External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G09970 466 / 1e-166
Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
AT3G09960 443 / 2e-157
Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
AT2G42500 46 / 7e-06
PP2A-3, PP2A-4
protein phosphatase 2A-3 (.1.2.3)
AT3G19980 45 / 4e-05
STPP, ATFYPP3, EMB2736
SERINE/THREONINE PROTEIN PHOSPHATASE, EMBRYO DEFECTIVE 2736, "flower-specific, phytochrome-associated protein phosphatase 3", flower-specific, phytochrome-associated protein phosphatase 3 (.1)
AT1G64040 44 / 0.0001
TOPP3
type one serine/threonine protein phosphatase 3 (.1)
AT1G03445 44 / 0.0001
BSU1
BRI1 SUPPRESSOR 1, Serine/threonine protein phosphatase family protein (.1)
AT3G58500 43 / 0.0002
PP2A-4, EP7, PP2A-3
protein phosphatase 2A-4 (.1)
AT3G46820 43 / 0.0002
TOPP5
type one serine/threonine protein phosphatase 5 (.1)
AT1G50370 43 / 0.0002
AtFYPP1
flower- specific, phytochrome-associated protein phosphatase 1, Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
AT2G39840 42 / 0.0003
TOPP4
type one serine/threonine protein phosphatase 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G156500
679 / 0
AT3G09970 478 / 4e-171
Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
Potri.008G003200
45 / 2e-05
AT1G50370 624 / 0.0
flower- specific, phytochrome-associated protein phosphatase 1, Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
Potri.001G215700
46 / 3e-05
AT2G27210 1626 / 0.0
BRI1 suppressor 1 (BSU1)-like 3 (.1), BRI1 suppressor 1 (BSU1)-like 3 (.2)
Potri.010G254500
45 / 3e-05
AT1G50370 617 / 0.0
flower- specific, phytochrome-associated protein phosphatase 1, Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
Potri.001G098600
45 / 3e-05
AT4G11240 582 / 0.0
Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
Potri.009G016900
46 / 4e-05
AT2G27210 1588 / 0.0
BRI1 suppressor 1 (BSU1)-like 3 (.1), BRI1 suppressor 1 (BSU1)-like 3 (.2)
Potri.001G245700
44 / 0.0001
AT2G29400 556 / 0.0
type one protein phosphatase 1 (.1)
Potri.013G045900
43 / 0.0002
AT2G39840 564 / 0.0
type one serine/threonine protein phosphatase 4 (.1)
Potri.001G196600
43 / 0.0003
AT5G10900 372 / 2e-119
Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014383
479 / 9e-172
AT3G09970 474 / 2e-170
Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
Lus10017361
46 / 2e-05
AT1G50370 617 / 0.0
flower- specific, phytochrome-associated protein phosphatase 1, Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
Lus10010156
46 / 2e-05
AT1G50370 617 / 0.0
flower- specific, phytochrome-associated protein phosphatase 1, Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
Lus10021872
46 / 2e-05
AT2G39840 523 / 0.0
type one serine/threonine protein phosphatase 4 (.1)
Lus10008369
46 / 4e-05
AT1G08420 1747 / 0.0
