External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G00080 164 / 6e-51
UNE11
unfertilized embryo sac 11, Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT1G62770 138 / 3e-41
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT3G47380 136 / 2e-40
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT5G62350 134 / 3e-39
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT4G25260 131 / 2e-38
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT4G12390 128 / 5e-37
PME1
pectin methylesterase inhibitor 1 (.1)
AT5G62360 117 / 9e-33
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT5G20740 112 / 6e-31
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT4G25250 104 / 7e-28
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT1G14890 100 / 7e-26
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G067500
320 / 3e-112
AT4G00080 156 / 1e-47
unfertilized embryo sac 11, Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.012G127500
158 / 6e-49
AT5G62350 220 / 2e-73
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128700
154 / 2e-47
AT5G62350 221 / 7e-74
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.003G113700
147 / 1e-44
AT4G12390 178 / 1e-56
pectin methylesterase inhibitor 1 (.1)
Potri.015G128100
121 / 2e-34
AT5G62360 221 / 1e-73
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128200
119 / 1e-33
AT5G62360 172 / 1e-54
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128300
115 / 5e-32
AT5G62360 192 / 1e-62
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.001G119300
114 / 8e-32
AT1G62760 167 / 3e-51
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.012G127400
109 / 6e-30
AT4G25250 150 / 7e-46
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030292
187 / 5e-60
AT4G00080 158 / 1e-48
unfertilized embryo sac 11, Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10001988
182 / 3e-58
AT4G00080 154 / 5e-47
unfertilized embryo sac 11, Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031133
149 / 3e-45
AT5G62350 211 / 8e-70
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031713
140 / 7e-42
AT5G62350 202 / 4e-66
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10027198
133 / 5e-39
AT5G62350 191 / 3e-62
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10038914
132 / 5e-39
AT5G62350 192 / 1e-62
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10032230
114 / 2e-31
AT1G62770 166 / 9e-52
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10024593
111 / 1e-30
AT1G62770 162 / 2e-50
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031138
107 / 9e-29
AT5G62360 172 / 3e-54
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10038915
100 / 3e-26
AT5G62360 174 / 5e-55
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04043
PMEI
Plant invertase/pectin methylesterase inhibitor
Representative CDS sequence
>Potri.002G145800.1 pacid=42779268 polypeptide=Potri.002G145800.1.p locus=Potri.002G145800 ID=Potri.002G145800.1.v4.1 annot-version=v4.1
ATGAATCGTCTCAGCCTTTCCATACCGCTACTGCTGCTATCCATATTTTGCTTTTCAGGCACAGTGGAACCGGCAAGGTTTGCCAGACATTCCCGGCCTC
GGGCCTATATCGAGACTGCGTGCACGAAGACCCTTTATCCATCCCTATGCACCCAATACCTATCAGTCTTTGCAAACTCAACAATACAAACCCCTCAACA
ACTAGCCCAAGCTGCCTTATCCGTGAGCCTATACAAGGCCCTCCAAACAAGAACATTCATGTTGAAAGTGGTCAAGGAGCTCAAGGCAATGAAATCTAAG
GATTACCAGGCTGTGAAGGACTGCTTGGATCAAATTGGTGATAGCGTGGATCAACTTAGCCAATCAGTTAGAGAGCTCCATCGCTTGGAACATCCAGGTG
CTGCGGGTGGCGGTGACGTCTTTTGGCACGTAAGTAATTTCGAGACCTGGGTAAGCTCTGCCATGACTGATGCCAGTACTTGTGTGGATGAGTTACCGGG
GAAGGATATGAACAAGTTAAAGGCTGTAATCAAGGCCAAGGTTTTGAATGTGGCACAGACTGCTAGCAATGCACTAGCCTTGTTTCAGCGGTACGCTGCA
AAGCACAAGCCATAA
AA sequence
>Potri.002G145800.1 pacid=42779268 polypeptide=Potri.002G145800.1.p locus=Potri.002G145800 ID=Potri.002G145800.1.v4.1 annot-version=v4.1
MNRLSLSIPLLLLSIFCFSGTVEPARFARHSRPRAYIETACTKTLYPSLCTQYLSVFANSTIQTPQQLAQAALSVSLYKALQTRTFMLKVVKELKAMKSK
DYQAVKDCLDQIGDSVDQLSQSVRELHRLEHPGAAGGGDVFWHVSNFETWVSSAMTDASTCVDELPGKDMNKLKAVIKAKVLNVAQTASNALALFQRYAA
KHKP
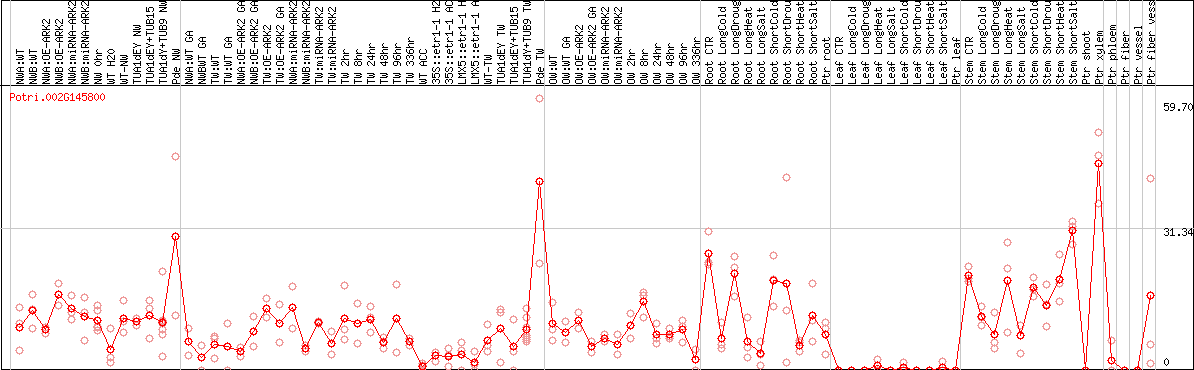
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G145800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.