Potri.002G189400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G189400.1 pacid=42777913 polypeptide=Potri.002G189400.1.p locus=Potri.002G189400 ID=Potri.002G189400.1.v4.1 annot-version=v4.1
ATGGAAGAGCAGCAGCAGAAACAGAGCAGGTGGAAACTAGAGAGAGGGCATTACCTAGGAGAAATATCAGCTCTCTGTTTCCTCCACCCTCCTTCTAACC
TCTCTTCTCTCCCTTTCCTCCTCGCTGGCACAGGATCGCAACTGCTGCTATATAATTTAGAATCAGGAAAAATAATAAAATCCTTTGAAGTTTTTGATGG
AATTCGTGTCCATGGAATTACTTGCAGCTCCAGTGAAGAAGAGTCATCGAACTTTCCTTCTTTAACTGTTTCTTTCAAAATTGCTGTCTTCGGTGAAAAA
AGACTCAAACTTTTCAACTTACACATTCAAACTCCTTCGCAAGTGAATGCAGATTTAGCTTTGATTCATTGCTTGCCCAAGTTCACTCATTGGGTCTTGG
ATGTTTCCTTCTTCAAGAACTCTGCAGTCTCTTCAAGTCAGGAAGAGAGACAATGTCTTGCAATTGGATGTAGTGATAACTCTGTCCATCTCTGGGATAT
GTCGGTGTCCAGTGTAGTTCTTCAAGTTCAATCTCCAGAGAGATGCCTTTTGTATTCCATGCGGTTATGGGGTGACAGTTTAGAAACTTTAAGAATTGCT
TCCGGTACCATATTCAATGAGATCATCGTTTGGAAAGTGGTTCCTGTAGAACCTCAACTTGGTGGGCTGCCTTCAACAAGTCTCTTGGAAGATGATATGT
ATTTGAGCTGTTCATTACCCGACTCTTCCCAGCTTCGTTTTCAGCAGCATAAATCTGCTCATATGTGCAGACTTATTGGACATGAAGGCTCAATTTTTCG
CATAGCATGGTCCTCAGATGGATCCAAACTGGTTTCCGTCTCTGATGATCGCAGTGCTCGTATCTGGGCTGTTAGGGATGAGCTGAAAGATTCAGATAAC
AGAGAGGAGAAGGTTGCAGGCCCTGTTCTGTTTGGTCATAATGCTAGAGTTTGGGACTGCTGTATCTGTGATTCTGTAATTGTCACTGCTGGTGAAGATT
GTACATGTCGTGTATGGAGATTGGATGGCAAACAGCTCAAGATGATTAAGGAACACATAGGAAGAGGTATATGGCGTTGTTTGTATGATCCAACCTCTTC
ACTCCTCATTACTGCTGGATTTGACTCTTCAATCAAAGTGCATCAAGTTTCTGCTTCTATTTCCCAGAGCTTGGAAGGACAAATTGAATCAAAACCATTC
ATTGACAGAATGGAGATATTTACCTGTCGCATCCCTAATTCATCTGAATATATTGGTCTCATGGACAGCAAAAGTGAATATGTACGTTGCTTGCATTTCA
CATGCGAAGATACTCTTTATGTTGCGACAAACAATGGTTATCTGTACCATGCTAGATTACATGGAACTGTGGATGTAAAATGGACAAAGCTTGCTCAGCT
CAGTGAAGAGGTGCCGATTGTTTGTATGGATTTGCTGTCCAAAAAACTCCCCAAACATTCCAATGGAGTTGATGACTGGGTTGCTTTAGGAGATGGTAAA
GGAAACATGACAATTGTGAGAATTATGGGTGACGTTTTCACTCCTGAAGTGGGTTTCACCGTCACCTGGTCAGCCGGGAAAGAGAGACAACTCTTAGGAA
CCTATTGGTGCAAGGCATTGGGATGTAGGTTCATTTTTACTGCTGATCCTAGAGGAATTTTGAAGCTATGGAGGTTAAGCGATCCTTTACCATCTGGTTC
TCTCACTTATGGAAGAACTTTTGATGCATCCCTTATAGCAGAATTCACATCATGTTTTGGAATTCGGATTATGTGTCTAGATGCATCATTTGAGGATGAG
GTGCTGGTATGTGGGGATTTGCGTGGAAATCTGGTTCTTTTCCCTTTGTCAAAGGGCTTGCTGCTGGACAAACCTACTCTTCCAGAAATAAAAATTTCAC
