External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G02930 263 / 4e-90
ATGST1, GST1, ERD11, ATGSTF3, ATGSTF6
EARLY RESPONSIVE TO DEHYDRATION 11, ARABIDOPSIS THALIANA GLUATIONE S-TRANSFERASE F3, ARABIDOPSIS GLUTATHIONE S-TRANSFERASE 1, glutathione S-transferase 6 (.1.2)
AT1G02920 259 / 2e-88
ATGST11, GST11, ATGSTF8, ATGSTF7
ARABIDOPSIS GLUTATHIONE S-TRANSFERASE 11, glutathione S-transferase 7 (.1)
AT2G02930 258 / 5e-88
GST16, ATGSTF3
GLUTATHIONE S-TRANSFERASE 16, glutathione S-transferase F3 (.1)
AT4G02520 254 / 1e-86
GST2, ATPM24.1, ATGSTF2
glutathione S-transferase PHI 2 (.1)
AT2G47730 256 / 2e-86
GST6, ATGSTF5, ATGSTF8
Arabidopsis thaliana glutathione S-transferase phi 8, GLUTATHIONE S-TRANSFERASE \(CLASS PHI\) 5, glutathione S-transferase phi 8 (.1)
AT1G02950 254 / 9e-86
GST31, ATGSTF4
GLUTATHIONE S-TRANSFERASE 31, glutathione S-transferase F4 (.1.2.3.4)
AT1G02940 220 / 2e-72
ATGSTF5
glutathione S-transferase (class phi) 5 (.1)
AT3G62760 184 / 8e-59
ATGSTF13
Glutathione S-transferase family protein (.1)
AT3G03190 184 / 9e-59
ATGSTF6, ATGSTF11
ARABIDOPSIS GLUTATHIONE-S-TRANSFERASE 6, glutathione S-transferase F11 (.1)
AT2G30860 170 / 3e-53
GLUTTR, ATGSTF7, ATGSTF9
glutathione S-transferase PHI 9 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G207479
425 / 4e-154
AT1G02930 267 / 1e-91
EARLY RESPONSIVE TO DEHYDRATION 11, ARABIDOPSIS THALIANA GLUATIONE S-TRANSFERASE F3, ARABIDOPSIS GLUTATHIONE S-TRANSFERASE 1, glutathione S-transferase 6 (.1.2)
Potri.002G207093
300 / 2e-104
AT1G02930 263 / 5e-90
EARLY RESPONSIVE TO DEHYDRATION 11, ARABIDOPSIS THALIANA GLUATIONE S-TRANSFERASE F3, ARABIDOPSIS GLUTATHIONE S-TRANSFERASE 1, glutathione S-transferase 6 (.1.2)
Potri.002G207672
201 / 3e-65
AT3G62760 302 / 2e-105
Glutathione S-transferase family protein (.1)
Potri.014G132200
197 / 7e-64
AT3G62760 295 / 2e-102
Glutathione S-transferase family protein (.1)
Potri.002G015200
181 / 2e-57
AT3G03190 208 / 4e-68
ARABIDOPSIS GLUTATHIONE-S-TRANSFERASE 6, glutathione S-transferase F11 (.1)
Potri.017G138800
171 / 2e-53
AT5G17220 265 / 1e-90
TRANSPARENT TESTA 19, GLUTATHIONE S-TRANSFERASE 26, ARABIDOPSIS THALIANA GLUTATHIONE S-TRANSFERASE PHI 12, glutathione S-transferase phi 12 (.1)
Potri.002G015100
166 / 1e-51
AT3G03190 196 / 2e-63
ARABIDOPSIS GLUTATHIONE-S-TRANSFERASE 6, glutathione S-transferase F11 (.1)
Potri.019G077000
61 / 7e-11
AT1G57720 303 / 1e-99
Translation elongation factor EF1B, gamma chain (.1.2)
Potri.001G105600
60 / 1e-10
AT5G41210 358 / 1e-126
glutathione S-transferase THETA 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026643
279 / 4e-96
AT2G47730 269 / 2e-91
