Potri.002G243400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G243400.18 pacid=42778360 polypeptide=Potri.002G243400.18.p locus=Potri.002G243400 ID=Potri.002G243400.18.v4.1 annot-version=v4.1
ATGCATGGATGTGGTTCAGGATATGCTCTCCTAGTAAATGCTGAGGTTGATTCCATGGGTGGGGTTGTTGATGGTGGAGTTGGTATTGCTACCAAAACCT
CTCCGCGGCGAGCAGCAATTGAGAAGGCCCAGGTGGAACTTAGACAAGAGTATGATGTTCGTGAGGAACGGAGAAGGGAGCTAGAATTTCTTGAGAAAGG
CGGCAATCCTTTGGATTTTAAATTTGTAAATGCAACTTCGGTCAGTGTCCAGTCTACTTCACTCACAGATCATCACGTTGAACAGTTTGTTACTAGTGAA
GCAAAAGGCAGTTTTCCACTGACTGCATCACCTCATGGGGATTCTGTAGAAAGTAGTGGCAGACCAGGGGCTACTCCAGTTTGTGAGCCCAATAGTGCTG
ACAATTTTGATGGTGAAAATGAGTTACTTGAAGTTGAAAGGAAACGTAAACATCCTAGTAGGAGGAATAATGTCACTCAATCAGAGCAGTCTTCACAGAT
GGATGGAATTCACAATGCCAAGGAATCAGAAGATTCTGCTATTTTTCGGCCATATGCTCGAAGGAATAGGTCCAGACCAAATCGTGACGGCGCGCGATCA
AGTTCAACTGATATTGTTCAGAGTTCAGTTGGCCATGGATCCTCTTTACCGGCTCGTGGGGAAGCAAGGGATGTGAAAGGGTTGGTAACTGAAACAGACG
ATCATAAAGCCCAGATTATTACTTCCATCTCTAATCCAAAGTCCACCACTTCAAACGGTGACCTGTTTTTTCAGATTGACACTTCCAATACACAGTCAAA
CACGGAGCTTGATTGTGTGCAGGCTCTGAAAACAGTAGTCAACCTGCCTGATGATAGGTTGGATGTTACAGAGTCCATTGTCTTGAGGGATAACCAACAT
GACCAACCTTCAGAAGCTGATGCTGAGAAAGCACCCAATGATATTGCTTCCAGGGAATGCGATCATGGTGGAGGAAAAGAGCTGGTTATTTCAGCTGGTC
CTGAATGTCCCCCTTGTGCTGAATCAGCCAAGACTGAGAATGAAACTGGTCCTGCTCTGCTCAATGGCTTGGAAAAAGATGGAAATGAAGGTCAAAATGG
TAATGCAGCAATGGGAACGGAACGATTCAATTCAGAGTCCTCTTGCACCCAAGATAGCTTAAGCCTGGATGCAAACAATGGCTGTGATCCTTGTGATAAT
CGTAGAAATGATGACACCAATGAAATTCTTTTGAAAGAATCATCAGAATTTGAAGGAACACGAAGTTTACCATCGGGTAATATAGGAAATGAAAAGAAAG
AGACTAACTCGATCTCTGCAATCAATGATGGTTCTGTGCATGAAAACTACTCTGGCAATGATTCTACTGTCAAAAATGAGGAAGAAAGGAGAACAACTTT
TCACTCTCTGGTGAAGTGTACTAATCTTGAAGGAGTGGAACAAAATGACCATGTTGCCTCTGAGGCTGATACAAAGGCGGGCAACATGTTGGCTGATAGT
TCTAATTCCAACAGGGAAATAATTTACCCCAGTGGTCCCCAGGGCTCTTTGGATCCTCCTGTTCAAGAGCTTCCCCAGCCTATTTTGTTAGAGAAAAACT
CTTTTGTTGCCACTGATCCTCAATCTTGTTCGAATACACATGTTAAAGTGGTAGACAAGTCTCATGAAGATTCTATTTTGGAAGAGGCACGGGTTATAGA
