Potri.002G157200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G157200.1 pacid=42779795 polypeptide=Potri.002G157200.1.p locus=Potri.002G157200 ID=Potri.002G157200.1.v4.1 annot-version=v4.1
ATGGAGTTTGCGTGCAATATTCAGCAGACGAATGCATTTTATAGCAGACAAGGTACTAGTTGTAGGGTATCAAATAGGTTATATTCGAGATTTAGACATA
AAAGTTATGGGTATAATGCTGGTGATTTGAAGATTGTTTCGAGAGAAAGACCTTCAAAGACATTAAAGAAAAGTGTTTTATATGGTAGTGGAAGTGGAAT
GCGTAGTCATTTGCTTGTTGGTGGTTATGCAAGTAATCCGTTGTTTTGCAATTTTATTGATGGGTTTGAGGGATTGAGAAGTGTTAAGTTGCGGTGTCAA
GGTAATGATTCGTTAACATATATTGATGGGAATGGTAGAAATGTAGAAATTGGTAAGGGTAATGATAAGAACTTGAGAGCAGGGTCTAATGGTGGCTTGG
GAGAAGAGGATGGAAGAGGAGAAAAGGTAATGGAGACTGAAATGGCGGCAGAGGCACTTAGTTTGGATGAATTGAGGGAATTGTTACAGAAAGCAATGAG
GGAATTGGAGGTTGCACGGCTTAATAGTACCATGTTCGAGGAAAAGGCACAGAGTATATCAGAAACTGCAATAGCTTTGCAAGATGAGGCATCGAGTGCT
TGGAATGATGTTAACTCAACACTTGATATGATTCAGGACATTGTGAACAAGGAGGGTGTTGCTAAAGAAGCTTTTCAAAAGGCAACTATGGCGCTTTCTT
TGGCTGAGGCAAGGCTTAAGGTGGCAGTGGAGTCGATAAAATCTACGAAAGAAGGGGTTGATTCATTGGAGGGTTCTGGAGAGAGTGATGTGCAAAATGA
TAATAAGGAAGATTACGAGACAATTTTGGCTGCTCAGAATGATATTAGGGAATGCCAGGCAAATTTGGCAAATTGTGAAGCAGAGCTGAGGCGCTTGCAA
AGTATAAAGGAAGAATTACAAAAGGAGGTAGACGTGTTGAATGAGAAAGCAGAGAAAGCACAAATGAATGCTTTGAAAGCAGAGGAGGATGTTGCAAACA
TTATGCTTCTAGCAGAGCAAGCTGTTGCCTTTGAACTTGAGGCTACCCAACGAGTGAGTGATGCAGAAATTGCGTTACAAAAGGCAGAGAAATCTCTCTC
TAGTTCTCATGTTGATATCCAGCAAACAGCTCGGGGATATGTGTCTGGTGATGAGGCAGTAGTCGAGGAGGAGAAGATGCGCGGAGGCAGTCTTTCCGAT
GTTGAGAAAGAAACAGATATGACTGTAAATGGTGATGTTTTAGTTGGCGAGCCTTCAATAGATAGACTATCTGATAAAACCAGCCCGAGTTCTGAAGAGT
TATATCTTTCTGATTATTCTAGTGACCACGAGAATGGAAAATCGAGTTTGCACTCTATTAAAGATACTGAAGCAGAAGCTGAAAAGTCAAAAGGTGGCAT
TCAGACAAAAAAGCAGGAATTACAGAAGGACCTGACCAGGGAGAGTTCATCTTCACCTCTCAGTGCTCCCAAGGCGTTGTTGAAGAAATCCTCTCGTTTC
TTTTCTGCATCCTTCTTCTCTTTCAGTGGAGATGAGACGGAGTTGACAGCAGCATCAGTTTTTCAGGGCTTAATGGAGCCTGCAAGGAAGCAACTGCCCA
