SIN1.1,DCL902 (Potri.002G181400) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | SIN1.1,DCL902 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G181400.1 pacid=42779272 polypeptide=Potri.002G181400.1.p locus=Potri.002G181400 ID=Potri.002G181400.1.v4.1 annot-version=v4.1
ATGGAGAGCGAGAGTAATGGTAGGGTTTCGGGGATTGGAGGTGGAGGTGGTCCGTCTTACTGGCTTGATGCATGTGAGGATATTTCTTGTGATATTATAG
ATGATTTTGTTGATTTTGATACTTCAATTGTTCCAGAATTATCTGTTGATAATAATTCCAATGTTAATAATGATTTCTTTGGAGGGATTGATCACATTCT
TGATAGTATCAAGAATGGTAGTGGTCTTCCTCCTCTTCATAATGCTAGTACGACAGCTAATGTTTCCAATGGAAATCGGGATTGCATTGTTGGTGATGGT
TGGTTTATTAATGTTGAAAATGGAGTTTGCCATGGGAGCTCGGTTTCACAATCAAATGGTGGTGATAAGGACAATATTGACAGAAAAGGGCAGGTGGAGA
ATGGTGGTAATGGGTTGAATTTGTCCAATGGGAAGAGAGAAGAGAGGTTCTCAAACAATTTTGTTAAAGAAAATGGGAAGAAGGATGAGCAGTCGACGGA
GCAAGGTATTGATGGTGATGAAAGGTGCGGTAAAAGGGCTAGACTTTGTTGTTATAGAAATGAGAGAGTGTATTCTAGTAGAGGGCAGCATGAGCATAGG
GATAGGGAGAGATGTTCTAGCAGGAAGAGGTCACGGGATTGGGATGAAAGCGATAGAAGGGATAGGGATATTAGTCGAAGAAGAGATAGATATAGTGGTA
GTAATAGGAGGGATGGTAGAGATAGGGATTGGAGGGAAAGAGAACTTAGGGGTTATTGGGAGAGGGATCGTTCAGGGTCCAAAGACATGGTTTTCCGATT
GGGCACTTGGGAAGCTGATCATAACAAAGAAGGAAGGGAGGCAAATGACAAAATTCAGGAGTGCAAAGGAGAGCTAGAGAAGAAATCAGAGGAGAGTAAA
GAGAAAGTCCCTGAAGAACAGGCCCGCCAGTATCAATTGGATGTTCTTGACCAGGCTAAGAAGAAAAATACAATAGCTTTTCTTGAAACAGGGGCAGGGA
AGACACTCATTGCTGTACTCCTCATCAGAAGCATTTGCAATGACTTGCAAAGGCAGAATAAAAAAATTCTAGCTGTATTTTTAGTTCCGAAGGTTCCACT
TGTTTATCAGCAAGCAGAAGTTATTCGTGAGCGCGGTTACCAAGTGGGCCATTATTGTGGTGAGATGGGTCAAGATTTCTGGGATACTCGAAGATGGCAG
CGTGAGTTTGAAACAAAACAGGTTTTAGTAATGACAGCCCAAATTCTTTTGAACATTCTGAGGCACAGCATAATCAAAATGGAAGCCATAAATCTCCTCA
TTCTGGATGAGTGTCATCATGCTGTGAAGAAACATCCATATTCTCTTGTGATGTCTGAGTTTTACCATACGACTCCCAAGGAGAAGAGACCGTCTGTTTT