Arabidopsis thaliana glutathione S-transferase phi 8, GLUTATHIONE S-TRANSFERASE \(CLASS PHI\) 5, glutathione S-transferase phi 8 (.1)
Lus10030414
271 / 5e-93
AT2G47730 267 / 1e-90
Arabidopsis thaliana glutathione S-transferase phi 8, GLUTATHIONE S-TRANSFERASE \(CLASS PHI\) 5, glutathione S-transferase phi 8 (.1)
Lus10037121
258 / 1e-87
AT2G47730 243 / 2e-81
Arabidopsis thaliana glutathione S-transferase phi 8, GLUTATHIONE S-TRANSFERASE \(CLASS PHI\) 5, glutathione S-transferase phi 8 (.1)
Lus10004151
183 / 2e-58
AT2G30860 194 / 1e-62
glutathione S-transferase PHI 9 (.1.2)
Lus10040393
177 / 5e-56
AT3G03190 263 / 1e-89
ARABIDOPSIS GLUTATHIONE-S-TRANSFERASE 6, glutathione S-transferase F11 (.1)
Lus10029815
176 / 8e-56
AT2G30860 332 / 2e-117
glutathione S-transferase PHI 9 (.1.2)
Lus10020735
176 / 1e-55
AT2G30860 334 / 4e-118
glutathione S-transferase PHI 9 (.1.2)
Lus10001419
173 / 2e-54
AT2G30860 297 / 3e-103
glutathione S-transferase PHI 9 (.1.2)
Lus10020736
166 / 2e-51
AT2G30860 308 / 6e-108
glutathione S-transferase PHI 9 (.1.2)
Lus10005634
164 / 8e-51
AT2G30860 177 / 1e-55
glutathione S-transferase PHI 9 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0497
GST_C
PF00043
GST_C
Glutathione S-transferase, C-terminal domain
CL0172
Thioredoxin
PF02798
GST_N
Glutathione S-transferase, N-terminal domain
Representative CDS sequence
>Potri.002G207286.1 pacid=42779524 polypeptide=Potri.002G207286.1.p locus=Potri.002G207286 ID=Potri.002G207286.1.v4.1 annot-version=v4.1
ATGGCCCCTTTGAAACTCCATGGAAGCGTTTTATCTACCAACACACAGCGAGTTCTAGCAACCCTTTACGAGAAAGAGGTAGAATTTGAGCTTGTTAACG
TCAATTTGGGAGCTGGGGAACACAAGCAAGAGCCCCATATTTCCCTCAATCCATTTGGTCAAGTTCCAGCTGCTGTTGATGGAGATCTTAAGCTCTTTGA
ATCAAGAGCAATTTCACAGTACGTAGCCCACCAATATGCCAGCAAGGGGACTCAACTGGGCGCTGCAGGCAACGGGTATGCAACAATTTTGGTATGGCAA
GAAGTGGAGTCTCACCAGTTCGACCCATCAGCTTCAAAACTGGTGTGGGAGCAAGTTTTTAAACCAGTTTTTGGGCTACCAACAGATGCAGCTCTAGTTG
CCGAGACGGAGGTGACTCTCGGTAAGGTTCTAGATGTGTACGAGGCAAGACTGTCTCAATCCAAGTACCTAGCCAGTGACAGCTTCACCTTGGCTGATCT
GCACCACCTCCCTAACATCCAGGCCTTGTTGGGGACTCCATCCAAGAAATTGTTCGATTCTCGCCCCCATGTTAGCGCATGGGTTGCCAGTATCACTGGA
AGGCCAGCTTGGGGCAAGGTGCTCGCTTTGCTTCCCAAGTAA
AA sequence
>Potri.002G207286.1 pacid=42779524 polypeptide=Potri.002G207286.1.p locus=Potri.002G207286 ID=Potri.002G207286.1.v4.1 annot-version=v4.1
MAPLKLHGSVLSTNTQRVLATLYEKEVEFELVNVNLGAGEHKQEPHISLNPFGQVPAAVDGDLKLFESRAISQYVAHQYASKGTQLGAAGNGYATILVWQ
EVESHQFDPSASKLVWEQVFKPVFGLPTDAALVAETEVTLGKVLDVYEARLSQSKYLASDSFTLADLHHLPNIQALLGTPSKKLFDSRPHVSAWVASITG
RPAWGKVLALLPK
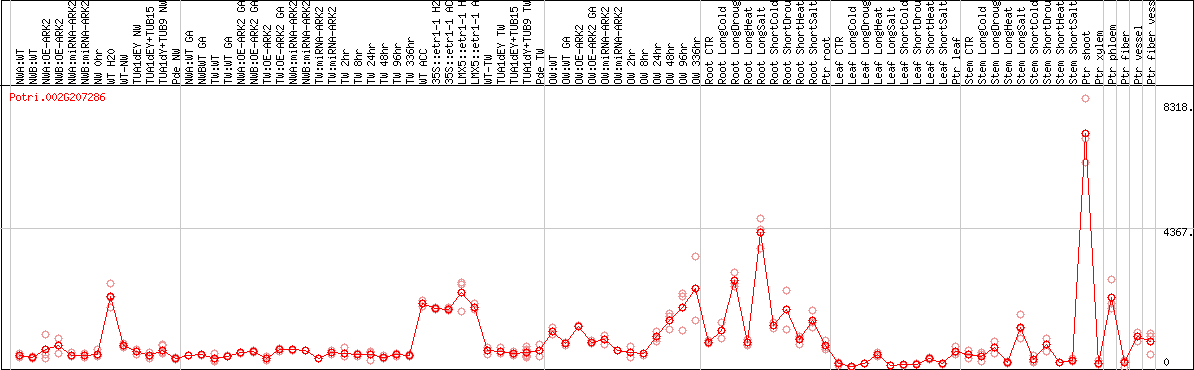
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G207286 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.