AATTTCTTCTTGGCCTATTGTTGTTTGGAGCAGGATTTGCCTTCTACTCTAATCGAGTGGAGAAGAGTACCCAGATGCTTCAAAAACCAGAAGTGGTCAC
CACAAGCATTGAAGAAGTTTCATCAAATGCAAAGCCTCTGATTCAACATATTCAGAAACTCCCAAAGAGAGTCAAAAAACTAATTGCAATGCTTCCTCAT
CAAGAGATGAATGAGGAGGAAGCATCTCTCTTTGATGTTTTGTGGTTGCTGCTTGCTAGTGTTATTTTTGTGCCTTTATTTCAGAAAATTCCAGGAGGCA
GTCCTGTTCTTGGATACTTGGCTGCTGGCATCTTGATTGGTCCTTATGGTCTCTCCATCATCCGTCATGTGCTTGGTACAAAAGCGATTGCTGAATTTGG
AGTTGTTTTCTTGCTATTCAATATTGGCCTTGAGCTCTCTGTTGAAAGGCTTAGTTCCATGAAAAAATATGTTTTTGGATTAGGCTCTGCTCAGGTCTTG
GTGACAGCAGTGGTTATTGGCTTGGTCACACATTTTGTTTCCAGGCTGCCTGGTCCTGCTGCAATTGTCATTGGAAATGGCCTGGCACTATCTTCTACTG
CTGTTGTTCTACAGGTGTTGCAGGAGCGAGGTGAGAGCACATCACGCCATGGACGTGCTACTTTCTCTGTCCTACTATTCCAGGATTTGGCTGTTGTGGT
GTTGCTAATACTCATTCCACTTATTTCACCAAATTCATCCAAAGGAGGGGTTGGTTTTCAAGCAATTGCTGAGGCTCTTGGGTTGGCTGCTGTTAAGGCA
GCAGTTGCCATCACCGCGATAATTGCTGGTGGAAGATTGCTGCTTCGACCAATTTATAAACAAATTGCAGAAAATCAAAATGCAGAGATATTCTCAGCCA
ATACACTTCTTGTTATTCTGGGGACCAGTCTCCTTACTGCCAGGGCTGGACTTTCCATGGCACTGGGGGCATTTCTGGCTGGTTTGCTTCTTGCAGAAAC
CGAATTTTCCTTGCAGGTTGAATCAGATATTGCTCCATATCGCGGGCTTCTATTGGGCCTTTTTTTCATGACGGTTGGAATGTCCATTGACCCAAAGCTT
CTGGTTTCAAATTTTCCTGCCATTATGGGATCATTAGGGCTTTTAATTGGAGGCAAGACTGTGTTGGTTGCTTTAGTTGGCAGGTTTTTTGGTATTTCTA
TTATATCTGCAATAAGGATTGGTCTTCTTCTTGCTCCGGGTGGAGAATTTGCATTTGTAGCATTTGGTGAAGCTGTTAACCAGGGTATAATGTCTCCTCA
GCTATCCTCTTTATTGTTTCTTGTGGTGGGGATCTCAATGGCCCTCACACCATGGCTGGCTGCTGGCGGCCAGTTAATAGCCTCTCGCTTTGAGCAGCAT
GATGTTCGTAGTTTGTTACCGGTTGAGAGTGAGACAGATGATTTGCAGGATCATATTATTATCTGTGGATTTGGACGTGTTGGCCAGATTATAGCACAGC
TTCTCTCAGAACGATTAATTCCATTTGTTGCTCTTGACGTGAGTAGCGATAGAGTGGCAGCAGGCCGTGCCCTGGATCTTCCCGTGTACTTTGGAGATGC
TGGTAGTCGAGAGGTCCTCCATAAAGTCGGTGCTGAAAGAGCATGTGCTGCTGCAATAACTCTAGATACACCTGGTGCAAATTATAGAACTGTCTGGGCT
CTGAGCAAGTACTTTCCCAATGTGAAGACGTTTGTTCGTGCTCATGATGTTGATCATGGCCTTAATTTGGAGAAGGCTGGGGCTACAGCTGTTGTACCTG