TGGAATGACAGCTTCTCCTGTTAACTTAAAGGGTGTTTCAAGTCAGGTGGATTGTGCAATAAAGATCCGCAATCTTGAAAGTAAACTGGACTCAATTGTT
TGTACGATAAAAGACCGCAAGGAACTGGAAAAGCATGTACCAATGCCGGCAGAGGTTGTGGTGGAGTACGACAAAGCAGCTAGCTTGTGGTCCCTTCATG
AACAAATAAAGCAAATCGAAGCAGCTGTTGAGGAAGCTGCACAGTCTAGCTCGAGAAGAAGTAAATGGCAATTTATGGGAGCTAGGGATGCTGGGGCCAA
GGAAGAGCTGCGCCAGGTTTATGGTGTCTCTGAAAGAACAGAAAGTGATGGGGCTGCTAACTTAATTCAAAAGTTGAGGGCAATTAATTATGCACTTGGT
GATCTGGGTCAGTGGTGTGCTTACAAGGTTGCACAATCCTTTCTGACTGCCTTACAAAATGACGAGAGAGCAAACTATCAGCTTGATGTTAAGTTTCAAG
AATCCTACCTCGAAAGAGTCGTGTTGCTCTTACAATGCCAATTAACAGAGGGAGCTGTTACAGACAAGGATACAAAAGTTTCAGACAATGGGAATGACAA
TATTCAGGATGGGCCTGGTTTTGATGAGATTGAGGAGGGAGAGCTCCCTGACAGTCATGTTGTCTCTGGTGGAGAACATGTTGATGTGATAATAGGAGCT
GCTGTAGCTGATGGAAAAGTGACCCCTAAAGTGCAGTCACTGATTAAAGTACTTCTCAGATATCAGCACACGGAGGATTTTCGTGCGATCATCTTTGTTG
AGCGCGTAGTAGCTGCTTTGGTTCTTCCTAAGGTCTTTGCAGAGCTTCCATCTCTAAGTTTTGTTAGGTGTGCAAGTTTGATAGGGCACAATAATAGTCA
GGAAATGAGAACAAGCCAAATGCAAGACACAATTGCCAAATTTCGAGATGGTCGTGTGACATTGTTAGTTGCTACCAGTGTTGCTGAGGAAGGACTTGAT
ATTAGGCAATGCAATGTTGTTATTCGCTTTGATCTTGCAAAAACTGTTCTTGCATATATTCAGTCTCGAGGTCGTGCAAGAAAGCCAGGCTCTGATTACA
TCTTGATGGTGGAGAGGGGAAACCTGTCTCATGGAGCATTCTTAAGGAATGCTAGGAATAGTGAGGAGACCCTGAGGAAAGAAGCGATTGAGAGAACTGA
CCTCAGTCATCTTAAGGATACTTCAAGGTTAATTGCAGTAGATTCAATACCAGGAACTGTGTATCAGGTGGAGTCTACTGGTGCTGTTGTGAGCTTAAAT
TCTGCTGTTGGACTTGTTCATTTCTATTGCTCTCAGTTACCCAGTGACAGGTATTCAATACTTCGTCCTGGGTTTATTATGGAGAAGCATGAGAAGCCAG
GAGGTCCCACGGAATATTCATGCAAGCTTCAGCTTCCTTGTAATGCACCATTTGAGGAGCTTGAGGGTCCTGTATGCAGTTCAATGCGCCTTGCACATCA
GGCTGTTTGTTTGGCTGCTTGCAAGAAGCTTCATGAGATGGGAGCATTTACTGACATGCTCTTGCCAGACAAGGGAAGTGAGGAAGAAAAGGACAAGGTT
GACCAGAATGATGAAGGTGAACCACTTCCTGGTACTGCTAGGCATCGGGAGTTCTATCCTGAAGGTGTAGCTAAGACACTCCAGGGAGAATGGATCTTAT
