External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G07490 248 / 9e-86
AtCML3, AGD11
calmodulin-like 3, ARF-GAP domain 11 (.1)
AT1G05990 223 / 6e-76
RHS1 ,RHS2
ROOT HAIR SPECIFIC 1, EF hand calcium-binding protein family (.1)
AT2G43290 209 / 2e-69
MSS3
multicopy suppressors of snf4 deficiency in yeast 3, Calcium-binding EF-hand family protein (.1)
AT4G12860 206 / 4e-69
UNE14
unfertilized embryo sac 14, EF hand calcium-binding protein family (.1)
AT4G03290 194 / 1e-64
EF hand calcium-binding protein family (.1)
AT3G59440 194 / 9e-64
Calcium-binding EF-hand family protein (.1)
AT5G37770 115 / 3e-33
CML24, TCH2
TOUCH 2, CALMODULIN-LIKE 24, EF hand calcium-binding protein family (.1)
AT1G18210 111 / 1e-31
Calcium-binding EF-hand family protein (.1.2)
AT1G73630 108 / 8e-31
EF hand calcium-binding protein family (.1)
AT1G24620 105 / 3e-29
EF hand calcium-binding protein family (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009564
244 / 2e-84
AT3G07490 233 / 7e-80
calmodulin-like 3, ARF-GAP domain 11 (.1)
Lus10027701
154 / 1e-47
AT2G43290 153 / 2e-46
multicopy suppressors of snf4 deficiency in yeast 3, Calcium-binding EF-hand family protein (.1)
Lus10038088
153 / 4e-47
AT3G07490 150 / 7e-46
calmodulin-like 3, ARF-GAP domain 11 (.1)
Lus10006644
152 / 9e-47
AT3G07490 148 / 3e-45
calmodulin-like 3, ARF-GAP domain 11 (.1)
Lus10039986
136 / 4e-41
AT3G07490 132 / 3e-39
calmodulin-like 3, ARF-GAP domain 11 (.1)
Lus10020389
132 / 7e-41
AT3G07490 130 / 1e-40
calmodulin-like 3, ARF-GAP domain 11 (.1)
Lus10018012
121 / 2e-35
AT1G18210 195 / 2e-64
Calcium-binding EF-hand family protein (.1.2)
Lus10009059
111 / 2e-31
AT1G18210 197 / 3e-65
Calcium-binding EF-hand family protein (.1.2)
Lus10031345
110 / 4e-31
AT1G18210 196 / 1e-64
Calcium-binding EF-hand family protein (.1.2)
Lus10009127
107 / 4e-29
AT1G66400 147 / 2e-44
calmodulin like 23 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0220
EF_hand
PF13499
EF-hand_7
EF-hand domain pair
Representative CDS sequence
>Potri.002G239100.1 pacid=42779313 polypeptide=Potri.002G239100.1.p locus=Potri.002G239100 ID=Potri.002G239100.1.v4.1 annot-version=v4.1
ATGGATCCGGCAGAGCTACGTCGGGTATTTCAAATGTTTGATAAGAATGGAGATGGGCAGATCACAAAGAAGGAACTAAGTGACTCGCTAAAAAACTTGG
GTATCTATATACCCGATAAGGATTTGATCCAAATGATTGAGAAGATAGATGTCAATGGAGATGGTTACGTAGACATTGAAGAGTTCGGGGCATTGTATCA
AACGATAATGGACGAGAGGGACGAGGAGGAGGACATGAGAGAAGCATTCAACGTGTTCGATCAAAATGGAGATGGTTTTATCACAGTGGAGGAGTTGAAG
TCAGTTTTGTCATCCTTGGGGCTAAAACAGGGAAGAACTTTAGAGGATTGCAAGAGAATGATAAAAAAGGTGGATGTGGATGGAGATGGCATGGTTAACT
TCAGGGAGTTCAAGCAAATGATGAAAGGAGGTGGATTTGCTGCCCTAGGTTCAAGTTGA
AA sequence
>Potri.002G239100.1 pacid=42779313 polypeptide=Potri.002G239100.1.p locus=Potri.002G239100 ID=Potri.002G239100.1.v4.1 annot-version=v4.1
MDPAELRRVFQMFDKNGDGQITKKELSDSLKNLGIYIPDKDLIQMIEKIDVNGDGYVDIEEFGALYQTIMDERDEEEDMREAFNVFDQNGDGFITVEELK
SVLSSLGLKQGRTLEDCKRMIKKVDVDGDGMVNFREFKQMMKGGGFAALGSS
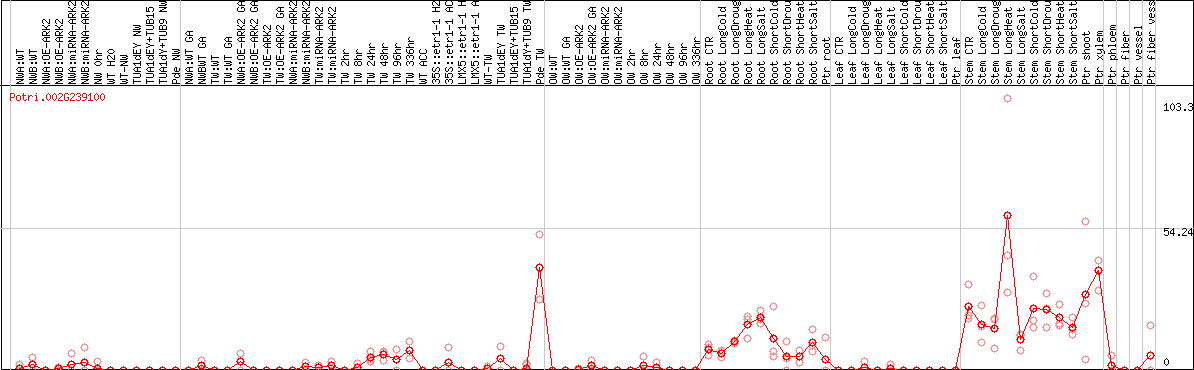
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G239100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.