AGACATTGGAACCAAGTCTACAGTTGGCAGCTGCTGTTCTTGCGCAGGCTAAACTGCCCATGTCAGAGATTGCAGCAACCATCAATGCATTTAGATCCCG
CCACTTGTCTGAGCTTACTGAGCTATGCGAAACTAGTGGAAGTTCTCTCGGATATGGATTTTCTCGTGTCATGACTAAACCCAAAAGTCAGTCCTTGGAT
TCTTCAGATGAAAACCAATTCAGTGAAGGAACATTAGCAATATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G157200.1 pacid=42779795 polypeptide=Potri.002G157200.1.p locus=Potri.002G157200 ID=Potri.002G157200.1.v4.1 annot-version=v4.1
MEFACNIQQTNAFYSRQGTSCRVSNRLYSRFRHKSYGYNAGDLKIVSRERPSKTLKKSVLYGSGSGMRSHLLVGGYASNPLFCNFIDGFEGLRSVKLRCQ
GNDSLTYIDGNGRNVEIGKGNDKNLRAGSNGGLGEEDGRGEKVMETEMAAEALSLDELRELLQKAMRELEVARLNSTMFEEKAQSISETAIALQDEASSA
WNDVNSTLDMIQDIVNKEGVAKEAFQKATMALSLAEARLKVAVESIKSTKEGVDSLEGSGESDVQNDNKEDYETILAAQNDIRECQANLANCEAELRRLQ
SIKEELQKEVDVLNEKAEKAQMNALKAEEDVANIMLLAEQAVAFELEATQRVSDAEIALQKAEKSLSSSHVDIQQTARGYVSGDEAVVEEEKMRGGSLSD
VEKETDMTVNGDVLVGEPSIDRLSDKTSPSSEELYLSDYSSDHENGKSSLHSIKDTEAEAEKSKGGIQTKKQELQKDLTRESSSSPLSAPKALLKKSSRF
FSASFFSFSGDETELTAASVFQGLMEPARKQLPKFLLGLLLFGAGFAFYSNRVEKSTQMLQKPEVVTTSIEEVSSNAKPLIQHIQKLPKRVKKLIAMLPH
QEMNEEEASLFDVLWLLLASVIFVPLFQKIPGGSPVLGYLAAGILIGPYGLSIIRHVLGTKAIAEFGVVFLLFNIGLELSVERLSSMKKYVFGLGSAQVL
VTAVVIGLVTHFVSRLPGPAAIVIGNGLALSSTAVVLQVLQERGESTSRHGRATFSVLLFQDLAVVVLLILIPLISPNSSKGGVGFQAIAEALGLAAVKA
AVAITAIIAGGRLLLRPIYKQIAENQNAEIFSANTLLVILGTSLLTARAGLSMALGAFLAGLLLAETEFSLQVESDIAPYRGLLLGLFFMTVGMSIDPKL
LVSNFPAIMGSLGLLIGGKTVLVALVGRFFGISIISAIRIGLLLAPGGEFAFVAFGEAVNQGIMSPQLSSLLFLVVGISMALTPWLAAGGQLIASRFEQH
DVRSLLPVESETDDLQDHIIICGFGRVGQIIAQLLSERLIPFVALDVSSDRVAAGRALDLPVYFGDAGSREVLHKVGAERACAAAITLDTPGANYRTVWA
LSKYFPNVKTFVRAHDVDHGLNLEKAGATAVVPETLEPSLQLAAAVLAQAKLPMSEIAATINAFRSRHLSELTELCETSGSSLGYGFSRVMTKPKSQSLD
SSDENQFSEGTLAI
|
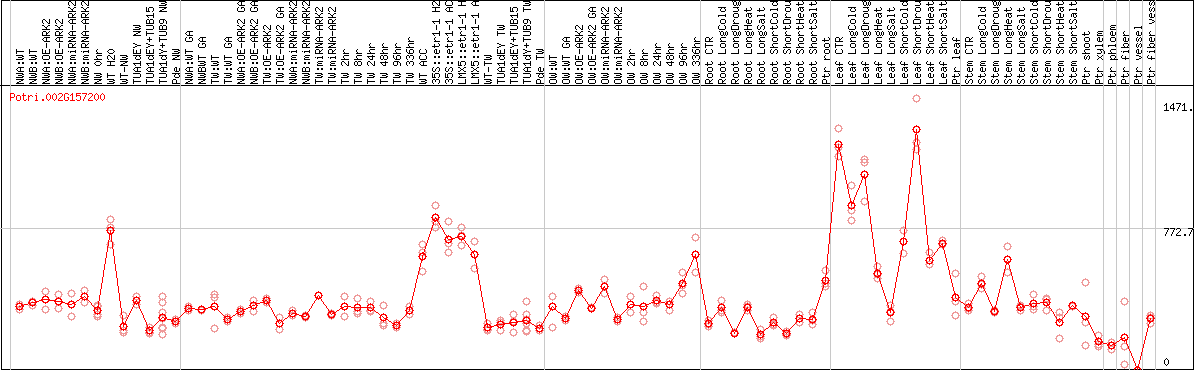
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G157200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
