External link
Symbol
ROC4.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G62030 294 / 2e-100
CYP20-3, ROC4
cyclophilin 20-3, rotamase CYP 4 (.1.2.3)
AT5G13120 235 / 2e-77
Pnsl5, ATCYP20-2
Photosynthetic NDH subcomplex L 5, ARABIDOPSIS THALIANA CYCLOPHILIN 20-2, cyclophilin 20-2 (.1.2)
AT5G58710 207 / 5e-67
ROC7
rotamase CYP 7 (.1)
AT2G29960 205 / 1e-66
CYP19-4, ATCYP5, CYP5
CYCLOPHILIN 19-4, ARABIDOPSIS THALIANA CYCLOPHILIN 5, cyclophilin 5 (.1.2)
AT2G21130 192 / 9e-62
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
AT4G38740 189 / 1e-60
ROC1
rotamase CYP 1 (.1)
AT3G55920 189 / 7e-60
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
AT2G16600 185 / 6e-59
ROC3
rotamase CYP 3 (.1.2)
AT3G56070 185 / 9e-59
ROC2
rotamase cyclophilin 2 (.1.2)
AT2G38730 177 / 1e-55
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G060200
246 / 7e-82
AT5G13120 297 / 7e-102
Photosynthetic NDH subcomplex L 5, ARABIDOPSIS THALIANA CYCLOPHILIN 20-2, cyclophilin 20-2 (.1.2)
Potri.003G167700
245 / 3e-81
AT5G13120 277 / 5e-94
Photosynthetic NDH subcomplex L 5, ARABIDOPSIS THALIANA CYCLOPHILIN 20-2, cyclophilin 20-2 (.1.2)
Potri.010G189000
211 / 3e-68
AT3G55920 303 / 4e-105
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Potri.009G046500
201 / 6e-65
AT2G29960 323 / 4e-114
CYCLOPHILIN 19-4, ARABIDOPSIS THALIANA CYCLOPHILIN 5, cyclophilin 5 (.1.2)
Potri.001G251700
201 / 1e-64
AT5G58710 313 / 6e-110
rotamase CYP 7 (.1)
Potri.009G130100
196 / 3e-63
AT2G21130 279 / 1e-97
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Potri.004G168800
196 / 5e-63
AT2G16600 278 / 4e-97
rotamase CYP 3 (.1.2)
Potri.009G106200
196 / 8e-61
AT2G15790 561 / 0.0
SQUINT, CYCLOPHILIN 40, peptidyl-prolyl cis-trans isomerase / cyclophilin-40 (CYP40) / rotamase (.1)
Potri.004G144300
191 / 1e-58
AT2G15790 590 / 0.0
SQUINT, CYCLOPHILIN 40, peptidyl-prolyl cis-trans isomerase / cyclophilin-40 (CYP40) / rotamase (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011330
247 / 3e-82
AT5G13120 336 / 2e-117
Photosynthetic NDH subcomplex L 5, ARABIDOPSIS THALIANA CYCLOPHILIN 20-2, cyclophilin 20-2 (.1.2)
Lus10003138
246 / 3e-81
AT5G13120 333 / 2e-115
Photosynthetic NDH subcomplex L 5, ARABIDOPSIS THALIANA CYCLOPHILIN 20-2, cyclophilin 20-2 (.1.2)
Lus10040666
206 / 1e-66
AT5G58710 347 / 2e-123
rotamase CYP 7 (.1)
Lus10018238
205 / 2e-66
AT2G29960 348 / 4e-124
CYCLOPHILIN 19-4, ARABIDOPSIS THALIANA CYCLOPHILIN 5, cyclophilin 5 (.1.2)
