Potri.002G254600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G254600.1 pacid=42779632 polypeptide=Potri.002G254600.1.p locus=Potri.002G254600 ID=Potri.002G254600.1.v4.1 annot-version=v4.1
ATGCATACCTTGGCGACATCGCTGCTTGTTCTTTGCTTGTCTAGCTTTGTTTTTACTTGTTTAGGACATCCATTACTTCATATGTATTTGTCTCCTTCTT
TGCAACCAAGCTGGCCATTATCTGTGAAAGACATTTTTGTGAGACATGGAATGTCGCTTGCAATGAAAGTGTCATTTGCTCGATCGCGGCCATCTCGAAA
TCACGTGGTTAAGCCTTCTTTGGGCCCTATCCTCTCACCTGCAACTTCTTTGGTGCATCAAGCTCTCCCCCCTGTTCCTAGCAGTGCACCCTTACCAAGA
CGTCATGGTGGTCATCATCATCGCCATGTGAAACCTGTAGTGACTGCTCCATTCCCTTCAGAAGAGCAAAGCTGTGATCAAATATGCACAGAGCCACTCA
CTGCATCTCTCTCTGGCTCACCTTGTGGCTGTGTTTATCCTATGAAAGTTAGACTTCTCCTAGATGTGGCTCCGTATGCTGTTTTTCCTGTAATGAGAGA
GCTAGAGAGTGAAGTTGCAGCAGGCACGTATCTGGAACAGAGTCAAGTGATAATCATGGGTGCAAGTGCTGACAGTCAGAATCAGGGGAAAACGGTCGTG
GATATCAACTTGGTTCCACTTGGAGAGAAGTTTGATAATACTACTGCAATACTTACATATGATAGATTTTGGAAAAACAAAATGCCTTTAAATATAACTC
TTTTTGGTAATTATGAAGTTATATACATTAGTTATCCAGGGATTCCTTCTTCACCACCATATCCAAACTACACGGGGAGTGGTCCGAGTGGAAGTGCAGG
AGATCTCCCCATTACTGCTAATTTTGCGAGCAAGACCCAGAGGATGAATCTTAGAACCATCACCACCATTGCTTTGTCAGCATTTGTGGTTTTAGTGGTT
TGCATTGGGGCAATTGCTGTTGTTGTGAAATGGAGGAAGTCTGGAAGACCATCAAGTGCTGTTGGTCCGGCATTTACATCCTCTATAAATAAAAGATCTG
GCATTGGGTCTTTCTTGTCAAGCAGCATTGCAAGCTCAACACCAATGTCCCTCATGTCCAACATGGCTACTTGTATGCTCTCCGTCAAAACATTTTCATT
CGCTGAGCTTGAGAAGGCAACAGACAAGTTCAGTTCAAAGAGGATTTTAGGTGAAGGAGGATTTGGACGTGTTTACCTCGGGAGTGTCGAAGATGGAACT
GAAGTTGCTTTTAAGCTGCTTACAAGGGACAATCAGAATGGAGACCGTGAATTTGTCGCTGAAGTTGAAATGTTAAGTCGATTGCATCATCGAAACCTTG
TGAAACTTATTGGCATATGTATTGAAGGGCGCACACGCTGCTTGGTGTATGAACTTATTCGGAATGGCAGTGTGGAATCGCACTTGCATGGTGTTGACAA
GAATAAGGGACCTCTTGACTGGGATGCACGGTTGAAGATTGCCCTTGGAGCAGCTAGGGGATTGGCCTATCTTCACGAAGATTCCAACCCTCGTGTAATT
CATCGAGATTTTAAGGCTAGTAATGTTTTACTAGAAGATGACTTCACCCCCAAGGTCTCTGATTTTGGCTTGGCGAGGGAAGCAACAGAAGGAAGTCATC
