGSSPPTRAQGPPFSQTTPT
GKLWPEERLRDMSTAAIKEFYSARDEKEVSLCIKELNSPSFHPSMISIWVTDSFERKDLERDLLAKLLVSLARSQNGILDSNQLIKGFESILTTLEDAVN
DAPKAPEFLGRIIGRVVVENVVPLSEIGPLLHEGGEEPGSLLKLGLAGDVLGSILEMIKVEKGEAVLNEIRGASNLRLEDFRPPDPNRSRILEKFI
|
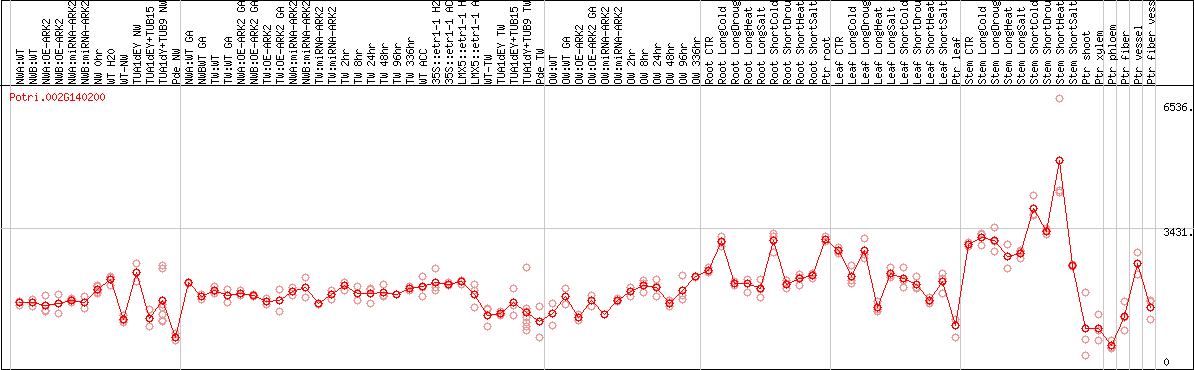
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G140200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
