Potri.003G018900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.003G018900.1 pacid=42784629 polypeptide=Potri.003G018900.1.p locus=Potri.003G018900 ID=Potri.003G018900.1.v4.1 annot-version=v4.1
ATGGGGACCCGTTTTAGATCCGGTTCTTGTGTTCTTGTCATTCTAGTTTGTGTTGTAGGATTTTGTGGGTTTGGTTCGGTTTCTTGTGGGGAGAATCGAA
GGCCCAAAAACGTTCAGGTTGCTGTTCGGGCTAAATGGGAGGGCACTCCAATTTTATTGGAAGCTGGGGAATTACTATCTAAGGAAAGGAAAGACATTTA
TTGGGAGTTCATTGATAGCTGGCTTCATTCAAAGAAGGAAGATAATGATTCTTATACCGCTAAAGATTGCCTCAAAAAAATTATGAAACATGGACATGCT
CTTTTAAGTGACACATTGGCCTCACTATTTGATTTTTCTCTTATTCTAAGATCTGCATCCCCAAGATTGGTGCTTTACAGACAGCTAGCAGAGGAGTCTC
TTTCTTCTTTTCCGCTTTTGGATGATAGTTTCTCAAACAATGCTAGTGGGGGCCTTGCCAAAACAAATGATACCAATGAAATGAAAAGGTCAGATCCCTT
GCTTGTGGGAAGAAATCCTGAAATTCCTGGTGGGAAATGCTGTTGGGTGGATACTGGTGCGGCATTGTTCTATGATGTTGCGGATTTGCTATTGTGGCTT
CATAGCCCTACTGGAATGGCAGAGGACTCGTTTCAGCAACCCGAACTATTTGATTTTGATCATGTTCATTTTGAATCACTTTCTGGAAGTCCAGTTACTA
TTCTGTATGGAGCTCTTGGAACTGATTGCTTTAAGGAGTTTCATTCCGCCTTGGTGGAAGCTGCCAAACAGGGGAAGGTAAAATATGTAGTTAGACCTGT
TTTACCTTCTGGCTGTGAATCAAAAGTTGGGCGGTGTGTTGCTGTTGGGGCGAGTGACTCCTTAAATTTAGGTGGTTATGGTGTAGAACTTGCTCTGAAG
AATATGGAATATAAGGCCATGGATGATAGTGCAATAAAGAAAGGTGTCACTTTGGAGGATCCTCGGACTGAAGATCTTAGCCAAGAAGTTAGAGGGTTCA
TTTTTTCCAAAATACTGGAACGTAAACCTGAGCTTACATCAGAGATTATGGCTTTCCGAGACTATTTATTGTCGTCAACAATATCTGATACACTTGATGT
TTGGGAACTAAAGGATCTAGGACATCAAACAGCACAAAGAATAGTTCATGCCTCTGACCCTCTACAATCAATGCAGGAAATCAACCAAAATTTTCCTAGT
GTAGTTTCTTCTTTGTCTCGCATGAAGCTCAAGGATTCTGTTAAAGATGAGATAACTGCAAACCAGCGAATGATCCCTCCCGGCAAGTCCTTAATGGCTC
TGAATGGTGCTTTAATCAATATTGAGGATATTGACCTTTATCTGTTGGTAGACATGGTCCAGCAGGAATTATCATTGGCCGATCAATTTTCTAAATTGAA
GGTTCCTCACAGCACAATAAGGAAACTTCTTTCGACAGCGTCTCCTCCTGAATCCAGTATGATCCGTGTTGATTTTCGTTCAAGTCATGTTCATTATCTT
AATAACTTAGAGGAGGATGCCATGTATAAGCGGTGGAGGAACAATATAAATGAGATTTTGATGCCTGTCTTCCCTGGACAGCTACGTTACATCCGCAAGA
