External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G43280 130 / 2e-37
FAR1_related
Far-red impaired responsive (FAR1) family protein (.1)
AT3G07500 123 / 3e-34
FAR1_related
Far-red impaired responsive (FAR1) family protein (.1)
AT3G59470 117 / 6e-32
FAR1_related
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2)
AT4G12850 101 / 2e-26
FAR1_related
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2), Far-red impaired responsive (FAR1) family protein (.3)
AT4G38180 91 / 2e-20
FAR1_related
FRS5
FAR1-related sequence 5 (.1)
AT5G18960 74 / 1e-14
FAR1_related
FRS12
FAR1-related sequence 12 (.1)
AT3G06250 69 / 3e-13
FAR1_related
FRS7
FAR1-related sequence 7 (.1)
AT1G52520 62 / 6e-11
FAR1_related
FRS6
FAR1-related sequence 6 (.1)
AT2G27110 62 / 1e-10
FAR1_related
FRS3
FAR1-related sequence 3 (.1.2.3)
AT1G80010 61 / 3e-10
FAR1_related
FRS8
FAR1-related sequence 8 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G128800
294 / 9e-101
AT3G07500 124 / 8e-35
Far-red impaired responsive (FAR1) family protein (.1)
Potri.007G128900
227 / 1e-74
AT3G07500 154 / 2e-46
Far-red impaired responsive (FAR1) family protein (.1)
Potri.007G128700
194 / 2e-61
AT3G07500 138 / 2e-40
Far-red impaired responsive (FAR1) family protein (.1)
Potri.014G176500
159 / 3e-48
AT3G07500 275 / 2e-94
Far-red impaired responsive (FAR1) family protein (.1)
Potri.017G029400
128 / 3e-36
AT2G43280 319 / 4e-112
Far-red impaired responsive (FAR1) family protein (.1)
Potri.002G239400
119 / 4e-33
AT4G12850 207 / 1e-68
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2), Far-red impaired responsive (FAR1) family protein (.3)
Potri.007G129500
114 / 1e-30
AT3G59470 260 / 9e-88
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2)
Potri.017G029100
114 / 2e-30
AT3G59470 323 / 1e-112
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2)
Potri.014G176600
107 / 2e-28
AT4G12850 178 / 5e-57
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2), Far-red impaired responsive (FAR1) family protein (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026854
70 / 1e-13
AT3G59470 121 / 1e-32
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2)
Lus10020226
66 / 2e-12
AT3G59470 120 / 7e-33
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2)
Lus10026732
61 / 1e-11
AT3G07500 88 / 2e-22
Far-red impaired responsive (FAR1) family protein (.1)
Lus10025517
55 / 1e-09
AT2G43280 87 / 1e-22
Far-red impaired responsive (FAR1) family protein (.1)
Lus10022979
46 / 6e-06
AT4G38180 85 / 8e-20
FAR1-related sequence 5 (.1)
Lus10023134
39 / 0.0005
AT3G59470 92 / 1e-24
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0274
WRKY-GCM1
PF03101
FAR1
FAR1 DNA-binding domain
Representative CDS sequence
>Potri.002G239300.1 pacid=42779105 polypeptide=Potri.002G239300.1.p locus=Potri.002G239300 ID=Potri.002G239300.1.v4.1 annot-version=v4.1
ATGGACAGCACTTCTAAGCAAGCATTTGATTTGGACGAAACCGAAAGTTGTTTTGTTGTTGAAAATTATGATGAAAATGAGTTTGTGGACGATGAACTTT
CAGAAAATATTGAATTATGCATGGACGAGGTTGACAAGATTATAGAACAGTCTAGTGAAACTTTAGGTTCAATAAATGATGCTATGGAACCCTGCATTGG
CATGGAGTTCAAGTCAAGAGATGATGCACGAGAATTTTATATTGCTTATGGAAGGCGGACAGGATTTACTGTGCGTATCCATCATAACCGGCGTTCTCGA
ATAAATAATATGGTCATTGGTCAGGATTTTGTTTGTTCGAAAGAAGGATTTCGTGAAAAGAAATATGTATACAGGAAAGATAGAGTTCTTCGTCCACCAC
CTGTAACCCGTGAAGGCTGTCAAGCAATGTTAAGACTTGCTTTAAAAGATGGGATTACGTGGGTTGTTACCAAATTCATTGCAGAGCACAATCATGCATT
GATGTCTCCTAGTAAAGTACCATGGCGAGGATCTGCAAAAAGTTTGGTCAGTGAGGATGAAAAGGATCGAAGAATTCGAGAACTGACTATTGAGCTAAAC
AATGAGAAGCAACGATGTAAACGTAGGTGTGCCGCTTACCAGGAACAATTACGCATGGTGTTGTCATATGTTGAGGAACATACTATTCATTTGTCAAATA
AAGTTCAGGATATTGTTAACAATGTGAAAGAACTTGAAAATGATATGGAGGATTCGGATTGCAAATATGTCTAG
AA sequence
>Potri.002G239300.1 pacid=42779105 polypeptide=Potri.002G239300.1.p locus=Potri.002G239300 ID=Potri.002G239300.1.v4.1 annot-version=v4.1
MDSTSKQAFDLDETESCFVVENYDENEFVDDELSENIELCMDEVDKIIEQSSETLGSINDAMEPCIGMEFKSRDDAREFYIAYGRRTGFTVRIHHNRRSR
INNMVIGQDFVCSKEGFREKKYVYRKDRVLRPPPVTREGCQAMLRLALKDGITWVVTKFIAEHNHALMSPSKVPWRGSAKSLVSEDEKDRRIRELTIELN
NEKQRCKRRCAAYQEQLRMVLSYVEEHTIHLSNKVQDIVNNVKELENDMEDSDCKYV
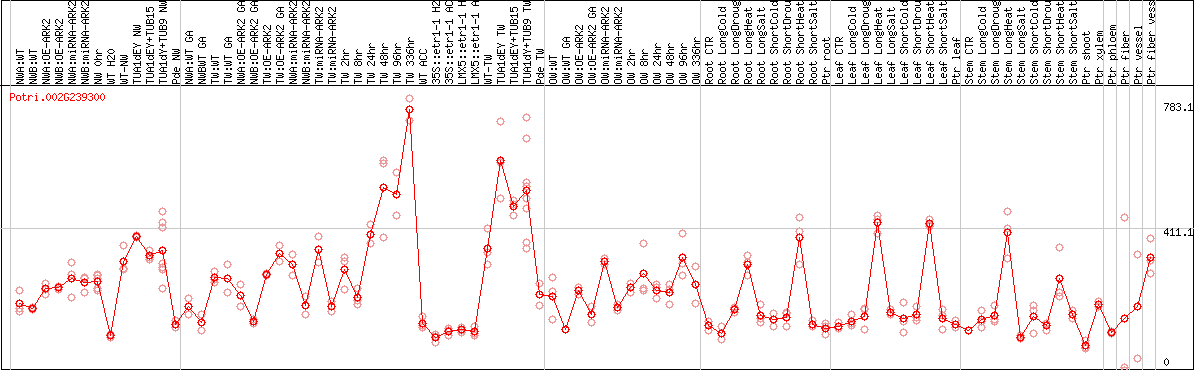
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G239300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.