Potri.002G149200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G149200.1 pacid=42778147 polypeptide=Potri.002G149200.1.p locus=Potri.002G149200 ID=Potri.002G149200.1.v4.1 annot-version=v4.1
ATGGCTTCCTCGGAAGCCGATTCCCGGTTAAGCCAAGTGGTATCCCCTGCTTTAGAAAAAATCGTCAAAAACGCATCGTGGCGTAAACACTCGAAATTAG
CACACGAATGCAAATCCGTCCTCGAGATTCTCACTTCTCGAAAACCCCAACAACAACATCCTCCGACTTCCCCGTCGGACGACTCCTCATCCGAATCCTC
TCTCCCTGGCCCACTCCACGACGGCGGGTCCATCGAGTACTCCCTCGCCGAATCTGAATCTATCCTCAGCCCTCTGATCAACGCGTGCAACACTCAATTC
CTCAAGATCGTCGATCCAGCCGTTGATTGCATCCAGAAATTGATCGCGCATGGGTACCTACGTGGCGAGGCGGACTCCACCGGAGGTACTGAAGCGAAAT
TGCTAGCTAAGTTGATTGAATCTGTGTGTAAATGTTATGATTTAGGGGATGATGGTGCTGAATTGCTGGTTTTGAAAACCCTATTATCAGCCGTTACGTC
GATTTCGTTGAGAATTCATGGAGATTGTTTGTTACAAATAGTGAGAACTTGTTATGACATTTATTTAGGGAGTAAAAATGTAATTAATCAAACGACGGCG
AAAGCGTCTTTGATTCAAATGTTAGTTATTGTGTTTAGGAGAATGGAGGCTGATTCTTCAACGGTTCCGGTTCAGCCGATTGTTGTGGCTGAATTAATGG
AACCAGTGGAGAAAACGGATGTCGATGGATCAATGGCGGTGTTTGTTCAAGGTTTTATAACGAAAATTATGCAGGATATAGATGGGGTTTTTAATCCGGG
GACACCCAGTAAGTCTTCGATGACAGTGGCACATGATGGGGCTTTCGAGACCACTACTGGTACGGTGGAGAGTACAAATCCGGCAGATTTGCTTGATTCG
ACGGATAAGGATATGTTAGATGCCAAATATTGGGAAATTAGTATGTATAAGACGGCTTTGGAAGGCAGGAAAGGGGAGTTGGCTGATGGGGAAGGGGAGA
GGGAGGATGATTTGGAAGTGCAGATTGGGAATAAGTTGAGGAGAGATGCATTTTTGGTGTTTCGAGCACTTTGTAAGTTATCTATGAAAACTCCGCCCAA
AGAAGCATTGGCGGATCCACAGTTGATGAGAGGGAAGATTGTTGCATTGGAGTTATTGAAGATTTTGTTGGAGAATGCTGGTGCTGTGTTTAGGACTAGT
GACAGGTTCTTAGGTGCCATCAAGCAATATCTATGTCTTTCACTGTTGAAGAACAGCTCGTCAAGTCTTATGATCATTTTCCAGCTTTCTTGCTCCATTT
TCATTAGTTTGGTTTCAAGGTTCAGAGCTGGACTGAAAGCAGAAATTGGAGTATTTTTTCCTATGATTGTTCTCAGGATTCTTGAAAATATTGTTCAACC
TAATTTTCAGCAGAAGATTATAGTACTTCGGTTTCTAGATAAGCTCTGTGTTGATTCCCAGATCTTGGTAGACATTTTTATTAATTATGACTGTGATATC
AACTCATCAAACATATTTGAGAGAATGGTCAATGGACTTCTTAAAACTGCTCAAGGTGCCCTGCCTGGTACTGCCACTACACTGGTGCCACCTCAGGAGG
TGACCATGAAACTTGAAGCTATGAAGTCCTTAGTGGCTATTCTGAAATCAATGGGAGACTGGATGAATAAACAATTGCGCATTCCCGATCCTCATTCTGC