ACCTGTTTCATGCTGTTTATGTTCTTGATCCAGCCACTAGTTGCGGTCTTGAGTCTGTCGACATGATTTTGTCTTTGTATGAGAACAACTTCCCAATGAG
ATTTGGATTGATTCTATATTCTTCAAAGTTTATCAAGAAAGCAACTAGTCGTGGCCTACATTTATCTGCTGAGGAGAATGATGGTGAAATTGAGGAGGAT
ATATCTAGTTTGATCATACGTCTTTTCATATATATTAAGGAAAGCTACGGGACACCAACAGCTTTCCAGTTTCTGAGCAATGTAAACAGATTACGCATGG
AGTCAGATTCTGAAGATGATGTTCCTGAAACACACCATGTGGACGGAGCATTTGTGGATACCATATTACCAAAGGTAAAAACCCCTCCTCAAGATATATT
GCTAAAACTGGCAAAGGAGCAAACCTATAAGGAACTGTCACAAGAAAGCTCCATGTTCGTGTTTAAGCTGGGTTTGAACAAGTTGCAGTGTTGCCTCTTA
ATGAACGGTCTTGTCTTTGATTCTAGTGAGGAGGTTCTTATGAACGCCATGAATGATGAGCTTCCCAGAATACAAGAGCAAGTTTATTATGGGCAAATAA
ATTCTCACACTGATGTGCTGGACAAATTCCTGTCAGAAAGTGGTATTGGCCGCTACAATCCACAGATCATTGCTGAAGGCAAGGCCAAACCAAGGTTTAT
TTCTCTGACTTCGGGAGTTCTTGGAGGGAAATCTGTGGTGAATGATATTAACTTCTTGCATTCCCCTGGAACTGTAGATGATGTGAAACCTGTAACTCAT
CTCCTTGCTGTTGACATCACATCAAAGAAGGGGATAAACTTGCTTCATGAAGGAATACGCTATCTGATAGAAGGGTCTAAGGGTGCTCGCTTGGGTGTCC
TCTTTAGTTCCAGTCAAGATTCTGATTTACCTGGTCTCCTTCTTGTGAAGGTTTTTGAAATCACCACTGCCTCTTACAGTCATAAGAAAAATGTGTTGAA
CTTTTTGGAGCATTTATGTTCATTTTATGAGCAAAAGTACATCCAGGCATCATCTGTAGCAGCTGAAAGCACTCAAACGTTTATAGACAAGGTGTATGAT
CTGGCTGATGCAAATGAGCTACCACAGAAGGCCTACAAGTCCATCCTTTCTGAATTTTCTGCTGACAAAGTGAAAAATCAATTGAATAAGGTGTCACAAT
TTTTTTACCTGCTTCTTGGGCTTGAATCTGGAGTGAATGCGGTCATTACCAATGGAAGGGTGATGTTTCCTGGTGATGAAGGCACCTTTTTGAGTCATGA
TTTACATCTTCTAGAGACGATGGAGTTCAAGCAAAGAGTAAAGCACATTGGAGAAATTATTGAAGAAGTGCAGTGGCAGGATGTAGACCCTGACATGCTA
ACAAGCAAATTTGTCAGCGATATTATAATGTATGTGTCATCTGCAATGGCAATGCGGGAAAGAAGTTCAGAAAGTGCACGCTTTGAGATTTTGAATGCTG
AACATAGTGCTGTTATTATAGATAATGAAAATTCCAGTGTTCATATTGATGCTGTTGTTGATCCTTTAAGCGCAGCTGGTCAGAAGGTTTCTTCACTTCT
TAGAGTTCTAAGGAAATATGTTCAACCAAGCATGAGAATCGTGCTAAACCCTATGAGTTCTCTTGTTGATCTTCCGCTGAAGAATTACTATAGATATGTG
GTCCCAACCATGGATGATTTCAGCAGCACTGATTTAACAGTAAATGGTCCTAAAGCATTTTTTGCAAATATGCCATTATCGAAGACTCTTACCATGAACC
