External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G01800 207 / 4e-67
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G24190 207 / 1e-66
SDR2
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT3G61220 203 / 2e-65
SDR1
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT5G51030 110 / 5e-29
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G59710 105 / 4e-27
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G61830 90 / 2e-21
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G156800
240 / 1e-79
AT3G61220 363 / 1e-126
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G157000
238 / 5e-79
AT3G61220 406 / 9e-144
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G156750
233 / 7e-79
AT1G01800 165 / 1e-51
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G156300
236 / 4e-78
AT3G61220 367 / 6e-128
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G156600
233 / 8e-77
AT3G61220 348 / 7e-121
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.014G080100
229 / 2e-75
AT3G61220 350 / 3e-121
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.014G080000
219 / 2e-71
AT3G61220 332 / 2e-114
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.004G235500
175 / 5e-54
AT3G61220 313 / 7e-107
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.004G235600
156 / 6e-47
AT3G61220 285 / 1e-95
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040933
207 / 7e-67
AT3G61220 342 / 1e-118
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10040932
208 / 9e-67
AT3G61220 327 / 3e-112
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10014694
202 / 1e-64
AT3G61220 369 / 4e-129
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10014690
201 / 2e-64
AT3G61220 335 / 2e-115
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10014688
200 / 5e-64
AT3G61220 348 / 8e-121
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10014689
202 / 7e-62
AT3G61220 336 / 2e-112
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10006894
191 / 3e-60
AT3G61220 325 / 1e-111
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10014693
186 / 2e-58
AT3G61220 347 / 3e-120
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10006893
184 / 1e-57
AT3G61220 313 / 4e-107
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10008413
162 / 4e-49
AT2G24190 273 / 3e-91
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
PFAM info
Representative CDS sequence
>Potri.002G156866.1 pacid=42777786 polypeptide=Potri.002G156866.1.p locus=Potri.002G156866 ID=Potri.002G156866.1.v4.1 annot-version=v4.1
ATGCAAGTGAACAATGCAGCGATCCGTGGAACCACAATAGACTCGGACGCTTCAGCAGCTTCAAAAATTACTGGTACAGAAGACGATTTGCAGAATGTTT
GGAAGAAGGTATCGATTCAAAATTATGAGTTAGCAGAAGAGTGCCTGAACACAAACTGCTGCGGTGCTAAAAGAACAGCTGAAGCTCTTATTCCGCTCCT
CCAGCTATCCGACTCTCCAAGGATTGTCAATGTTTCTTCCTTCTTGTCAATGCTGAAGAATATACCAAATGAATGGGCTAAAGGAGTGTTCAGTGATGTT
GATACCTTTACAGAGGAGAGAATAGATGAACTATTAAGTGTATTTCTAAAAGATTTCAAGGAGGATTCTTTAGAAACCAAGGGCTGGCCAGCCTCACTGT
CAGCCTATGCACTCTCAAAAGCAGCCATGAATGCTCACACAAGGATTGTAGCCAAGAAGCACCCCAATTTCTGCATTAATTGCATCTGTCCAGGGTTTGT
CAAAACAGATATGAGCAATAACACCGGCACCTTATCGGTTGATGAAGCTGCAGTGTATCCTGTGAAATTGGCGCTGCTTCCTGATGGAGGACCTTCTGGC
CTCTTTTTCATTCTGGATAAATTATCTTGTTTCTGA
AA sequence
>Potri.002G156866.1 pacid=42777786 polypeptide=Potri.002G156866.1.p locus=Potri.002G156866 ID=Potri.002G156866.1.v4.1 annot-version=v4.1
MQVNNAAIRGTTIDSDASAASKITGTEDDLQNVWKKVSIQNYELAEECLNTNCCGAKRTAEALIPLLQLSDSPRIVNVSSFLSMLKNIPNEWAKGVFSDV
DTFTEERIDELLSVFLKDFKEDSLETKGWPASLSAYALSKAAMNAHTRIVAKKHPNFCINCICPGFVKTDMSNNTGTLSVDEAAVYPVKLALLPDGGPSG
LFFILDKLSCF
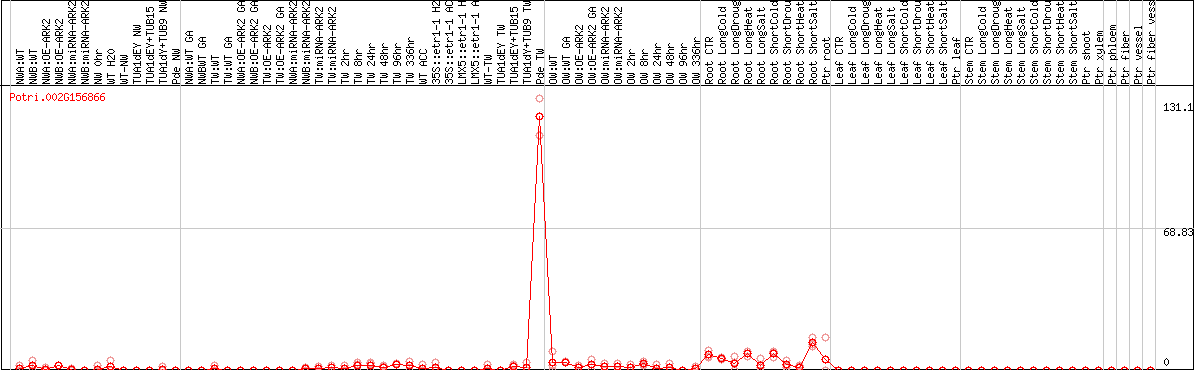
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G156866 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.