Potri.002G257900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.002G257900.1 pacid=42778028 polypeptide=Potri.002G257900.1.p locus=Potri.002G257900 ID=Potri.002G257900.1.v4.1 annot-version=v4.1
ATGGCTGGCCTTGTCACGGGCAGTTCACAGACCTTGCATGCCAAAGATGAGCTGAGGCCTCCAACTCGCCAGTCTGCAACGTTGAAAAAATGTAGAGTTT
GTGGGGATGAGATTGGAGTTAAGGAAGATGGAGAGGTGTTTGTTGCTTGTCATGTGTGTGGCTTTCCTGTTTGTAGGCCTTGTTATGAGTATGAGAGGAG
TGAAGGCAACCAGTCCTGTCCTCAGTGCAACACTCGATATAAGCGTCACAAAGGTTGTCCTAGAGTTCCTGGAGATAATGACGATGAGGATGCCAATTTT
GATGATTTTGAAGACGAATTTCAGATTAAGCATCATGATCATGACGAATCCAATCAGAAAAATGTTTTTAGTCACACGGAAATTGAGCACTACAATGAAC
AGGAAATGCAACCCATTCGTCCGGCCTTTTCGTCAGCAGGAAGTGTTGCTGGTAAGGATCTTGAAGGCGAGAAAGAGGGTTATAGCAATGCAGAATGGCA
AGAGAGGGTGGAGAAATGGAAAGTTAGGCAAGAAAAGAGAGGTTTGGTGAGCAAAGATGATGGAGGAAATGATCAAGGAGAAGAAGATGAGTACCTCATG
GCTGAAGCCAGGCAACCACTATGGAGAAAAATCCCAATTCCCTCAAGCAGAATCAACCCGTATCGAATTGTCATTGTCCTTCGACTTATCATTCTTTGCT
TCTTTTTCCGTTTTCGGATCTTAACTCCAGCATACGATGCTTATGCATTGTGGCTTATCTCTGTAATATGTGAGGTATGGTTTGGCCTCTCCTGGATCCT
TGACCAGTTCCCAAAATGGAACCCCATTGAACGTGAAACTTATCTTGATCGCCTATCCATGAGGTTTGAGCGTGAGGGTGAGCCTAATCGTCTGGGCCCA
GTTGATGTGTTTGTGAGTACTGTGGATCCTCTCAAGGAACCACCAATCATAACTGCAAATACAGTCCTTTCAATCCTATCCGTTGATTATCCTGTCGACA
AGGTCAGTTGTTATGTATCAGATGATGGTGCATCCATGCTCCTTTTCGACTCCCTAGCAGAAACTGCTGAGTTTGCTAGAAGATGGGTTCCATTTTGCAA
GAAGCATAACATTGAGCCAAGGGCTCCTGAGTTCTACTTCACTCAGAAGATTGACTACTTGAAAGACAAAGTGCATCCCAACTTTGTGAAGGAGCGCAGA
GCTATGAAAAGAGAATATGAAGAATTCAAAGTAAGGATCAACGCATTGGTGTCAAAGGCCCAAAAGAAACCAGAAGAAGGATGGGTGATGCAGGATGGTA
CCCCATGGCCTGGAAACATCACACGTGATCATCCTGGAATGATTCAGGTATATCTAGGAAGTGAGGGTGCGCTCGACGTGGAAGGCAAGGAGCTTCCGAG
GCTTGTGTATGTTTCCCGTGAGAAACGACCTGGATATAACCACCACAAGAAAGCAGGTGCCATGAATGCTCTGATTCGAGTCTCAGCAGTGCTCACCAAT
GCACCTTTTATGTTGAATTTGGATTGTGACCATTACATCAATAACAGCAAGGCTGTAAGAGAAGCCATGTGCTTTTTGATGGATCCCCAACTTGGGAAGA
AGCTCTGCTATGTCCAGTTTCCGCAGAGGTTTGATGGTATTGATCGCCATGATAGATATGCTAATCGCAATGTTGTGTTCTTTGATATAAACATGAAAGG
TCTAGATGGGGTTCAAGGGCCAGTATATGTTGGTACTGGATGTGTCTTCAACAGGCAGTCCTTGTATGGCTATGATCCTCCAGTGTCTGAGAAGAGACCC
AAGATGACATGCGATTGCTGGCCTTCATGGTGTTGCTGTTGCTGTGGTGGTTCAAGGAAAAAGTCTAAGAAGAAAGGGCAAAGAAGTCTTCTTGGAGGAC
TATACCCCATGAAAAAGAAAATGATGGGGAAGAAGTACACAAGGAAAGCATCTGCACCAGTCTTTGATCTTGAAGAGATTGAAGAAGGGCTTGAAGGCTA
CGAAGAGTTGGAGAAATCATCACTCATGTCACAAAAAAGTTTCGAGAAACGGTTTGGGCAATCACCGGTATTTATTGCCTCTACCCTCATGGAAAATGGT
GGCCTGCCTGAAGGAACTAACTCTCAATCACACATTAAGGAAGCCATTCATGTTATAAGTTGCGGGTATGAAGAAAAAACCGAATGGGGTAAAGAGGTTG