Lus10014846
201 / 8e-64
AT2G29960 310 / 6e-107
CYCLOPHILIN 19-4, ARABIDOPSIS THALIANA CYCLOPHILIN 5, cyclophilin 5 (.1.2)
Lus10012167
196 / 3e-63
AT2G16600 313 / 4e-111
rotamase CYP 3 (.1.2)
Lus10007579
196 / 3e-63
AT2G16600 311 / 2e-110
rotamase CYP 3 (.1.2)
Lus10038315
196 / 2e-62
AT2G29960 313 / 1e-109
CYCLOPHILIN 19-4, ARABIDOPSIS THALIANA CYCLOPHILIN 5, cyclophilin 5 (.1.2)
Lus10023860
196 / 9e-61
AT2G15790 599 / 0.0
SQUINT, CYCLOPHILIN 40, peptidyl-prolyl cis-trans isomerase / cyclophilin-40 (CYP40) / rotamase (.1)
Lus10020992
194 / 7e-60
AT2G15790 598 / 0.0
SQUINT, CYCLOPHILIN 40, peptidyl-prolyl cis-trans isomerase / cyclophilin-40 (CYP40) / rotamase (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0475
Cyclophil-like
PF00160
Pro_isomerase
Cyclophilin type peptidyl-prolyl cis-trans isomerase/CLD
Representative CDS sequence
>Potri.002G185200.1 pacid=42778066 polypeptide=Potri.002G185200.1.p locus=Potri.002G185200 ID=Potri.002G185200.1.v4.1 annot-version=v4.1
ATGGCCTCTACATCGTCAATGCAAATGGCACACAGTCCTCGCCTTGTTATTCAGGGTAGTTATGAACAAGATTTTTTGAATACAAGCAGGCCACATAAGA
TATCTACCATCAGAAATGCCCGGGGTGGTTACTCTCAAACTCCAACCTTGAACTGCCTTGGAAGAGCATTGACATCAAGATCTCACTATGCATCGAAATT
CCCTATCATACGGCTTCCTAAAGCACCTTATATTAAGAATCAGAGAATGGCATGCGCCAACTCAATGGCAAATGATGTTGATCTACAAGCAAAAGTGACG
ACCAAGTGTTTCTTTGATGTTGAAGTTGGGGGTGAACTTGTGGGTAGAATTGTAATGGGCCTTTTTGGAGATGTTGTCCCAAAAACTGCTGAGAACTTTC
GTGCCTTGTGCACAGGAGATAAAGGGTATGGTTACAAAGGATGCTCCTTCCATCGTATTATCAAAGATTTCATGATCCAGGGTGGTGACTTCACGAGAGG
AGATGGAACTGGAGGAAAAAGCATATATGGTTCTAGTTTTGAAGATGAGAGCTTTTCCTTGAAGCATGTTGGACCTGGTGTTTTGAGCATGGCAAATGCA
GGTCCCAATACCAATGGGAGCCAGTTTTTCATTTGCTCAGTAAAGACTCCATGGCTGGACAACCGCCACGTTGTCTTTGGACATGTCATTGATGGAATGG
ATGTTGTGCGCAAGCTTGAATCTGTAGAGACAAGCAGGTCAGATAATCCTCGGAAGCCATGCAGAGTTGTTAATTCTGGGGAGCTACCGCTAGATAGCTG
A
AA sequence
>Potri.002G185200.1 pacid=42778066 polypeptide=Potri.002G185200.1.p locus=Potri.002G185200 ID=Potri.002G185200.1.v4.1 annot-version=v4.1
MASTSSMQMAHSPRLVIQGSYEQDFLNTSRPHKISTIRNARGGYSQTPTLNCLGRALTSRSHYASKFPIIRLPKAPYIKNQRMACANSMANDVDLQAKVT
TKCFFDVEVGGELVGRIVMGLFGDVVPKTAENFRALCTGDKGYGYKGCSFHRIIKDFMIQGGDFTRGDGTGGKSIYGSSFEDESFSLKHVGPGVLSMANA
GPNTNGSQFFICSVKTPWLDNRHVVFGHVIDGMDVVRKLESVETSRSDNPRKPCRVVNSGELPLDS
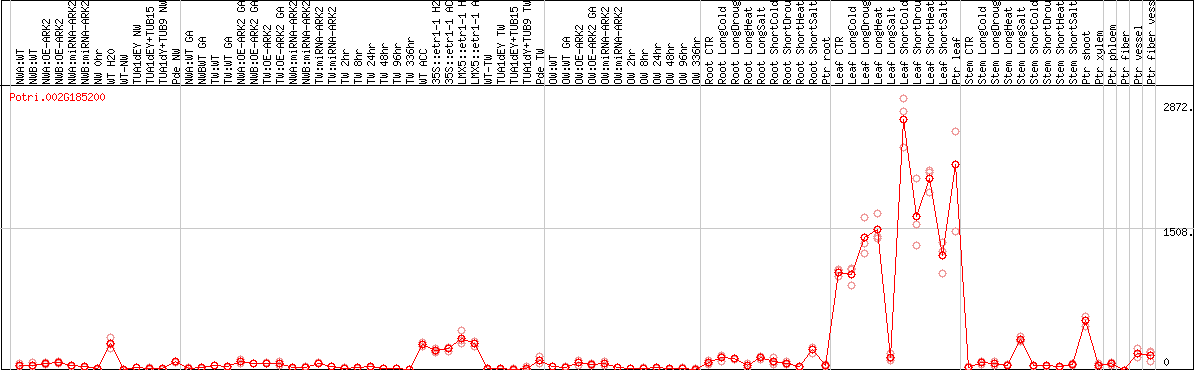
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G185200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.