External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G12915 114 / 4e-30
Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
AT1G56070 119 / 6e-10
LOS1, AT1G56075.1
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
AT1G06220 45 / 6e-06
GFA1, CLO, MEE5
MATERNAL EFFECT EMBRYO ARREST 5, GAMETOPHYTE FACTOR 1, CLOTHO, Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
AT5G25230 39 / 0.0007
Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G098100
128 / 3e-13
AT1G56070 1648 / 0.0
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Potri.007G065600
123 / 3e-12
AT1G56070 1620 / 0.0
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Potri.007G065700
123 / 1e-11
AT1G56070 1623 / 0.0
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Potri.018G114900
45 / 3e-06
AT1G06220 1710 / 0.0
MATERNAL EFFECT EMBRYO ARREST 5, GAMETOPHYTE FACTOR 1, CLOTHO, Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
Potri.006G190600
45 / 3e-06
AT1G06220 1744 / 0.0
MATERNAL EFFECT EMBRYO ARREST 5, GAMETOPHYTE FACTOR 1, CLOTHO, Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031779
124 / 1e-11
AT3G12915 760 / 0.0
Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
Lus10031186
124 / 6e-11
AT1G56070 902 / 0.0
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Lus10031764
52 / 2e-09
AT1G56070 158 / 8e-46
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Lus10031129
46 / 3e-06
AT1G06220 1751 / 0.0
MATERNAL EFFECT EMBRYO ARREST 5, GAMETOPHYTE FACTOR 1, CLOTHO, Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
Lus10043478
45 / 3e-06
AT1G06220 729 / 0.0
MATERNAL EFFECT EMBRYO ARREST 5, GAMETOPHYTE FACTOR 1, CLOTHO, Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
Lus10034111
45 / 5e-06
AT1G06220 1761 / 0.0
MATERNAL EFFECT EMBRYO ARREST 5, GAMETOPHYTE FACTOR 1, CLOTHO, Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
Lus10031763
0 / 1
AT1G56070 1418 / 0.0
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Lus10031201
0 / 1
AT1G56070 1415 / 0.0
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0329
S5
PF03764
EFG_IV
Elongation factor G, domain IV
Representative CDS sequence
>Potri.002G176966.1 pacid=42779132 polypeptide=Potri.002G176966.1.p locus=Potri.002G176966 ID=Potri.002G176966.1.v4.1 annot-version=v4.1
ATGGAGGCAGGGCCCACGGAAGAAGGCCTTGCTGAGGCTATTGACGTGGGCCATATTGGTCCAAGAGACGATCCTAAGAATCGCGCCAAGATCTTGTCGG
AGGAGTTTGGTTGGGACAAGCATCTTGCGAAGAAAATCTGGTGTGTTGGTCCTGAGACTACGGGACCCAACGTGGTTGTGGATATGTGTAAGGGAGTTCT
GTTGTTGCTGGGTTCCAATGGGCTTCAAAGGAAGGTGCCTAGGCTGAAGAAAACAAGAGAGGTATCTGCTTTGAAGTCTGTGATGTGGTTCTCCCTGCTG
ATGCCATTCACACAACCTTTGGAGGCTGGCACACAAGCAGCTCAGCTTGTTACTGATATCACGAACCGAAAGGGTTTGAAGGAGCAGATGACACCTCTAT
CTGAGTTTGAGAACAAGCTATAA
AA sequence
>Potri.002G176966.1 pacid=42779132 polypeptide=Potri.002G176966.1.p locus=Potri.002G176966 ID=Potri.002G176966.1.v4.1 annot-version=v4.1
MEAGPTEEGLAEAIDVGHIGPRDDPKNRAKILSEEFGWDKHLAKKIWCVGPETTGPNVVVDMCKGVLLLLGSNGLQRKVPRLKKTREVSALKSVMWFSLL
MPFTQPLEAGTQAAQLVTDITNRKGLKEQMTPLSEFENKL
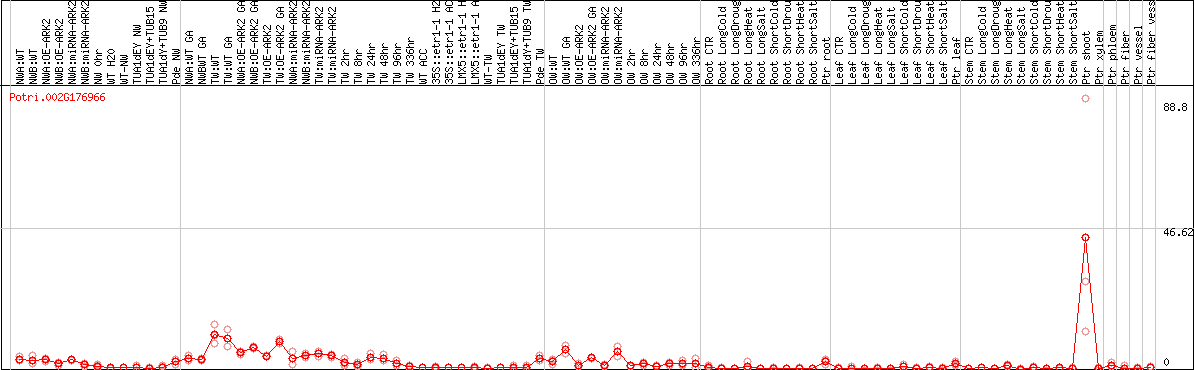
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G176966 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.