TTGATGTTCCTGAGCCCTGGCTTGTTGAGCCTGTTATTGCTGTCCATGATTTGGATAATATTTTGCTTGAGAACCTGGGTGACACTAGAACATTGCAAGC
AGTTTTTGAACTTGAAGCACTTGTTCTTACTGGTCATTGTTCAGAGAAAGATCATGAACCCCCCCGAGGACTTCAGCTAATTCTTGGAACTAAGAGTAAT
CCTCACTTGGTTGATACACTTGTGATGGCCAATTTGGGTTATTGGCAAATGAAAGTCTCTCCTGGGGTGTGGTATCTTCAACTTGCTCCAGGTCGAAGCT
CTGAGCTGTATGCTTTTAGGGAAGGTGGTGATGGAAGTCAGGAAAAGCATCTGTCGAAACTTATCACTATTAATGATTTGCGAGGCAAGGTAGTTCATCT
AGAAGTGGTGAAGAAAAAGGGCATGGAGCATGAGAAACTGCTCATTTCAAGTGATGATGATAACAATTCACAAAGGAAGGGAACTCATGATAGCTGGAAC
TCCAATTTGTTTAAATGGGCTTCTGGCTTTATTGGCGGCGGCGGGCTTTCCAAGAAGAATGAGAGTGCTTTAATGGAGCATGAGAAGCGTGGACGGCATG
GAAAAACTATTAATATCTTCTCTATTGCTTCTGGGCATCTATATGAACGTTTCCTTAAAATCATGATTCTAAGTGTCTGGAAGAACACCCAGCGTCCAGT
GAAATTTTGGTTTATAAAGAACTATCTCTCTCCGCAATTTAAGGATGTGATTCCACATATGGCACAAGAGTATGGTTTTGAGTACGAACTGGTCACTTAT
AAATGGCCATCATGGTTGCATAAGCAGACAGAAAAGCAGCGAATTATTTGGGCATATAAGATTTTGTTCCTTGATGTTATCTTTCCACTTTCTTTGGAGA
GGGTTATATTTGTTGATGCTGATCAGGTTGTGCGAGCAGACATGGGAGAACTGTATGACATGGATATAAAAGGAAGACCTCTTGCTTATACACCATTCTG
TGATAACAACAGAGATATGGATGGATATCGATTTTGGAGCCAAGGATTCTGGAAAGAACATTTGCGGGGAAGACCATATCATATCAGTGCTTTGTACATT
GTTGATCTGGTTAAATTTCGAGAGACAGCTGCAGGAGATAATCTAAGGGTTTTCTATGAAACACTGAGCAAGGATCCAAATAGTCTTTCAAATTTGGATC
AGGATCTTCCGAACTATGCTCAGCATACAGTACCCATCTTTTCACTGCCTCAAGAATGGTTGTGGTGCGAATCATGGTGTGGTAATGCTACGAAATCCAG
AGCAAAAACTATTGATCTTTGCAACAATCCCATGACAAAGGAACCCAAGCTTCAGGGCGCAAAAAGAATAGTTTCGGAATGGGTTAATCTTGACTCTGAG
GCGAGACACTTCACTGCAAAAATCTTAGGGGATGAAGTAAATCCTCAAGAACTTGTTTCCCCCAACCAATCACAGGCAAAAATCTTAGGTGATGAAGTAA
ATCCTCAAGAACTTGTGTCCCCCAACCAATCACAGGACTATCAAACTGATAATTCACTGGAGGAAGACGCTGAATCCAAGTCTGAATTATGA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.003G018900.1 pacid=42784629 polypeptide=Potri.003G018900.1.p locus=Potri.003G018900 ID=Potri.003G018900.1.v4.1 annot-version=v4.1