GCGGAAGGGATGGTTGCAACAACTCCAAAGTGCTTCATCTCTACTTGTATGGAGTCAGATGCTTAAACATTGGCACTTCAAATGATCCATTCTTGACTCA
AGTTTCAAATTTTGCAGTGCTTTTTGGCAATGAGCTGGATGCAGAGGTCTTATCGATGTCGATGGATCTATTTATCGCTCGAACCATGATAACGAAGGCA
TCTCTTGTCTTTAGGGGTCGCATACCTATTACAGAAAGTCAGCTGGCATCCCTTAAAAATTTTCATGTAAGGTTGATGAGCATTGTACTGGATGTGGATG
TTGAACCTTCCACCACTCCATGGGATCCTGCAAAAGCTTATTTATTCGTCCCTATGGTTAGCGATAAGTCTGTAGATCCTATAAAAGAAATTGACTGGGA
TCTGGTTGAAAATATAATTGGAACAGATGCTTGGAGCAATCGTCTTCAGAGAGCTCGACCAGATGTTTACCTTGGTACAAATGAGAGGACACTTGGTGGA
GACAGGAGGGAGTATGGTTTTGGGAAATTGCGACATGGAATTGCTTTTGGACAGAAACCTCATCCCACTTATGGCATCAGGGGAGCTGTAGCACAGTTTG
ATGTTGTGAAAGCTTCTGGATTGATTCCTAAGAGGGGATGGGATGCAACTGAGACGCAAAAACTGGAGCTGACCAAAGGCAAATTGATGATGGCTGATAC
TTGTGTCAATGCTGATGCTTTGATGGGGAGAATTGTAACAGCTGCTCACTCTGGGAAGAGGTTTTATGTGGATTCCATATGCTATGATATGACTGCAGAG
ATCTCATTTCCAAGAAAAGAGGGTTATCTTGGTCCTCTGGAGTACAGCTCATATGCTGATTACTACAAGCAAAAGTATGGAGTTGAATTGAAATTTAAGC
AGCAGCCTTTACTAAGAGGACGTGGTGTTTCATACTGCAAGAATCTCTTATCTCCTCGGTTTGAACACTCAGATTCAAATGAAGGCGATGCTGAAGAGAA
TCTTGACAAGACTTACTATGTGTTTCTTCCTCCTGAGCTGTGTCTGGTGCACCCTCTTCCTGGATCACTTGTTCGAGGTGCCCAGAGATTACCATCGATA
ATGAGGAGGGTTGAAAGCATGCTGCTCGCTGTTGAACTTAAGGATATAATAAATTATCCAGTCCCTGCTTCAAAGATCTTGGAAGCCTTGACTGCTGCTT
CATGCCAAGAGACATTCTGCTATGAAAGAGCAGAGCTTCTAGGAGATGCTTACCTGAAATGGGTGGTTAGTCGATTTCTTTTTCTGAAGTATCCCCAGAA
ACATGAGGGTCAACTTACTAGAATGAGGCAACAAATGGTAAGTAATATGGTTTTGTATCAATATGCACTTAACAAAGGACTTCAGTCATATATCCAAGCA
GATCGCTTTGCTCCATCTCGATGGGCTGCTCCTGGAGTGTTGCCTGTCTTTGATGAGGAAACAAAAGATGGAGACTCGTACATATTTGATCAGGAGAAGT
CACTTGCTGAGGATAGAACCGGAATGAATCATCTTGATGATGGGTATGAAAATGAAATTGAAGATGGTGAACTTGAGAGTGATGCTAGTTCTTACAGAGT
CCTCTCTAGCAAGACATTGGCAGATGTTGTTGAAGCATTAATTGGTGTTTATTATGTTGAAGGTGGGAAAAATGCTGTTAACCACCTTATGAAATGGATT
