Pt-MSJ1.3 (Potri.002G141100) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-MSJ1.3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G141100.6 pacid=42780008 polypeptide=Potri.002G141100.6.p locus=Potri.002G141100 ID=Potri.002G141100.6.v4.1 annot-version=v4.1
ATGTTTGGGCGTGCTCCAAGGAGGAGTGACAACACCAAGTATTATGAGGTTCTGGGTGTGTCAAAGGGTGCAAGTCAAGATGAACTGAAGAAGGCTTATA
GGAAAGCTGCCATAAAGAATCATCCTGATAAAGGTGGAGATCCTGAAAAGTTCAAGGAGTTGTCTCAAGCTTATGAAGTCCTTAGTGATCCAGATAAAAG
AGATATTTATGATCAATATGGGGAAGATGCACTTAAGGAGGGGATGGGACCTGGTGGTGGTGGTGGTGGTCATAATCCATATGATATATTTGAATCATTT
TTTGGTGGAGGTGGTTTTGGTGGTGGTGGCAGCTCAAGAGGAAGAAGGCAGAAGCAAGGTGAAGATGTAGTGCACCCTCTGAAGGTTTCCTTGGAGGATT
TGTACAATGGAACTTCCAAGAAACTCTCTCTTTCTAGAAACATTTTGTGTGCCAAATGTAAAGGGAAAGGCTCAAAGAGTGGAGCCTCTGGGACATGTCG
TGGCTGCCAAGGTACTGGAATGAAAGTTTCAATCCGACAAATTGGATTGGGCATGGTCCAACAAATGCAACATGTGTGTCCTGAATGCAGAGGCTCAGGT
GAGCTAATTAGTGAGAAGGATAAATGCCCTCATTGCAGAGGGAATAAGGTAACGCAGGAAAAGAGGGTGCTGGAAGTGCATGTTGAAAGGGGAATGCGAC
ATGGCCAGAAGATAGTTTTTGAAGGTCAAGCTGATGAAGCTCCTGACACAATTACAGGGGATATTGTTTTTGTATTGCAACTGAAAGAGCACTCCAAGTT
TGAACGGAAAATGGATGATCTCTTTGTGGAACACTCTGTCAGTTTAACAGAGGCTCTTTGCGGGTATCAGTTTGCCCTTACCCATCTCGATGGTCGGCAG
CTTCTTATTAAATCAAATCCTGGGGAGATTGTAAAACCTGGTCAATACAAAGCAATTAATGATGAAGGAATGCCACATCATCATAGGCCCTTCATGAAGG
GCAAGCTCTATATCCATTTTAATGTGGAGTTCCCCGAGTCGGGGACTCTATCCCCTGAGCAGTGCCGTACTTTAGAGACTATATTACCCCCAAGGCAAAG
CAAAAACTTGTCAGAGATGGAGTTGGATAATTGCGAAGAGACAATTATGCATGATGTCAATATTGAGGAGGAGAAAAGGCGCAAACAGCAGCAGCGCCAG
CAGCAGGAGGCATATGATGAGGATGATGATGATGAAGAATCACCTATGCCCCGAGTGCAGTGTGCCCAGCAGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G141100.6 pacid=42780008 polypeptide=Potri.002G141100.6.p locus=Potri.002G141100 ID=Potri.002G141100.6.v4.1 annot-version=v4.1
MFGRAPRRSDNTKYYEVLGVSKGASQDELKKAYRKAAIKNHPDKGGDPEKFKELSQAYEVLSDPDKRDIYDQYGEDALKEGMGPGGGGGGHNPYDIFESF
FGGGGFGGGGSSRGRRQKQGEDVVHPLKVSLEDLYNGTSKKLSLSRNILCAKCKGKGSKSGASGTCRGCQGTGMKVSIRQIGLGMVQQMQHVCPECRGSG
ELISEKDKCPHCRGNKVTQEKRVLEVHVERGMRHGQKIVFEGQADEAPDTITGDIVFVLQLKEHSKFERKMDDLFVEHSVSLTEALCGYQFALTHLDGRQ
LLIKSNPGEIVKPGQYKAINDEGMPHHHRPFMKGKLYIHFNVEFPESGTLSPEQCRTLETILPPRQSKNLSEMELDNCEETIMHDVNIEEEKRRKQQQRQ
QQEAYDEDDDDEESPMPRVQCAQQ
|
DESeq2's median of ratios [POPLAR]
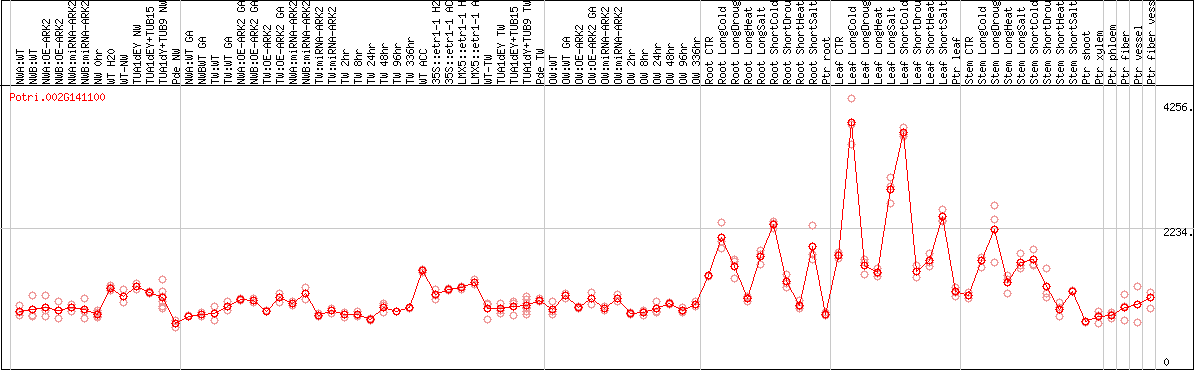
Coexpressed genes
Potri.002G141100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