CATTGTGTTATTTTAAAGGATCTCATGGAATATCAACTGTATCCAACATTTCTGTTGCTAAATTGAGTGACACGATTGAAATACGCTCGACTGGAGGTGA
TGGTTGCATATGCTATTTAGAATATGACCCAGACCAGTGTGGTTTGGAATTTATTGGGATGAAACAAGTAAAAGAATTGAGTTTAGTTCAATCTGTCTCT
GCCGATAACAACTGCCTCGATGATTTGGCTAATTGTGGATATGCTATTGGTTTTGCATCAACAGATTTTATAATCTGGAACTTAATAAGTGAGGCAAAGG
TTGTGCAAATTCCATGTGGTGGATGGCGACGTCCTCATTCTTATTATCTTGGTGATGTACCAGAGGCGATGAGCTGTTTTGCATATGTCAAGGATGAGAT
AATCTATATTCATCGAAAATGGGTACCAGAAAGGGAGTGGAAGATATTTCCCCAGAATTTGCACACCCAGTTTCATGGGAGAGAGATGCACTCCTTATGC
TTTGTTTCTAAAAATACACTTGTTGAGGCAAATGGGAATGATTTTCAGTTTGATAGATCTAGTTGGATTGCTACTGGGTGTGAGGATGGAACTGTGAGGT
TGACTAGGTATATTCCTGGAGTTGAGGGTTGGCTTACATCAAAATTGCTTGGGGAACATGTTGGTGGATCTGCTGTGAGATCAATATGCTCAGTATCAAA
GATGCACATAATTGCTTCGGACTTGACCAACTTGTCAGATTGGACAAAGAGACAAAACACTTGTGCAGGGGACATGGATAATCCTTTCTTGTTGATCTCG
GTTGGGGCAAAGCGGGTGCTAACTTCTTGGCTACTGAGAGATAGGAATCTAGACAAGGAAAATGTATTTATTGAACAAGAAAAAATGGAAAACGAAAATG
GCTATAAGCCTTCGTCAGAAGTATCCTCATTGATGTCTTTCAAATGGCTGTCAACTGACATGCCTCCCAGAAATTCCAGTTCTCGTGGAAAAACAAAAGT
AGCTGAGAACATACAAGGAATAACCAAAGAACTTAATGTGAATATTGATGTAACATCAGGACCACTTTTACTGGAAAAGGGAGAGGGATATTCAAAAATT
TCTTATGATGATAAGTATGAGGATGACTGGAGATATCTGGCTGTCACTGCTTTTCTGGTGAAGTGTGCTGGTTCCAGGTTGACTGTCTGTTTTGTTGTTG
TTGCTTGCTCAGATGCAACACTTGCACTGCGAGCTCTTGTTTTACCACATCGGTTATGGTTCGATGTGGCTTTGTTGGTTCCTCTATCATCACCAGTTTT
AACATTGCAGCATGTCATCATTCCTAGCTGTTTGCCTTTTGAAGAGAACATTCGAATTGGAAATGTGTATATTGTGATCAGTGGAGCTACAGATGGAAGT
ATTGCCTTTTGGGATCTAACTGACAATATTGAGGCTTTTGTTCAGCGGTTATCAACACTGAACATAGAGAAGTCCATAAATTGTCAAACACGGCCACGCA
CTGGAAGAGGAAGTCAGGGTGGACGATGGTGGAGAACATTAAGCAGCGGTGTGCCTAAAAATAGACCAGGTGATGGTTTAGTTGCAATAAAAGCTGGAGA
GAGGACTAACTGCAATTTGGCAAATCATCCTATGAATGAAGCATCAACAGCAGTAAGTGACGCGGAAAACTGCACAATAGTTTGTTCTCAAGCTGTAGAC
AACACACATCATGAACCAGAAGTGAATAGTGTCAATTCCTTGCCAGGAATATGTGAAATAAGGCCTTTCCATGTCTTCAATAATGTTCACCAATCAGGGG
TCAACTCTCTTCATATTTCAGACATACAGGATATTCAAAGTTCTGAAAATGGTTTTGCATTCAGTGTAATAAGTGGAGGCGATGATCAAGCACTCCACTG
TCTTAAATTTGATTTGTCACCATTATCAACAGGGAAGGATTCTGATGTCGTGACATCAAATCTCATCAATTTATTCACTAGTTCTGAAAGCATGAAGAAT