GGAATTCAGGTAGAGTTTGATCATGAAGAGATAGATGGTGCATCTAGGCCATTCAATGTTCCAGAAAGTGTTCTTAGGAGTGTTGACTTTGATACTTTAG
AAGGTGCCTTGGATATTAAGTTCAATGATCGTGGCTTGTTGATAGAAGCCATAACGCATGCTTCACGACCATCTTCTGGAGTATCCTGCTATCAACGATT
AGAATTTGTTGGTGACGCAGTCTTAGATCATCTTATTACTAGGCACTTGTTTTTTACATATACAAATCTGCCCCCAGGTCGCCTGACAGACTTGCGAGCC
GCTGCTGTAAACAATGAAAATTTTGCACGTGTTGCAGTTAAACATAAGCTTCATGTGCACCTTAGACATGGGTCAAGTGCCCTTGAAAAGCAGATTCGGG
ACTTTGTTAGGGAAGTTCAAGATGAACTATTGAAGCCAGTGTTCAACTCCTTTGGTTTGGGAGATTGCAAAGCTCCAAAGGTTCTTGGTGACATCGTTGA
ATCCATTGCTGGTGCTATTTTTCTTGACAGCGGACGAGATACAGCGGTTGTGTGGAAGGTTTTTCAACCTCTGTTGCATCCAATGGTCACTCCAGAAACC
CTGCCAATGCATCCTGTAAGGGAGCTGCAGGAACGGTGCCAGCAACAGGCTGAAGGATTGGAATACAAAGCCACTCGAAGTGGCAATTTGGCTACTGTTG
AAGTTTTCATTGATGGTGTTCAGGTAGGAGTTGCCCAAAATCCACAAAAGAAAATGGCACAGAAACTCGCAGCTAGGAATGCACTTGTTGTTTTGAAAGA
GAAGGAAACAGCTGAAGCCAAGGAAAAGAGTGATGAAAATGGGAAGAAGAAAAGGAACGGCAACCAAACCTTTACTAGACAAACATTAAATGATATCTGC
TTGCGTAGAAATTGGCCTATGCCATCATATCGGTGTGTCAATGAGGGTGGTCCTGCACATGCAAAGAGGTTTACTTTTGCAGTTCGTGTGAATACTACCG
ATAGAGGATGGACAGACGAATGTGTGGGAGAGCCGATGCCGAGTGTTAAGAAGGCCAAAGACTCTGCTGCAGTTCTTCTCTTGGAACTATTAAATAAACG
GTACTCATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G181400.1 pacid=42779272 polypeptide=Potri.002G181400.1.p locus=Potri.002G181400 ID=Potri.002G181400.1.v4.1 annot-version=v4.1
MESESNGRVSGIGGGGGPSYWLDACEDISCDIIDDFVDFDTSIVPELSVDNNSNVNNDFFGGIDHILDSIKNGSGLPPLHNASTTANVSNGNRDCIVGDG
WFINVENGVCHGSSVSQSNGGDKDNIDRKGQVENGGNGLNLSNGKREERFSNNFVKENGKKDEQSTEQGIDGDERCGKRARLCCYRNERVYSSRGQHEHR
DRERCSSRKRSRDWDESDRRDRDISRRRDRYSGSNRRDGRDRDWRERELRGYWERDRSGSKDMVFRLGTWEADHNKEGREANDKIQECKGELEKKSEESK
EKVPEEQARQYQLDVLDQAKKKNTIAFLETGAGKTLIAVLLIRSICNDLQRQNKKILAVFLVPKVPLVYQQAEVIRERGYQVGHYCGEMGQDFWDTRRWQ
