Potri.003G031000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.003G031000.4 pacid=42787321 polypeptide=Potri.003G031000.4.p locus=Potri.003G031000 ID=Potri.003G031000.4.v4.1 annot-version=v4.1
ATGGCGCCAAGGTCAGGGAGGGGGAAATCAAACAAAGCTAGAGCAGAGAGGAAAAGGAAGGAGGAGAAATCTGTTCCTAGTGTTGTTGATGTCACCGTAA
TCACCCCATATGAATCGCAAGTTGTGCTCAAGGTGGTGCCACTACTCCTGGGCATCTCCACTGATAGGATACTTGATGTGAAGAAGTTATTGGCTGCAAG
TGTGCAAACATGTCATCTCACAAACTATTCTTTATCTCACGAGGTGAAAGGGCATGGGTTGCATGACAGAGTAGAAATCATCTCCTTGAAGCCCTGCCTG
CTCAAAATCATCGAAGAGGACTACACAGAGGAGTCACAAGCCGTAGCTCATGTACGGAGGCTGCTGGACATCGTAGCCTGCACAACCAGGTTCTCCAACA
AATCCAGGCGACCGTCACAGTCAATATCACAATCCAAACGGTCTAATAGTTCTAGGTCTCCACGGACCAGTACTCCTGCAACGCCACTGTCCGACGATGC
AGCATCCGAAACGACGTCGGTGTCGGCGGCGATGTCGGAGAGTATGGACATGGCGGCGATTCACCCGACGCCGAAGCTGTCTGAGTTTTATGACTTCTTC
TCCTTTTCTCACCTCCCGCCTCCTATTCTCGATTTGAGACGATGCAGTGAGGTTAAGGATGGAGAGGAAAGGTCACGACCGGGTGATTATTTTGAATTTC
AGGTTAAGATATGCAATGGAAAGCTAATAAAGGTGGTTGCTTCAGTAAAAGGCTTTTATGCAGTGGGAAAGCAATTTTCCCAAAGCCATTCCGTGGTGGA
TCTTCTTCAAAATCTCAGCCGAGCTTTTGCCAATGCATATGATTCCCTTATGAAGGCTTTTGTAGAGCATAATAAGTTCGGTAATCTTCCTTACGGATTT
CGTGCAAATACATGGCTTGTTCCTCCATCTGTTGCAGACTCTCCATCAAACTTTCCATCCCTTCCTGTAGAAGACGAGAGCTGGGGTGGCAATGGTGGAG
GGCAAGGAAGATATGGTGGATATGATCTTCGACCTTGGGCCACTGATTTCGCGATATTGGCTAGCCTACCATGCAAGACAGAGGAGGAGAGAGTGGTGCG
AGACCGGAAGGCCCTTTTGCTTCATAGTCAATTTGTTGATGTTTCAATTTTTAAAGCTGTTGGAGCGATACAAGGTGTTATTGATTCCAATCTTCAAGCA
AGAGATACAATTTCAGGCTCTTTCTTGCTTGAAGATCATGTAGGAGACTTGTCCATAGTTGTAGAACGTGATGCAGCAGATGCAAGCTTGAAGACTGTGG
TCAAAGTCAACGGCAATCACTTATCTGGTATCCCTGCTAAGGAAATTGCTCAAAGGAATTTGCTTAAAGGGGTAACCGCAGATGAGAGTGTAGTTGTTCA
TGATACCTCTTCATTGAGCACTGTCATTGTAAGACTTTGTGGATACACTGCAACAGTGAAAGTTGTTGGCAATGTGAAGAAGAAAAAGTTTGATGCTCAA
GATATTGAAATTGATGATCTACCAGATGGAGGAGCTAATGCTCTTAATATTAACAGTTTAAGGGTGCTGCTGCACAAGTGTTGCAGTGCTGAATCATCTC