AATTGTTGTAGACAAAGCCAGACAAACAAGTACAGGATCAGATTTTTATATCATGACCGAATCATCTCAGCTCATAGCTCTGCCATAAAAGGTGTTTGGA
CAGATGGCATGTGGGTGTTTTCTACTGGTCTTGATCAGCGCATCAGATGCTGGCTTCTTCAGGATAACTGTAAATTAACTGAACAAGCTTATTTGATCAT
TAGTGTTCCAGAGCCAGAAGCACTGCATGCTAGAGCCCGTGGCAGGAACCACTACGAAATTGCAGTGGCTGGGAGAGGAATGCAAATGGTTGAGTTCTCT
GCGTCTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G189400.1 pacid=42777913 polypeptide=Potri.002G189400.1.p locus=Potri.002G189400 ID=Potri.002G189400.1.v4.1 annot-version=v4.1
MEEQQQKQSRWKLERGHYLGEISALCFLHPPSNLSSLPFLLAGTGSQLLLYNLESGKIIKSFEVFDGIRVHGITCSSSEEESSNFPSLTVSFKIAVFGEK
RLKLFNLHIQTPSQVNADLALIHCLPKFTHWVLDVSFFKNSAVSSSQEERQCLAIGCSDNSVHLWDMSVSSVVLQVQSPERCLLYSMRLWGDSLETLRIA
SGTIFNEIIVWKVVPVEPQLGGLPSTSLLEDDMYLSCSLPDSSQLRFQQHKSAHMCRLIGHEGSIFRIAWSSDGSKLVSVSDDRSARIWAVRDELKDSDN
REEKVAGPVLFGHNARVWDCCICDSVIVTAGEDCTCRVWRLDGKQLKMIKEHIGRGIWRCLYDPTSSLLITAGFDSSIKVHQVSASISQSLEGQIESKPF
IDRMEIFTCRIPNSSEYIGLMDSKSEYVRCLHFTCEDTLYVATNNGYLYHARLHGTVDVKWTKLAQLSEEVPIVCMDLLSKKLPKHSNGVDDWVALGDGK
GNMTIVRIMGDVFTPEVGFTVTWSAGKERQLLGTYWCKALGCRFIFTADPRGILKLWRLSDPLPSGSLTYGRTFDASLIAEFTSCFGIRIMCLDASFEDE
VLVCGDLRGNLVLFPLSKGLLLDKPTLPEIKISPLCYFKGSHGISTVSNISVAKLSDTIEIRSTGGDGCICYLEYDPDQCGLEFIGMKQVKELSLVQSVS
ADNNCLDDLANCGYAIGFASTDFIIWNLISEAKVVQIPCGGWRRPHSYYLGDVPEAMSCFAYVKDEIIYIHRKWVPEREWKIFPQNLHTQFHGREMHSLC
FVSKNTLVEANGNDFQFDRSSWIATGCEDGTVRLTRYIPGVEGWLTSKLLGEHVGGSAVRSICSVSKMHIIASDLTNLSDWTKRQNTCAGDMDNPFLLIS
VGAKRVLTSWLLRDRNLDKENVFIEQEKMENENGYKPSSEVSSLMSFKWLSTDMPPRNSSSRGKTKVAENIQGITKELNVNIDVTSGPLLLEKGEGYSKI
SYDDKYEDDWRYLAVTAFLVKCAGSRLTVCFVVVACSDATLALRALVLPHRLWFDVALLVPLSSPVLTLQHVIIPSCLPFEENIRIGNVYIVISGATDGS
IAFWDLTDNIEAFVQRLSTLNIEKSINCQTRPRTGRGSQGGRWWRTLSSGVPKNRPGDGLVAIKAGERTNCNLANHPMNEASTAVSDAENCTIVCSQAVD
NTHHEPEVNSVNSLPGICEIRPFHVFNNVHQSGVNSLHISDIQDIQSSENGFAFSVISGGDDQALHCLKFDLSPLSTGKDSDVVTSNLINLFTSSESMKN
NCCRQSQTNKYRIRFLYHDRIISAHSSAIKGVWTDGMWVFSTGLDQRIRCWLLQDNCKLTEQAYLIISVPEPEALHARARGRNHYEIAVAGRGMQMVEFS
AS
|
DESeq2's median of ratios [POPLAR]
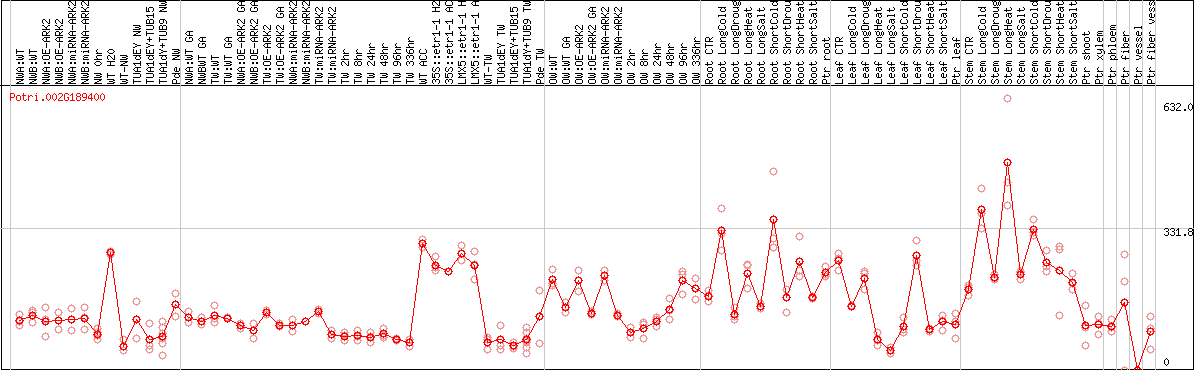
Coexpressed genes
Potri.002G189400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