REFETKQVLVMTAQILLNILRHSIIKMEAINLLILDECHHAVKKHPYSLVMSEFYHTTPKEKRPSVFGMTASPVNLKGVSSQVDCAIKIRNLESKLDSIV
CTIKDRKELEKHVPMPAEVVVEYDKAASLWSLHEQIKQIEAAVEEAAQSSSRRSKWQFMGARDAGAKEELRQVYGVSERTESDGAANLIQKLRAINYALG
DLGQWCAYKVAQSFLTALQNDERANYQLDVKFQESYLERVVLLLQCQLTEGAVTDKDTKVSDNGNDNIQDGPGFDEIEEGELPDSHVVSGGEHVDVIIGA
AVADGKVTPKVQSLIKVLLRYQHTEDFRAIIFVERVVAALVLPKVFAELPSLSFVRCASLIGHNNSQEMRTSQMQDTIAKFRDGRVTLLVATSVAEEGLD
IRQCNVVIRFDLAKTVLAYIQSRGRARKPGSDYILMVERGNLSHGAFLRNARNSEETLRKEAIERTDLSHLKDTSRLIAVDSIPGTVYQVESTGAVVSLN
SAVGLVHFYCSQLPSDRYSILRPGFIMEKHEKPGGPTEYSCKLQLPCNAPFEELEGPVCSSMRLAHQAVCLAACKKLHEMGAFTDMLLPDKGSEEEKDKV
DQNDEGEPLPGTARHREFYPEGVAKTLQGEWILCGRDGCNNSKVLHLYLYGVRCLNIGTSNDPFLTQVSNFAVLFGNELDAEVLSMSMDLFIARTMITKA
SLVFRGRIPITESQLASLKNFHVRLMSIVLDVDVEPSTTPWDPAKAYLFVPMVSDKSVDPIKEIDWDLVENIIGTDAWSNRLQRARPDVYLGTNERTLGG
DRREYGFGKLRHGIAFGQKPHPTYGIRGAVAQFDVVKASGLIPKRGWDATETQKLELTKGKLMMADTCVNADALMGRIVTAAHSGKRFYVDSICYDMTAE
ISFPRKEGYLGPLEYSSYADYYKQKYGVELKFKQQPLLRGRGVSYCKNLLSPRFEHSDSNEGDAEENLDKTYYVFLPPELCLVHPLPGSLVRGAQRLPSI
MRRVESMLLAVELKDIINYPVPASKILEALTAASCQETFCYERAELLGDAYLKWVVSRFLFLKYPQKHEGQLTRMRQQMVSNMVLYQYALNKGLQSYIQA
DRFAPSRWAAPGVLPVFDEETKDGDSYIFDQEKSLAEDRTGMNHLDDGYENEIEDGELESDASSYRVLSSKTLADVVEALIGVYYVEGGKNAVNHLMKWI
GIQVEFDHEEIDGASRPFNVPESVLRSVDFDTLEGALDIKFNDRGLLIEAITHASRPSSGVSCYQRLEFVGDAVLDHLITRHLFFTYTNLPPGRLTDLRA
AAVNNENFARVAVKHKLHVHLRHGSSALEKQIRDFVREVQDELLKPVFNSFGLGDCKAPKVLGDIVESIAGAIFLDSGRDTAVVWKVFQPLLHPMVTPET
LPMHPVRELQERCQQQAEGLEYKATRSGNLATVEVFIDGVQVGVAQNPQKKMAQKLAARNALVVLKEKETAEAKEKSDENGKKKRNGNQTFTRQTLNDIC
LRRNWPMPSYRCVNEGGPAHAKRFTFAVRVNTTDRGWTDECVGEPMPSVKKAKDSAAVLLLELLNKRYS
|
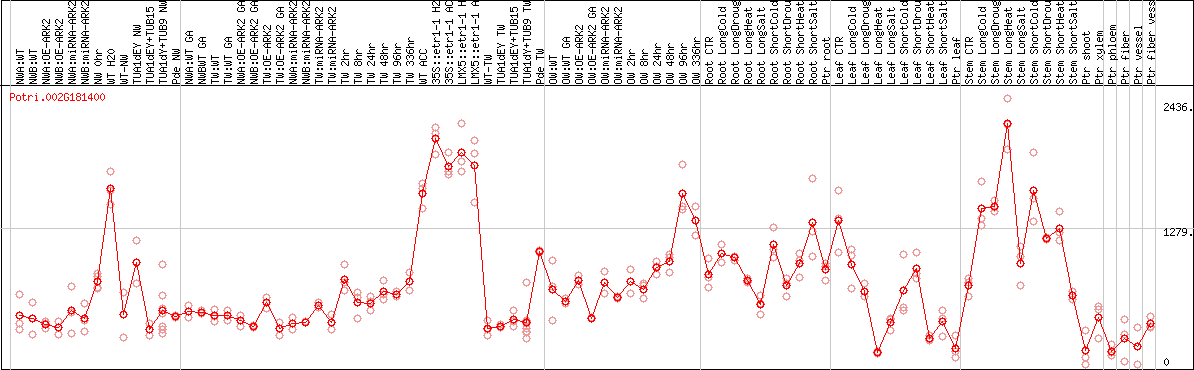
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G181400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