CAAGAAATCTGATGCAGCCGAAAACAGTCCTGGACCTGGAAGTCTTCCCATGACAAATGGTAATGGGGATGAGCCTGTTGAAGGATCAGATTCTCATTCT
GAAACCTCAACTGAAGCATCGGATGTTTCAGCTATTGAGCAACGACGGGCTTACAAGCTGGAATTTCAGGAAGGTATATCTCTTTTCAATCGGAAGCCTA
AGAAAGGAATTGAATTTCTAATCAATGCAAATAAAGTGGGGAATTCAGCAGAAGAGATAGCAGCCTTTCTTAAAAACGCATCTGGCTTGAACAAGACTCT
GATTGGCGATTACCTTGGGGAAAGAGAAGATTTTTCACTGAAAGTAATGCATGCTTATGTAGATTCTTTTGACTTTCGCGGCCTGGAGTTCGATGAGGCT
ATCAGAGTCTTTCTACAGGGCTTTAGGTTACCAGGTGAGGCACAAAAGATTGATCGAATTATGGAGAAGTTTGCTGAACGTTACTGCAAATGTAACCCAA
AGGTTTTTAGCAGTGCTGACACAGCCTATGTCCTTGCTTACTCCGTGATACTGCTCAACACTGATGCTCATAATCCCATGGTGAAGAATAAGATGTCAGC
TGATGACTTCATAAGAAACAACCGTGGCATTGATGATGGGAAAGATCTACCTGAGGAGTACTTGAGATCTTTGTTTGAAAGAATATCAAAAAATGAGATC
AAAATGAAAGAATATGATTTAGCTCTCCAACAAAAGCAGTCTTTGAATTCAAACAGAGTTTTAGGCTTGGATAGTATCCTGAACATTGTGATCCGCAAGC
GGGGTGAAGAAAAGAATATGGAAACAAGTGATGATCTTATCAGGCACATGCAGGAACAATTTAAAGAAAAAGCTCGAAAATCTGAGTCAGTTTATTATGC
TGCAACAGATGTGGTGATTCTTAGATTCATGATTGAGGTATGCTGGGCTCCTATGCTGGCAGCCTTCAGTGTGCCTCTTGACCAAAGTGATGATGAAGTG
GTGATAGCTTTGTGTCTTGAAGGTATCCGCTATGCAATCCATGTTACTGCTGTGATGTCCATGAAGACTCATAGAGATGCTTTTGTGACTTCATTAGCAA
AGTTTACATCGCTCCACTCCCCTGCTGATATAAAGCAGAAAAATATTGATGCTATCAAGGCAATAGTTACAATTGCAGATGAAGATGGGAATTATTTGCA
AGAAGCTTGGGAACATATTTTGACATGTGTCTCACGCTTTGAACATTTGCATCTCCTGGGAGAGGGTGCTCCTCCAGATGCCACTTTTTTTGCTTTTCCC
CAAAATAACTCTGAAAAGTCAAAACAAAGCAAGTCAACCATCCTCCCTGTTCTGAAAAAGAAGGGACCCGGGAGGATGCAGCATGCAGCTGCTAGTGTGC
TGAGAGGTTCATATGATAGTGCTGGTATTGGTGGAAACGCTGCTGGGGCTGTCACAAGTGAGCAGATGAACAATTTAGTTTCCAACTTAAACAAGTTAGA
GCAAGTTGGCAGCTCTGAGATGAACCGCATCTTCACACGGAGCCAAAAGTTAAACAGTGAGGCAATAATTGACTTTGTGAAGGCTCTTTGCAAGGTTTCT
GTGGAGGAATTGAGATCAGCTTCTGATCCACGGGTCTTTAGCCTTACAAAGATTGTTGAGATTGCGCACTTTAACATGAATCGCATCAGGCTTGTATGGT
CAAGCATCTGGCATGTGCTTTCTGATTTCTTTGTAACCATTGGCTGTTCTGAAAACCTTTCCATTGCAATTTTTGCAATGGACTCTTTACGCCAATTATC
AATGAAATTCTTAGACCGAGAAGAGTTGGCAAACTATAATTTTCAGAATGAGTTTATGAAGCCGTTTGTTATTGTTATGCGGAAGAGTAATGCTGTTGAA