BRI1 suppressor 1 (BSU1)-like 2 (.1), BRI1 suppressor 1 (BSU1)-like 2 (.2)
Lus10004100
46 / 4e-05
AT1G08420 1686 / 0.0
BRI1 suppressor 1 (BSU1)-like 2 (.1), BRI1 suppressor 1 (BSU1)-like 2 (.2)
Lus10013357
46 / 4e-05
AT1G08420 1713 / 0.0
BRI1 suppressor 1 (BSU1)-like 2 (.1), BRI1 suppressor 1 (BSU1)-like 2 (.2)
Lus10015192
45 / 4e-05
AT3G05580 412 / 1e-144
type one protein phosphatase 9, Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
Lus10013456
45 / 4e-05
AT3G19980 623 / 0.0
SERINE/THREONINE PROTEIN PHOSPHATASE, EMBRYO DEFECTIVE 2736, "flower-specific, phytochrome-associated protein phosphatase 3", flower-specific, phytochrome-associated protein phosphatase 3 (.1)
Lus10011827
45 / 4e-05
AT2G39840 513 / 0.0
type one serine/threonine protein phosphatase 4 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0163
Calcineurin
PF00149
Metallophos
Calcineurin-like phosphoesterase
Representative CDS sequence
>Potri.002G156200.1 pacid=42777171 polypeptide=Potri.002G156200.1.p locus=Potri.002G156200 ID=Potri.002G156200.1.v4.1 annot-version=v4.1
ATGTACCCAAAAGCATTCTCTTTATCAATCGAAGGACAGAACAGAACCACCACGGAACCCAAAATGACAGAGGCAGAGCCAAGCAACAACAGCACGGTCG
CAAAACCCAGATTAGTAATCTGCATAGGCGACATCCATGGCCACATCACAAAGCTTCAAAATCTTTGGTCAAATCTCGAAACCCAATTCGACCCCCAACA
CTTTAACGCTGCAACAATTATCTTCTTAGGCGATTACTGTGACAGAGGCCCAGATACCAAGAAAGTCCTAGATTTCTTGATAGACTTACCTTCAAGGTAT
CCAAATCAAAAGCATGTGTTTCTTTCGGGAAATCATGACTTTGCTTTTGCTGCTTTTGTTGGTGTCCTGCCTGAACCGCAAAATGGGGTGTCATTTAAGG
AAGGATGGGAGGAGTATGAAGAGAGTGAGGACAGGGAAGGGTGGTATAAAGGAGACGGGTATGAGAATATGCATTTGGAAGGGAGAAGGTGGGCTGGTCA
TATTAAGGTGGGATTCAATACCATTAAAGGGACTGAATGTAAGGGCTCTATTTATGATGCTGGTCCCACTTTCACTTCTTATGGTGTCCCTCATGGATCT
TCTGACTTATTGAAGGCAGTTCCTGATGATCACAAGAAGTTTCTAGCTGATACGGTTTGGGTCCATGAAGAGGATGATGTGTGCATAGAGGATGAAGAAG
GAATTAGACACTGTAAATTGATTGCCGCGCATGCTGGTTTAGAGGAAGGGAAAAATGTAGGCGAACAACTGAGATTTTTGAAAGCTAAGGAGACTCATGT
CCCCAAGATAGAGGCTCTAAGTGGGAGGAAAACAGTTTGGGATATACCAAAGGAATTAACTGAGAAACCGACCATTGTGGTTAGCGGCCACCACGGGAAA
CTTCATATTGAAGGTCTAAGATTGATTATTGATGAAAGTGGTGGATTTGAGAATAAACCATTGGCTGCAATTGCGCTTCCCTCGATGAAGCTTGTTCGTG
ATACTGATAATTTGACAAAATAG
AA sequence
>Potri.002G156200.1 pacid=42777171 polypeptide=Potri.002G156200.1.p locus=Potri.002G156200 ID=Potri.002G156200.1.v4.1 annot-version=v4.1
MYPKAFSLSIEGQNRTTTEPKMTEAEPSNNSTVAKPRLVICIGDIHGHITKLQNLWSNLETQFDPQHFNAATIIFLGDYCDRGPDTKKVLDFLIDLPSRY
PNQKHVFLSGNHDFAFAAFVGVLPEPQNGVSFKEGWEEYEESEDREGWYKGDGYENMHLEGRRWAGHIKVGFNTIKGTECKGSIYDAGPTFTSYGVPHGS
SDLLKAVPDDHKKFLADTVWVHEEDDVCIEDEEGIRHCKLIAAHAGLEEGKNVGEQLRFLKAKETHVPKIEALSGRKTVWDIPKELTEKPTIVVSGHHGK
LHIEGLRLIIDESGGFENKPLAAIALPSMKLVRDTDNLTK
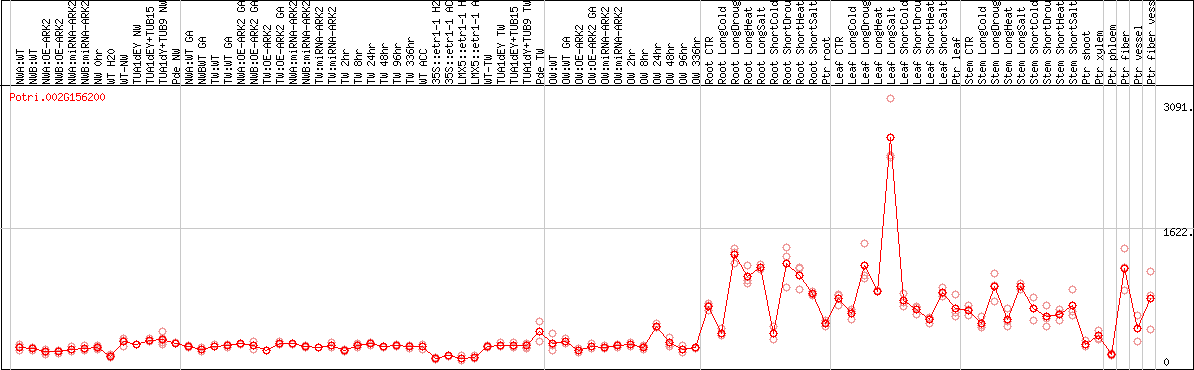
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G156200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.