GGCTAAGCGCAAGAGGATTGCAGAATTATCTGTTGCATCTGTGCATTCAGAGAATCGCCGGAGATCTCACTGGGATTTTGTCCTTGAAGAAATGGCATGG
CTGGCAAATGATGTTGCACAGGAGCGTCTCTGGAAGATGACTGCTGCTGCTCAAATATGTCGACGAATTGCTTTTACTTCTCGATTGAGAGTTGAAGAGC
AGAATCATCATTTGAAGCTAAAAAACGTCGCTTACAGTCTTGCTAAGGCTGTCATGCAGTTCTGGCATTCAGCAAAGGTGTATCTAAGTAACAATTGTCA
CAGTGTTGGTTCAAAGAATGGGAAGCATGAGGTGGGGATGTTTGTTGGGAATGAATTTTCTGTCAACAAGTTTGGAGATATTGATAAGGAGCAGGTGGCA
TGCAAAGAGTTAGAGAAACAAAATCGTGCCAAAAACATTGCGCATTCCATTCATGGATATGCTGTAAGGTTTTTGAAATATAACAGTTCTCCATTTCCTT
CCTTTCAAGCAGAAGCACCAGCAACTCCTGATAGAATAGCTGATTTGGGCATTGTGGACACTTCATGGGATGATCGCCTTACTGAAGAAAGCCTGTTCTA
CGCAGTTCCCTCAGGTGCCATGGCAATGTACAGATTGTCCATTGAATCTCATATTGCGCAGAGTGAGAAAACTCGCAGTAGCATGCAGGAGGAAGTCGAT
ACATCCATGTATGACACTCCAGCAGATTTTGGATATCATGATACTGCTGCATACGATGAGGAGGAAGGAGAAACAAGTGCTTACTATATGCATGGAGTCT
TTGAAGGTAGCAAGTCAGCAAAACATGACCAAAAGAAACGGAAAAGCTTAACAAAGTCACCCTCTGCAAGATCTTATGATCTGGGAACTGATTCACCTTA
TGGACACTGCACCACTGGGCCACAACAAAATGTATTGATGGGGAAAAGACCTGCTAGTAATCTTAATGCTGGCTCAATCCCAACAAAACGCATGCGTACT
GCTTCTAGACAGAGGTTTACAAGTCCTTTTACTGCTGGAACTGCTGGGGTTTTACTCCAGGCTCCAGTGAAGACAGATGCATCGAGTGGAGATACCAATT
CTTTTCAGGATGATCAGAGTATTTTGCATGGTGGGTCACAAATTCAGAAAAGTGTGGAGGTTGAGTCCGCTGCGCATTTCGAAAGGCAGTTACCATATGA
CTACGCAGAGACATCAACAAAACCGAAGAAGAAGAAGAAGGCAAAGCACCTGGGTTCTGCTTATGAGCAGGGGTGGCAGTTGGATTCTACTGGTCATAAT
GAACAGAGGGATAATTTCAAAAAGAGATCGGAGAGTCATCATTTGGACTCTAATGGAACCAGTGGTTTATATGGGCAACATACCACAAAGAAGCCAAAGA
TATCTAAGCAATTGCTAGATAATACTTTTGACAACATGGTTCAAATGACCGGGTCCATCCCATCTCCAGCTGCTTCCCAGATGAGTAATATGTCAAACAC
AAATAGATTCATCAAATTGATTGGTGGCAGGGAACGAGGCAGAAAAAATAAATCAATGAAGATGTCTGTTGGGCAGCCAGGTTCTGGAAGTCCCTGGTCA
CTTTTTGAAGACCAGGCTCTTGTTGTTCTTGTGCATGACATGGGTCCTAATTGGGAGCTTATAAGCGATGCTATCAACAGCACTGTTCAATTCAAGTGTA
TATTCCGCAAGCCTAAAGAATGCAAAGATCGTCACAAAATCTTAATGGATAAGGGTGCTGGTGATGGGGCTGATAGTGCTGAGGATTCAGGATCTTCTCA