MGTRFRSGSCVLVILVCVVGFCGFGSVSCGENRRPKNVQVAVRAKWEGTPILLEAGELLSKERKDIYWEFIDSWLHSKKEDNDSYTAKDCLKKIMKHGHA
LLSDTLASLFDFSLILRSASPRLVLYRQLAEESLSSFPLLDDSFSNNASGGLAKTNDTNEMKRSDPLLVGRNPEIPGGKCCWVDTGAALFYDVADLLLWL
HSPTGMAEDSFQQPELFDFDHVHFESLSGSPVTILYGALGTDCFKEFHSALVEAAKQGKVKYVVRPVLPSGCESKVGRCVAVGASDSLNLGGYGVELALK
NMEYKAMDDSAIKKGVTLEDPRTEDLSQEVRGFIFSKILERKPELTSEIMAFRDYLLSSTISDTLDVWELKDLGHQTAQRIVHASDPLQSMQEINQNFPS
VVSSLSRMKLKDSVKDEITANQRMIPPGKSLMALNGALINIEDIDLYLLVDMVQQELSLADQFSKLKVPHSTIRKLLSTASPPESSMIRVDFRSSHVHYL
NNLEEDAMYKRWRNNINEILMPVFPGQLRYIRKNLFHAVYVLDPATSCGLESVDMILSLYENNFPMRFGLILYSSKFIKKATSRGLHLSAEENDGEIEED
ISSLIIRLFIYIKESYGTPTAFQFLSNVNRLRMESDSEDDVPETHHVDGAFVDTILPKVKTPPQDILLKLAKEQTYKELSQESSMFVFKLGLNKLQCCLL
MNGLVFDSSEEVLMNAMNDELPRIQEQVYYGQINSHTDVLDKFLSESGIGRYNPQIIAEGKAKPRFISLTSGVLGGKSVVNDINFLHSPGTVDDVKPVTH
LLAVDITSKKGINLLHEGIRYLIEGSKGARLGVLFSSSQDSDLPGLLLVKVFEITTASYSHKKNVLNFLEHLCSFYEQKYIQASSVAAESTQTFIDKVYD
LADANELPQKAYKSILSEFSADKVKNQLNKVSQFFYLLLGLESGVNAVITNGRVMFPGDEGTFLSHDLHLLETMEFKQRVKHIGEIIEEVQWQDVDPDML
TSKFVSDIIMYVSSAMAMRERSSESARFEILNAEHSAVIIDNENSSVHIDAVVDPLSAAGQKVSSLLRVLRKYVQPSMRIVLNPMSSLVDLPLKNYYRYV
VPTMDDFSSTDLTVNGPKAFFANMPLSKTLTMNLDVPEPWLVEPVIAVHDLDNILLENLGDTRTLQAVFELEALVLTGHCSEKDHEPPRGLQLILGTKSN
PHLVDTLVMANLGYWQMKVSPGVWYLQLAPGRSSELYAFREGGDGSQEKHLSKLITINDLRGKVVHLEVVKKKGMEHEKLLISSDDDNNSQRKGTHDSWN
SNLFKWASGFIGGGGLSKKNESALMEHEKRGRHGKTINIFSIASGHLYERFLKIMILSVWKNTQRPVKFWFIKNYLSPQFKDVIPHMAQEYGFEYELVTY
KWPSWLHKQTEKQRIIWAYKILFLDVIFPLSLERVIFVDADQVVRADMGELYDMDIKGRPLAYTPFCDNNRDMDGYRFWSQGFWKEHLRGRPYHISALYI
VDLVKFRETAAGDNLRVFYETLSKDPNSLSNLDQDLPNYAQHTVPIFSLPQEWLWCESWCGNATKSRAKTIDLCNNPMTKEPKLQGAKRIVSEWVNLDSE
ARHFTAKILGDEVNPQELVSPNQSQAKILGDEVNPQELVSPNQSQDYQTDNSLEEDAESKSEL
|
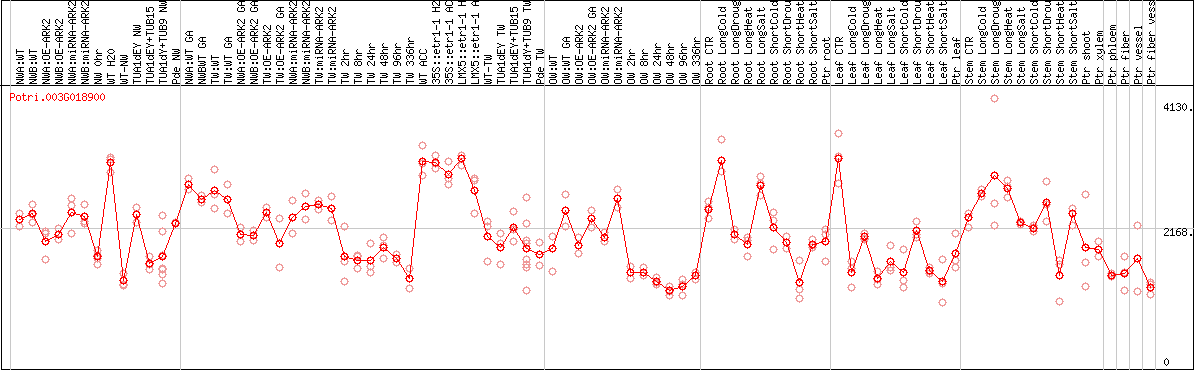
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G018900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