TAGGACAGTCTTCTCACTCTACTTTGGAAGAGTTAGAAGCATCTAGATGTCTTATTCGGAAAGTAATTAAAGAGAGTTTAACCAAGCAAGAGGAGAAGCC
AATTGCTTCAGAAAGGTCTATTAGATGGGAACTTGGCTCGTGTTGGTTGCAACATCTGCAAAAGCATGAAGCCTCAAAAGATACCAATTCTAAAAGCCCT
GAGGATAATAGTGAGAACGAACAAGCTGTGAAAGGTCTTGGAAAGGAATTCAAATTTTTGAAGAAGAGGGATATGAAGCTTACTGTGACTTCCACACATG
ATAGAGAAGAAATTGAATCTGGCCTGTGCAGCCAAGCTATGGGAATTAATGCCGGACAACATAGCAATGATGAGTCCAACATTGGGTGCGAATTGAGGAG
ATTGGTTTCTGAAGAAGCCTTCTTGCGTCTGAAAGAATCTGGAACTGGCCTTCATTTAAAGTCAGCAGATGAGCTTCTTCAAACAGCATACAGATATTAT
GATGAAGTTGCATTGCCTAAGCTGGTAACGGATTTTGGCTCCCTTGAACTTTCGCCTGTTGATGGTCGCACGCTGACTGACTTCATGCATTTTAGGGGAC
TGCAGATGCGCTCTTTGGGACGGGTGGTTGAACTTGCAGAGAAGCTTCCTCACATACAATCACTTTGTGTTCATGAGATGGTTACTCGAGCTTTCAAGCA
CATACTTAAAGTGGTTATTGCATCTATCAACAATATATCAGACTTGTCTGCAGCCATAGCTTCATCCCTAAATTTCTTGCTTGGAAGTTGTGGGGTTGAA
GGCAGTGATCAAACTATGAAGGATGACCATGCTCTCAAATTGCAGTGGTTGCGAACATTTCTTTCCCAGAGATTTGGTTGGACACTGAAAGATGAATTTC
AGCATTTGAGAAAGCTTTCAATTCTTCGTGGACTTTGCCATAAGGTTGGATTGGAGTTGGTTCCAAGAGATTATGACATGGAGTGCTCAAACCCCTTTAG
GAAATGCGACATTATCAGTGTAGTTCCCGTGTGTAAGAATGTAGGATGCTCTTCAGCTGATGGACGGACTCTGTTGGAGTCGTCGAAGGTCGCTCTTGAT
AAAGGAAAGCTTGAGGATGCTGTTAATTATGGAACGAAGGCTCTAGCAAAGATGATAGCTGTATGTGGTCCCTATCATCGGACTACTGCCAGTGCTTACA
GTCTTCTAGCTGTTGTTCTCTACCACACAGGAGACTTTAATCAGGCTACTATATACCAGCAAAAGGCTTTGGATATCAATGAGAGGGAACTTGGTCTTGA
TCATCCAGATACAATGAAAAGCTATGGTGATCTTTCTGTGTTTTATTATCGTCTTCAACATGTTGAGTTAGCTTTGAAATATGTGAACCGTGCTTTATTC
CTTCTTCAATTTGCTTGTGGTCTGTCTCACCCTAATACCGCCGCTACATACATCAATGTGGCTATGATGGAAGAGGGTATGGGAAATGTTCATGTTGCTC
TCAGATACCTGCATGAAGCTCTAAAGTGCAATCAAAGATTGTTAGGGGCAGATCATATACAGACAGCTGCAAGCTACCATGCTATAGCCATTGCTCTTTC
TCTGATGGAGGCATATTCCTTAAGTGTTCAACATGAGCAAACGACACTCAAGATACTTCAGGCAAAACTTGGAACGGAGGATCTTCGTACTCAGGATGCT
GCTGCATGGCTTGAATATTTTGAATCAAAAGCCCTAGAGCAGCAAGAAGCAGCACGAAATGGAACTCCAAAGCCAGATGCATCCATCGCAAGCAAAGGTC
