External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G01850 105 / 5e-27
ATXTH27, EXGT-A3
XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE 27, endoxyloglucan transferase A3 (.1)
AT1G14720 103 / 4e-26
ATXTH28, EXGT-A2, XTR2
xyloglucan endotransglycosylase related 2, ENDOXYLOGLUCAN TRANSFERASE A2, xyloglucan endotransglucosylase/hydrolase 28 (.1)
AT4G18990 102 / 8e-26
XTH29, XTR13
xyloglucan endotransglucosylase/hydrolase 29 (.1)
AT1G10550 98 / 2e-24
XTH33, XET
xyloglucan:xyloglucosyl transferase 33 (.1)
AT2G36870 97 / 5e-24
XTH32
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
AT2G18800 96 / 1e-23
ATXTH21, XTH21, XTR17
xyloglucan endotransglucosylase/hydrolase 21 (.1)
AT4G30270 95 / 1e-23
XTH24, SEN4, BRU1, MERI-5, MERI5B
SENESCENCE 4, meristem-5, MERISTEM 5, xyloglucan endotransglucosylase/hydrolase 24 (.1)
AT1G32170 96 / 2e-23
XTH30, XTR4
xyloglucan endotransglycosylase 4, xyloglucan endotransglucosylase/hydrolase 30 (.1)
AT5G48070 95 / 2e-23
XTH20, ATXTH20
xyloglucan endotransglucosylase/hydrolase 20 (.1)
AT4G03210 95 / 3e-23
XTH9, EXGT-A6, XTR16
xyloglucan endotransglucosylase/hydrolase 9 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G138400
104 / 1e-26
AT1G14720 467 / 2e-166
xyloglucan endotransglycosylase related 2, ENDOXYLOGLUCAN TRANSFERASE A2, xyloglucan endotransglucosylase/hydrolase 28 (.1)
Potri.010G102300
103 / 3e-26
AT1G14720 470 / 7e-168
xyloglucan endotransglycosylase related 2, ENDOXYLOGLUCAN TRANSFERASE A2, xyloglucan endotransglucosylase/hydrolase 28 (.1)
Potri.006G071200
98 / 1e-24
AT4G25810 379 / 3e-133
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.013G005700
97 / 1e-23
AT4G25810 431 / 5e-153
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.003G097300
97 / 1e-23
AT1G32170 436 / 8e-154
xyloglucan endotransglycosylase 4, xyloglucan endotransglucosylase/hydrolase 30 (.1)
Potri.018G094900
95 / 2e-23
AT4G25810 410 / 8e-146
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.018G095200
94 / 3e-23
AT4G25810 414 / 3e-147
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.018G095100
94 / 4e-23
AT4G25810 420 / 1e-149
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.016G098600
94 / 5e-23
AT2G36870 477 / 1e-171
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041373
150 / 1e-46
ND 39 / 3e-04
Lus10013822
103 / 1e-26
AT2G36870 444 / 1e-158