ATCAGAGAATTGATTATCAGATGTGTCTCGCAAATGGTCCTGTCTCGTGTCAATAATGTTAAATCTGGATGGAAGAGCATGTTCATGGTTTTCACAGCAG
CAGCTTATGATGACCACAAAAACATTGTACTCTTGGCCTTTGAAATAATTGAAAAGATTATTAGAGATTACTTCCCATACATTACGGAGACTGAAACAAC
CACCTTTACTGATTGTGTGAATTGCCTGATAGCATTCACCAATAGTAGATTCAACAAAGACATTAGCCTTAATGCAATTGCTTTTCTTCAATTCTGTGCC
ACAAAACTTGCTGAAGGAGATCTTGGATCTTCATCAAGGAACAAAGACAAGGAAGTCTCTGTAAAAATTTCATCCCCTTCACCTCGGACTGGAAAAGATG
GAAAACAAGAGAATGGAGAAATCAAAGACAAGGAAGATCATCTTTATTTCTGGTTCCCTCTGTTGGCTGGTTTATCAGAACTTAGCTTTGACCCTAGGCC
TGAAGTCAGGAAGAGTGCCTTGCAAGTTCTGTTTGAAACTTTACGCAATCATGGTCATCTTTTTTCTCTACCTTTATGGGAAAGAGTCTTTGAGTCGGTC
TTATTTCCTATATTTGACTATGTGCGGCATGCTATTGATCCTCCTGGAGGCAACTCCCCTGAGCAGGGGATTGATGGTGATATGGGTGAACTTGATCAAG
ATGCATGGCTCTATGGGACATGTACCTTGGCTCTCCAACTGGTTGTAGATCTTTTTGTGAAATTCTACAATACTGTCAATCCGCTTTTAAGAAAGGTGCT
ATCGCTTCTGGTTAGCTTCATTAGGCGTCCTCACCAAAGTCTTGCTGGTATTGGTATTGCGGCATTTGTGCGTTTGATGAGCAATGCTGGAGATATGTTT
TCAGAGGAGAAGTGGCTGGAAGTGGTTTTGTCATTGAAAGATGCTGCCAATGCAACACTCCCTGATTTCTCTTACATTGTAAGTGGAGAATCATCGGTAA
TAGCCGATGAACAAAATAATGGAGAGACTGCTGGATCTGACATGCCCGAAGATGAATCAGAGGGACTGGTGACACACCATCTTTATGCTTCTATATCTGA
TGCAAAGTGTCGAGCTGCTGTTCAACTTCTGTTGATTCAGGCGGTGATGGAGATCTATAGCATGTACCGGTCTCAACTTTCAGCAAAATGTGCCTTGGTG
CTCTTTGATGCTTTGCATGAGGTGGCATCCCATGCTCACAGTATCAACACAAATACAACATTACGTTCAAAACTACAAGAGTTTGGCTCTATGACCCAAA
TGCAAGACCCTCCACTATTACGCCTGGAGAATGAGTCCTATCAAATTTGCCTCACATTCTTACAAAATCTTATGATGGACAGACCTCCACCTTTTGATGA
AGCTGAAGTGGAATCCTGCCTTGTCAACCTCTGTGAAGAGGTTTTACAGTTTTACGTTGTAACTGCCTGTTCTGGACAGGCATCAGAAACTTCTACCAGT
GGACAATGCCTGTGGCTGATTCCTTTAGGTTCTGGAAAACGAAGGGAATTGGCTGCACGTGCACCTCTTATTGTTGCAACTCTCCAGGCCATCTGTAGTT
TGGGAGACTCTTCGTTTGAGAAGAAATTGCCTCATTTCTTCCCTCTTCTTTCGAGTTTGATAAGTTGTGAGCATGGGTCAAATGAGGTCCAGGTAGCTCT
CAGTGACATGCTCAGCTCATCAGTGGGTCCTGTTTTGCTTCGGTCTTGTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G149200.1 pacid=42778147 polypeptide=Potri.002G149200.1.p locus=Potri.002G149200 ID=Potri.002G149200.1.v4.1 annot-version=v4.1