ACCTTAGTGTATCAGATCTCCTGGATTACATTAGTCCTGATCAAGACTCTAGAGGAAGTGATGCACTCAGGAAACAGCGTCGTGCAAAGGTACTCCAGGT
TAGTGATAAGTCCTATCAAGTCCATCAAGATGTAATGGTCAAAGATGGCTTGGGAAATGCAATGGTTATGACAGATGATGGCAACACTCAAGAACAAGGG
GTGGATATGATTCATAATGAAGAAGCAGAGGAAAATGATGATATCACCAAGTACAGGCCCACAGTTGCGGGGGAAGTTGTTGAAGAAACAACTTCAGATG
AAGGATGGCTAGAAGCCAATCCAAAAGGACGGTCATGGAAAGCTGCTGGCAGGAAATCTGGCCGTAGACGACCAGCTCTTGCAAAGCTAAACATAAATAC
TGCAGAATATTCGAGTAATAGAGAAAGAAGGTATAGGAGTCAAATCATTTCCCCGGCACAAAGAAAAACTCCAAGGACAATTACAATGGAGGTGTCACCT
GCGAAGCAATCAATTGAACTGCAGGCAAAAGCTACTGTCTCCAAACCTTTTTGTGCTCCGGCAAATCTCACTGCTATGGCATCAAAATCGCTCTCTTACA
AAGAAGTAGCTGTGGCACCCCCTGGTATGGCTTTGAAGCCATCTCAGGAGATAGTAGAAGAATCAAGTGGGGCAAAACCTGAAACTCAGATATGCGGCGT
TGTACCTGAGACATTTAAGGAGGAGGAAAGCAATGATATCCCTGTTATTGATAATAAACCAGGTCCTGATGAAGCAGAAGGAACTCATGAAAGTGAAACT
CAACCCGAAAAATCTGGCCCTGAAGTTGAGGAAATTTCTTCTTCTAATCAGGAAAAGTACATTGAGAAAAATGGAAGTAAGCTTTCAGCTGCAGCTGAAC
CATTCAATCCAGGAGTGTGCCCACTGGTTCACCCTTTAAACTCAGCTTCTGCACCAAGCATATATGATGCGACAGCTAGTCAAGGAATGCTTGTAGTCCC
AGCAGTGGCACCTCCACTTGCAAGAGTTCCTCGTGGGCCTAGGTCACCGCTGTATTATAGAACTGCTCAATCTTACCACATGAGACAAGGTCTCCTGAAA
TACCGAACTCATTTGGCAACACAACCTAGAAGCATGAACCCACATGCACCTGAGTTTGTTCCCAGTAGAGCTTGGCAAACAAATCCTGAAAATGGAGATT
CTGCTATTTCAACCGAGATGAAATCTTTGCTTGAGACAAGCAAGGCAAGAGAGGAAGAGGAGGACTTTGATGAGGAATCCGGTAATGAAGTTCAAGATTG
TTCTACAAAAAGGACAACCTCTGAGACCGAGAAGGCAGAACTTGCAAGGCAGATATTGCTCAGTTTCATTGTGAAGTCAGTACAGAATAATATTGATGGT
GGAAGCGAGACTCTAGGCAGTAAAAGGCTTGACAGTTCAGAAAGTTCTTCGGATGCAATAGCTAATGACACTGCAATTATAAAAATCCTCTATGGAAATG
AAGGAAAGACAAAATTGGTTACTCAGTCCAGCGACGGTGAACAGCTGAAAACCCCAGATGCTAATAAAAACAATCACGGTGACGGTGAAGGATTTATAGT
TGTGACAAAGCGGAGAAGAAACAAACAGCAGTTCACAAACGGGGTTGCTGGGTTGTACAACCAGCAATCACTCTGCGCACCTGTTCTTTGA
|