ATCCTACCCATCAACTTTGCCTGGCATTCCAAAGGGCAGTGCCAGACAGTTGTTTCAACATTTGCAAGGGCCGATGCAAGAGGATACCCTGAAGTCTCAT
TTTGAGAAAATCATCATTATTGGGAAGAAACATCATTACAAAAGGAGTCAGAATGAAAACCAGGATCCCAAGCAAATAGCAGCAACTCACAATTCTCATT
TTATTGCTCTTTCCCAAGTTTGCCCAAACACTAAATGGAGGTGTCCTAACGCCCCTTGA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G243400.18 pacid=42778360 polypeptide=Potri.002G243400.18.p locus=Potri.002G243400 ID=Potri.002G243400.18.v4.1 annot-version=v4.1
MHGCGSGYALLVNAEVDSMGGVVDGGVGIATKTSPRRAAIEKAQVELRQEYDVREERRRELEFLEKGGNPLDFKFVNATSVSVQSTSLTDHHVEQFVTSE
AKGSFPLTASPHGDSVESSGRPGATPVCEPNSADNFDGENELLEVERKRKHPSRRNNVTQSEQSSQMDGIHNAKESEDSAIFRPYARRNRSRPNRDGARS
SSTDIVQSSVGHGSSLPARGEARDVKGLVTETDDHKAQIITSISNPKSTTSNGDLFFQIDTSNTQSNTELDCVQALKTVVNLPDDRLDVTESIVLRDNQH
DQPSEADAEKAPNDIASRECDHGGGKELVISAGPECPPCAESAKTENETGPALLNGLEKDGNEGQNGNAAMGTERFNSESSCTQDSLSLDANNGCDPCDN
RRNDDTNEILLKESSEFEGTRSLPSGNIGNEKKETNSISAINDGSVHENYSGNDSTVKNEEERRTTFHSLVKCTNLEGVEQNDHVASEADTKAGNMLADS
SNSNREIIYPSGPQGSLDPPVQELPQPILLEKNSFVATDPQSCSNTHVKVVDKSHEDSILEEARVIEAKRKRIAELSVASVHSENRRRSHWDFVLEEMAW
LANDVAQERLWKMTAAAQICRRIAFTSRLRVEEQNHHLKLKNVAYSLAKAVMQFWHSAKVYLSNNCHSVGSKNGKHEVGMFVGNEFSVNKFGDIDKEQVA
CKELEKQNRAKNIAHSIHGYAVRFLKYNSSPFPSFQAEAPATPDRIADLGIVDTSWDDRLTEESLFYAVPSGAMAMYRLSIESHIAQSEKTRSSMQEEVD
TSMYDTPADFGYHDTAAYDEEEGETSAYYMHGVFEGSKSAKHDQKKRKSLTKSPSARSYDLGTDSPYGHCTTGPQQNVLMGKRPASNLNAGSIPTKRMRT
ASRQRFTSPFTAGTAGVLLQAPVKTDASSGDTNSFQDDQSILHGGSQIQKSVEVESAAHFERQLPYDYAETSTKPKKKKKAKHLGSAYEQGWQLDSTGHN
EQRDNFKKRSESHHLDSNGTSGLYGQHTTKKPKISKQLLDNTFDNMVQMTGSIPSPAASQMSNMSNTNRFIKLIGGRERGRKNKSMKMSVGQPGSGSPWS
LFEDQALVVLVHDMGPNWELISDAINSTVQFKCIFRKPKECKDRHKILMDKGAGDGADSAEDSGSSQSYPSTLPGIPKGSARQLFQHLQGPMQEDTLKSH
FEKIIIIGKKHHYKRSQNENQDPKQIAATHNSHFIALSQVCPNTKWRCPNAP
|
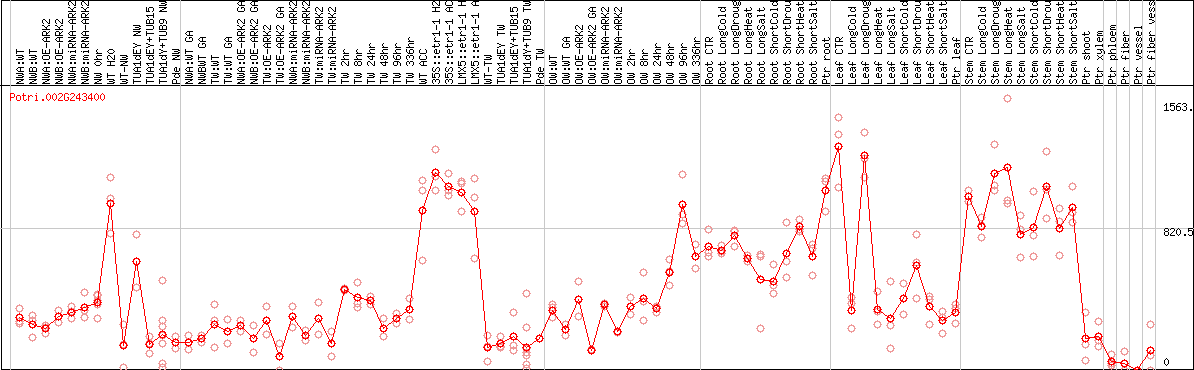
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G243400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