MASSEADSRLSQVVSPALEKIVKNASWRKHSKLAHECKSVLEILTSRKPQQQHPPTSPSDDSSSESSLPGPLHDGGSIEYSLAESESILSPLINACNTQF
LKIVDPAVDCIQKLIAHGYLRGEADSTGGTEAKLLAKLIESVCKCYDLGDDGAELLVLKTLLSAVTSISLRIHGDCLLQIVRTCYDIYLGSKNVINQTTA
KASLIQMLVIVFRRMEADSSTVPVQPIVVAELMEPVEKTDVDGSMAVFVQGFITKIMQDIDGVFNPGTPSKSSMTVAHDGAFETTTGTVESTNPADLLDS
TDKDMLDAKYWEISMYKTALEGRKGELADGEGEREDDLEVQIGNKLRRDAFLVFRALCKLSMKTPPKEALADPQLMRGKIVALELLKILLENAGAVFRTS
DRFLGAIKQYLCLSLLKNSSSSLMIIFQLSCSIFISLVSRFRAGLKAEIGVFFPMIVLRILENIVQPNFQQKIIVLRFLDKLCVDSQILVDIFINYDCDI
NSSNIFERMVNGLLKTAQGALPGTATTLVPPQEVTMKLEAMKSLVAILKSMGDWMNKQLRIPDPHSAKKSDAAENSPGPGSLPMTNGNGDEPVEGSDSHS
ETSTEASDVSAIEQRRAYKLEFQEGISLFNRKPKKGIEFLINANKVGNSAEEIAAFLKNASGLNKTLIGDYLGEREDFSLKVMHAYVDSFDFRGLEFDEA
IRVFLQGFRLPGEAQKIDRIMEKFAERYCKCNPKVFSSADTAYVLAYSVILLNTDAHNPMVKNKMSADDFIRNNRGIDDGKDLPEEYLRSLFERISKNEI
KMKEYDLALQQKQSLNSNRVLGLDSILNIVIRKRGEEKNMETSDDLIRHMQEQFKEKARKSESVYYAATDVVILRFMIEVCWAPMLAAFSVPLDQSDDEV
VIALCLEGIRYAIHVTAVMSMKTHRDAFVTSLAKFTSLHSPADIKQKNIDAIKAIVTIADEDGNYLQEAWEHILTCVSRFEHLHLLGEGAPPDATFFAFP
QNNSEKSKQSKSTILPVLKKKGPGRMQHAAASVLRGSYDSAGIGGNAAGAVTSEQMNNLVSNLNKLEQVGSSEMNRIFTRSQKLNSEAIIDFVKALCKVS
VEELRSASDPRVFSLTKIVEIAHFNMNRIRLVWSSIWHVLSDFFVTIGCSENLSIAIFAMDSLRQLSMKFLDREELANYNFQNEFMKPFVIVMRKSNAVE
IRELIIRCVSQMVLSRVNNVKSGWKSMFMVFTAAAYDDHKNIVLLAFEIIEKIIRDYFPYITETETTTFTDCVNCLIAFTNSRFNKDISLNAIAFLQFCA
TKLAEGDLGSSSRNKDKEVSVKISSPSPRTGKDGKQENGEIKDKEDHLYFWFPLLAGLSELSFDPRPEVRKSALQVLFETLRNHGHLFSLPLWERVFESV
LFPIFDYVRHAIDPPGGNSPEQGIDGDMGELDQDAWLYGTCTLALQLVVDLFVKFYNTVNPLLRKVLSLLVSFIRRPHQSLAGIGIAAFVRLMSNAGDMF
SEEKWLEVVLSLKDAANATLPDFSYIVSGESSVIADEQNNGETAGSDMPEDESEGLVTHHLYASISDAKCRAAVQLLLIQAVMEIYSMYRSQLSAKCALV
LFDALHEVASHAHSINTNTTLRSKLQEFGSMTQMQDPPLLRLENESYQICLTFLQNLMMDRPPPFDEAEVESCLVNLCEEVLQFYVVTACSGQASETSTS
GQCLWLIPLGSGKRRELAARAPLIVATLQAICSLGDSSFEKKLPHFFPLLSSLISCEHGSNEVQVALSDMLSSSVGPVLLRSC
|
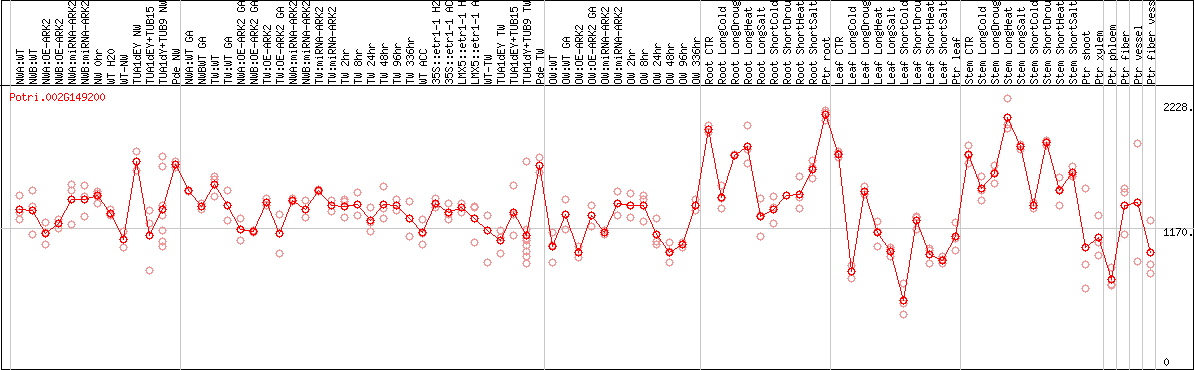
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G149200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