ATATATCTACTAGGGTCATGGGAACTTTTGGGTATGTTGCCCCTGAATATGCAATGACAGGACATTTACTTGTAAAGAGTGATGTCTACAGTTATGGGGT
TGTGCTGCTTGAGCTTCTGTCTGGAAGAAAACCTGTGGATATGTCCCAACCTCCGGGACAGGAAAATTTAGTGACTTGGGCACGCCCGCTGCTAACTACT
AGAGAAGGATTGGAGCAGTTGGTGGATCCTTCCTTGGCAGGAAGCTATGACTTTGATGACATGGCTAAAGTGGCAGCCATTGCTGCCATGTGTGTTCACT
CAGAGGTGACTAACAGGCCATTCATGGGTGAAGTTGTCCAGGCTCTGAAACTGATATACAATGACACAGATGAGACTTGTGGAGACTACTGCAGTCAAAA
GGAGTCCTCTGTTCTGGAATCTGACTTCAAATGTGATCTTGTTCCTTCGGACAGTAGCTGGTGGAATGCTGGCGGGACCTCCCCTCGGTTAACTTATGGG
CAGGCCTCATCCTTCATTACGATGGATTACAGTTCCGGTCCACTTGAAGAGATGGAAAGCAGACCGTTTTCAGCTTCAAGTTTAGCTGGGGATCGCCTGT
CTCTGCCAATTAGACACATGAACCGTTCCGGCCCCTTGAGAACTGTAAGAAGTAAACCAGCCTTCTATAGATTAAGGGGCAGCGTTAGCGAACACTGGGG
GCTCCTGTCAAGGCGTTTTCGGAATGACGGATATTGGGTATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G254600.1 pacid=42779632 polypeptide=Potri.002G254600.1.p locus=Potri.002G254600 ID=Potri.002G254600.1.v4.1 annot-version=v4.1
MHTLATSLLVLCLSSFVFTCLGHPLLHMYLSPSLQPSWPLSVKDIFVRHGMSLAMKVSFARSRPSRNHVVKPSLGPILSPATSLVHQALPPVPSSAPLPR
RHGGHHHRHVKPVVTAPFPSEEQSCDQICTEPLTASLSGSPCGCVYPMKVRLLLDVAPYAVFPVMRELESEVAAGTYLEQSQVIIMGASADSQNQGKTVV
DINLVPLGEKFDNTTAILTYDRFWKNKMPLNITLFGNYEVIYISYPGIPSSPPYPNYTGSGPSGSAGDLPITANFASKTQRMNLRTITTIALSAFVVLVV
CIGAIAVVVKWRKSGRPSSAVGPAFTSSINKRSGIGSFLSSSIASSTPMSLMSNMATCMLSVKTFSFAELEKATDKFSSKRILGEGGFGRVYLGSVEDGT
EVAFKLLTRDNQNGDREFVAEVEMLSRLHHRNLVKLIGICIEGRTRCLVYELIRNGSVESHLHGVDKNKGPLDWDARLKIALGAARGLAYLHEDSNPRVI
HRDFKASNVLLEDDFTPKVSDFGLAREATEGSHHISTRVMGTFGYVAPEYAMTGHLLVKSDVYSYGVVLLELLSGRKPVDMSQPPGQENLVTWARPLLTT
REGLEQLVDPSLAGSYDFDDMAKVAAIAAMCVHSEVTNRPFMGEVVQALKLIYNDTDETCGDYCSQKESSVLESDFKCDLVPSDSSWWNAGGTSPRLTYG
QASSFITMDYSSGPLEEMESRPFSASSLAGDRLSLPIRHMNRSGPLRTVRSKPAFYRLRGSVSEHWGLLSRRFRNDGYWV
|
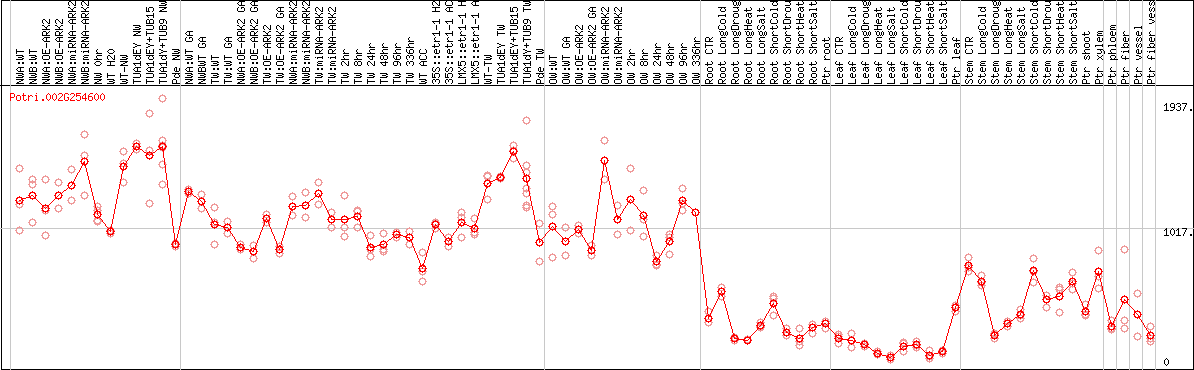
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.002G254600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