GATGGATTTATGGTTCTGTTACAGAAGATATCCTGACAGGCTTCAAGATGCATTGTAGAGGGTGGAGGTCTGTCTACTGTTCTCCCCAGAGACCAGCTTT
TAAGGGATCTGCTCCCATTAATCTATCAGATAGGTTGCACCAAGTTCTGCGATGGGCACTAGGCTCTATTGAGATTTTCCTTAGTCATCACTGTCCTCTA
TGGTATGGCTATGGAGGAAAGTTGAAGTTGCTGGAGAGGCTTGCTTACATCAACACCATCGTTTACCCTTTCACCTCCATTCCCTTACTTGCCTACTGTA
CTATTCCAGCAGTCTGCCTTCTGACAGGAAAATTTATCATCCCTACTCTGAACAACCTTGCTAGCATATGGTTCCTAGCCCTTTTCATCTCAATCATAGC
AACATCTGTGTTAGAACTTCGATGGAGTGGAGTCAGCATCCAGGACTTGTGGCGTAATGAGCAATTTTGGGTTATCGGCGGTGTCTCAGCTCATCTTTTT
GCTGTCTTCCAAGGCCTCCTCAAGGTCCTTGGAGGAGTAGACACTAACTTCACTGTTACATCAAAATCAGCAGACGATGCCGAGTTTGGAGAGCTGTACC
TCTTCAAATGGACCACCCTCCTCATCCCACCAACCACTCTAATCATCTTGAATATGGTTGGAGTTGTAGCAGGAGTATCCGATGCAATAAACAACGGATA
TGGATCATGGGGTCCTTTATTTGGGAAGCTATTTTTCGCTTTCTGGGTCATTGTCCATCTCTATCCCTTCCTCAAAGGTCTGATGGGAAGGCAAAACAGG
ACTCCTACAATTGTTGTCCTCTGGTCTATACTTCTTGCATCTATTTTCTCATTGATTTGGGTTAGAATTGATCCCTTCTTGCCCAAGCAAACTGGCCCAA
TTCTCAAACAATGTGGGGTGGAGTGCTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.002G257900.1 pacid=42778028 polypeptide=Potri.002G257900.1.p locus=Potri.002G257900 ID=Potri.002G257900.1.v4.1 annot-version=v4.1
MAGLVTGSSQTLHAKDELRPPTRQSATLKKCRVCGDEIGVKEDGEVFVACHVCGFPVCRPCYEYERSEGNQSCPQCNTRYKRHKGCPRVPGDNDDEDANF
DDFEDEFQIKHHDHDESNQKNVFSHTEIEHYNEQEMQPIRPAFSSAGSVAGKDLEGEKEGYSNAEWQERVEKWKVRQEKRGLVSKDDGGNDQGEEDEYLM
AEARQPLWRKIPIPSSRINPYRIVIVLRLIILCFFFRFRILTPAYDAYALWLISVICEVWFGLSWILDQFPKWNPIERETYLDRLSMRFEREGEPNRLGP
VDVFVSTVDPLKEPPIITANTVLSILSVDYPVDKVSCYVSDDGASMLLFDSLAETAEFARRWVPFCKKHNIEPRAPEFYFTQKIDYLKDKVHPNFVKERR
AMKREYEEFKVRINALVSKAQKKPEEGWVMQDGTPWPGNITRDHPGMIQVYLGSEGALDVEGKELPRLVYVSREKRPGYNHHKKAGAMNALIRVSAVLTN
APFMLNLDCDHYINNSKAVREAMCFLMDPQLGKKLCYVQFPQRFDGIDRHDRYANRNVVFFDINMKGLDGVQGPVYVGTGCVFNRQSLYGYDPPVSEKRP
KMTCDCWPSWCCCCCGGSRKKSKKKGQRSLLGGLYPMKKKMMGKKYTRKASAPVFDLEEIEEGLEGYEELEKSSLMSQKSFEKRFGQSPVFIASTLMENG
GLPEGTNSQSHIKEAIHVISCGYEEKTEWGKEVGWIYGSVTEDILTGFKMHCRGWRSVYCSPQRPAFKGSAPINLSDRLHQVLRWALGSIEIFLSHHCPL
WYGYGGKLKLLERLAYINTIVYPFTSIPLLAYCTIPAVCLLTGKFIIPTLNNLASIWFLALFISIIATSVLELRWSGVSIQDLWRNEQFWVIGGVSAHLF
AVFQGLLKVLGGVDTNFTVTSKSADDAEFGELYLFKWTTLLIPPTTLIILNMVGVVAGVSDAINNGYGSWGPLFGKLFFAFWVIVHLYPFLKGLMGRQNR
TPTIVVLWSILLASIFSLIWVRIDPFLPKQTGPILKQCGVEC
|
DESeq2's median of ratios [POPLAR]
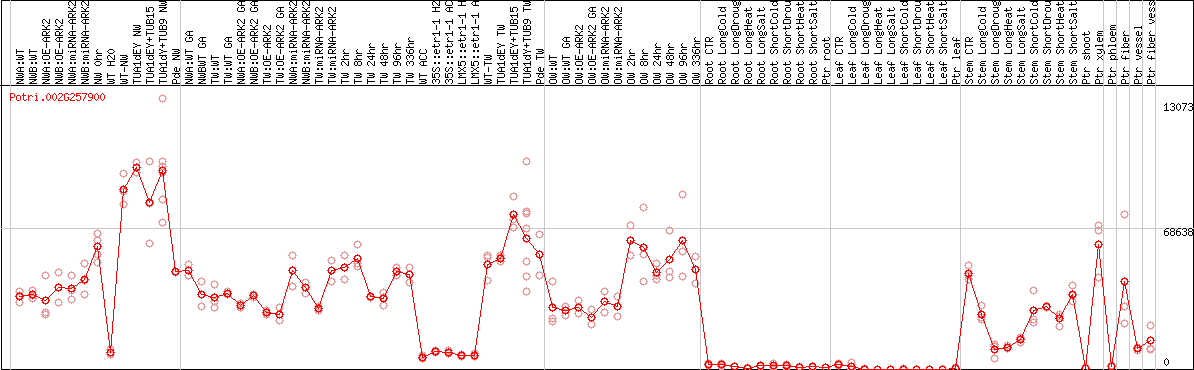
Coexpressed genes
Potri.002G257900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
