External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G61770 331 / 3e-114
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1)
AT1G67600 126 / 7e-36
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1)
AT1G24350 123 / 8e-35
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1.2.3)
AT3G21610 114 / 3e-31
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1.2)
AT3G12685 72 / 5e-15
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G056600
126 / 1e-35
AT1G67600 254 / 8e-88
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1)
Potri.011G087100
120 / 2e-33
AT3G21610 204 / 9e-68
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1.2)
Potri.014G156000
104 / 4e-27
AT3G21610 184 / 5e-60
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1.2)
Potri.002G226900
100 / 7e-26
AT3G21610 218 / 2e-73
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1.2)
Potri.010G176100
64 / 8e-12
AT3G12685 193 / 2e-61
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030253
242 / 2e-81
AT3G61770 245 / 5e-83
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1)
Lus10014550
117 / 2e-33
AT3G61770 115 / 4e-33
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1)
Lus10006432
119 / 4e-33
AT1G67600 245 / 3e-84
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1)
Lus10011373
119 / 5e-33
AT1G67600 245 / 2e-84
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1)
Lus10003670
117 / 7e-32
AT3G21610 219 / 1e-73
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1.2)
Lus10000883
116 / 7e-32
AT3G21610 224 / 3e-76
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1.2)
Lus10035299
111 / 1e-29
AT3G21610 275 / 1e-95
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1.2)
Lus10030025
112 / 2e-29
AT3G21610 243 / 6e-82
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1.2)
Lus10036984
98 / 2e-25
AT1G24350 185 / 2e-61
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1.2.3)
Lus10001841
76 / 1e-15
AT3G12685 206 / 2e-66
Acid phosphatase/vanadium-dependent haloperoxidase-related protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0525
pap2
PF02681
DUF212
Divergent PAP2 family
Representative CDS sequence
>Potri.002G171300.1 pacid=42776934 polypeptide=Potri.002G171300.1.p locus=Potri.002G171300 ID=Potri.002G171300.1.v4.1 annot-version=v4.1
ATGGATTCTCTAACACTCACCAAACCCATCTTTAACTCTTTCTACTTCACTCCCACAAATCCAAGAACCAAACTTTGGTGTTCTGGCCACTTTTCATTTT
TCAAGTCCACTCACCTTACCAAATTCAAAAACTATCCAATTTGTTTTTCGTATAAAAACCCTTCAAATCATCAAAACCCAAAACCAAGAAACAAAAATAA
CAGAACTTTCTTATCAAACCCTTTGCGGGCTTTGGTCCCGATAATGGAACACATCAAAGCCTTAGCTTCTTCACAAACAAAAAAATGGGCTTCACAGCTA
CAAGCATACAGTGATGACTCGGAGAAGGTGATGGGTGAACACAACGGGGATTTTACGCTAAATGGAGCGTTTGGATTGGCTTTATTGAGTATAACATCGC
ATGCGAAAGTAAAGATCAGTCCTTTTGTGGCAACATTAGCTGCAAACCCAACTTTTATATCGGGTTTGTTTGCTTGGTTTATTGCACAGTCAATGAAAGT
GTTTTTGAATTTTTTTGTGGAGAGGAAATGGGATTTGAGATTATTGTTTGCCTCTGGAGGGATGCCGTCTTCCCACTCTGCTTTGTGTACTGCTTTGACA
ACATCAGTTGCGTTTTGTCATGGAGTGGCTGATTCGTTATTTCCTGTTTGTTTGGGGTTTAGTCTTATAGTTATGTATGATGCAATTGGTGTCAGGCGTC
ATGCTGGGATGCAAGCTGAGGTTCTTAATATGATCGTTGAGGACCTGTTTCAAGGGCATCCTATCAGTCAAAGAAAGCTCAAGGAACTTCTTGGACATAA
CCCATCACAAGTTCTTGCTGGAGCTTTGCTTGGCATCTTGGTAGCCTGTGTATGTTGCCAAGGTTGCTTAGTGGTAACCTAG
AA sequence
>Potri.002G171300.1 pacid=42776934 polypeptide=Potri.002G171300.1.p locus=Potri.002G171300 ID=Potri.002G171300.1.v4.1 annot-version=v4.1
MDSLTLTKPIFNSFYFTPTNPRTKLWCSGHFSFFKSTHLTKFKNYPICFSYKNPSNHQNPKPRNKNNRTFLSNPLRALVPIMEHIKALASSQTKKWASQL
QAYSDDSEKVMGEHNGDFTLNGAFGLALLSITSHAKVKISPFVATLAANPTFISGLFAWFIAQSMKVFLNFFVERKWDLRLLFASGGMPSSHSALCTALT
TSVAFCHGVADSLFPVCLGFSLIVMYDAIGVRRHAGMQAEVLNMIVEDLFQGHPISQRKLKELLGHNPSQVLAGALLGILVACVCCQGCLVVT
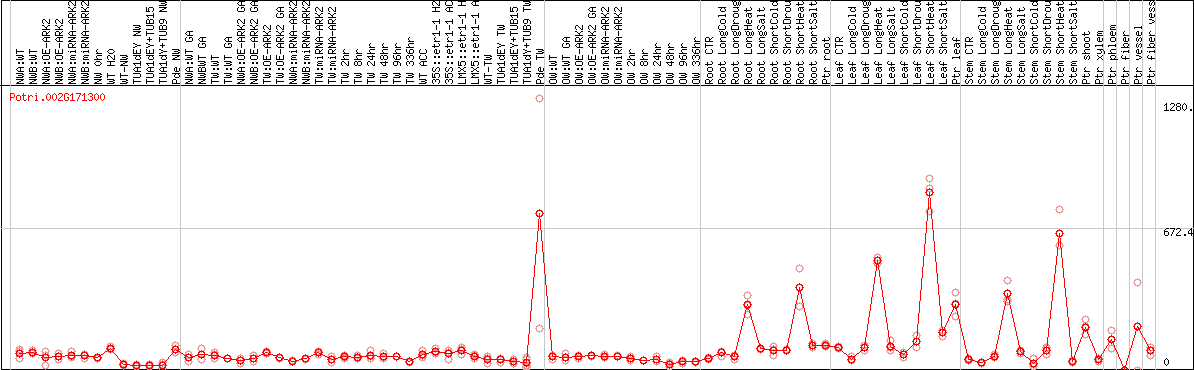
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G171300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.