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Lus10026535
103 / 1e-26
AT2G36870 444 / 9e-159
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Lus10020772
102 / 5e-26
AT3G23730 393 / 7e-139
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10007349
101 / 9e-26
AT3G23730 394 / 4e-139
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10013240
102 / 1e-25
AT2G01850 456 / 2e-162
XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE 27, endoxyloglucan transferase A3 (.1)
Lus10030760
101 / 2e-25
AT1G14720 456 / 3e-162
xyloglucan endotransglycosylase related 2, ENDOXYLOGLUCAN TRANSFERASE A2, xyloglucan endotransglucosylase/hydrolase 28 (.1)
Lus10030484
100 / 3e-25
AT4G25810 398 / 9e-141
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Lus10023009
100 / 4e-25
AT2G36870 466 / 3e-167
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Lus10001396
100 / 4e-25
AT2G36870 460 / 8e-165
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0004
Concanavalin
PF00722
Glyco_hydro_16
Glycosyl hydrolases family 16
Representative CDS sequence
>Potri.002G153200.1 pacid=42778835 polypeptide=Potri.002G153200.1.p locus=Potri.002G153200 ID=Potri.002G153200.1.v4.1 annot-version=v4.1
ATGGCGGATCCAGCAATCCACCACGAGACCCAACCCATTAACCAAATCGCCATAGACTACACCCCAGAGGCATGCACTCACTGCCCGGAGTCCAATTCCA
TCACTCTCACATACGACCACCGTGGGGGCGCACGGTGGCGCAGTACCACCCGTTTTCTTTACGGCACTTTCAGTTCTCTCATCCAATGTCCCAAAGGTAA
CACGAGTGGCCTTAACTTCAACATCTATCTTTCTTCTCTTGAAGGTGACAAATCCCAAGATGAGATCGACTTTGAGTTTCTTGGTAAAGACAAGACCATC
GTACAAACTAATTACTACGCTTCAGGGACTGGGAACAGAGAAGAAATTCATGATCTGGGGTTTGATTGTTCTGATGCGTTTCATGAGTATGTAATAAAGT
GGTGTCCAAATTTCATAGAGTGGTTGATTGATGGAAAAGTGGTGAGGAAAGTGGAGAAAAGGGAAGGGGAGGGATTCCCTGAGAAACCCATGTTTTTGTA
TGCTTCAATTTGGGATGCAAGCTACATTGGTGACGCAACTTGGACTGGACCGTATATGGGGTGTGATGCACCTTATCTTTGTCTCTACAAGGACATTTGT
GTGCCTGTTGGGACTGCGGTGGAGTGTTCTTGTGATTCTTGA
AA sequence
>Potri.002G153200.1 pacid=42778835 polypeptide=Potri.002G153200.1.p locus=Potri.002G153200 ID=Potri.002G153200.1.v4.1 annot-version=v4.1
MADPAIHHETQPINQIAIDYTPEACTHCPESNSITLTYDHRGGARWRSTTRFLYGTFSSLIQCPKGNTSGLNFNIYLSSLEGDKSQDEIDFEFLGKDKTI
VQTNYYASGTGNREEIHDLGFDCSDAFHEYVIKWCPNFIEWLIDGKVVRKVEKREGEGFPEKPMFLYASIWDASYIGDATWTGPYMGCDAPYLCLYKDIC
VPVGTAVECSCDS
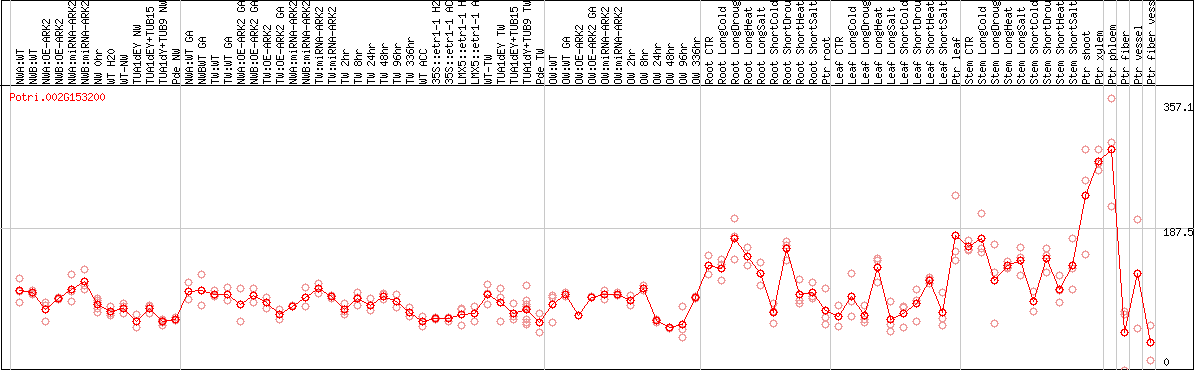
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G153200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.