|||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.003G031000.4 pacid=42787321 polypeptide=Potri.003G031000.4.p locus=Potri.003G031000 ID=Potri.003G031000.4.v4.1 annot-version=v4.1
MAPRSGRGKSNKARAERKRKEEKSVPSVVDVTVITPYESQVVLKVVPLLLGISTDRILDVKKLLAASVQTCHLTNYSLSHEVKGHGLHDRVEIISLKPCL
LKIIEEDYTEESQAVAHVRRLLDIVACTTRFSNKSRRPSQSISQSKRSNSSRSPRTSTPATPLSDDAASETTSVSAAMSESMDMAAIHPTPKLSEFYDFF
SFSHLPPPILDLRRCSEVKDGEERSRPGDYFEFQVKICNGKLIKVVASVKGFYAVGKQFSQSHSVVDLLQNLSRAFANAYDSLMKAFVEHNKFGNLPYGF
RANTWLVPPSVADSPSNFPSLPVEDESWGGNGGGQGRYGGYDLRPWATDFAILASLPCKTEEERVVRDRKALLLHSQFVDVSIFKAVGAIQGVIDSNLQA
RDTISGSFLLEDHVGDLSIVVERDAADASLKTVVKVNGNHLSGIPAKEIAQRNLLKGVTADESVVVHDTSSLSTVIVRLCGYTATVKVVGNVKKKKFDAQ
DIEIDDLPDGGANALNINSLRVLLHKCCSAESSLGQSSHSTLEELEASRCLIRKVIKESLTKQEEKPIASERSIRWELGSCWLQHLQKHEASKDTNSKSP
EDNSENEQAVKGLGKEFKFLKKRDMKLTVTSTHDREEIESGLCSQAMGINAGQHSNDESNIGCELRRLVSEEAFLRLKESGTGLHLKSADELLQTAYRYY
DEVALPKLVTDFGSLELSPVDGRTLTDFMHFRGLQMRSLGRVVELAEKLPHIQSLCVHEMVTRAFKHILKVVIASINNISDLSAAIASSLNFLLGSCGVE
GSDQTMKDDHALKLQWLRTFLSQRFGWTLKDEFQHLRKLSILRGLCHKVGLELVPRDYDMECSNPFRKCDIISVVPVCKNVGCSSADGRTLLESSKVALD
KGKLEDAVNYGTKALAKMIAVCGPYHRTTASAYSLLAVVLYHTGDFNQATIYQQKALDINERELGLDHPDTMKSYGDLSVFYYRLQHVELALKYVNRALF
LLQFACGLSHPNTAATYINVAMMEEGMGNVHVALRYLHEALKCNQRLLGADHIQTAASYHAIAIALSLMEAYSLSVQHEQTTLKILQAKLGTEDLRTQDA
AAWLEYFESKALEQQEAARNGTPKPDASIASKGHLSVSDLLDYISPDQDSRGSDALRKQRRAKVLQVSDKSYQVHQDVMVKDGLGNAMVMTDDGNTQEQG
VDMIHNEEAEENDDITKYRPTVAGEVVEETTSDEGWLEANPKGRSWKAAGRKSGRRRPALAKLNINTAEYSSNRERRYRSQIISPAQRKTPRTITMEVSP
AKQSIELQAKATVSKPFCAPANLTAMASKSLSYKEVAVAPPGMALKPSQEIVEESSGAKPETQICGVVPETFKEEESNDIPVIDNKPGPDEAEGTHESET
QPEKSGPEVEEISSSNQEKYIEKNGSKLSAAAEPFNPGVCPLVHPLNSASAPSIYDATASQGMLVVPAVAPPLARVPRGPRSPLYYRTAQSYHMRQGLLK
YRTHLATQPRSMNPHAPEFVPSRAWQTNPENGDSAISTEMKSLLETSKAREEEEDFDEESGNEVQDCSTKRTTSETEKAELARQILLSFIVKSVQNNIDG
GSETLGSKRLDSSESSSDAIANDTAIIKILYGNEGKTKLVTQSSDGEQLKTPDANKNNHGDGEGFIVVTKRRRNKQQFTNGVAGLYNQQSLCAPVL
|
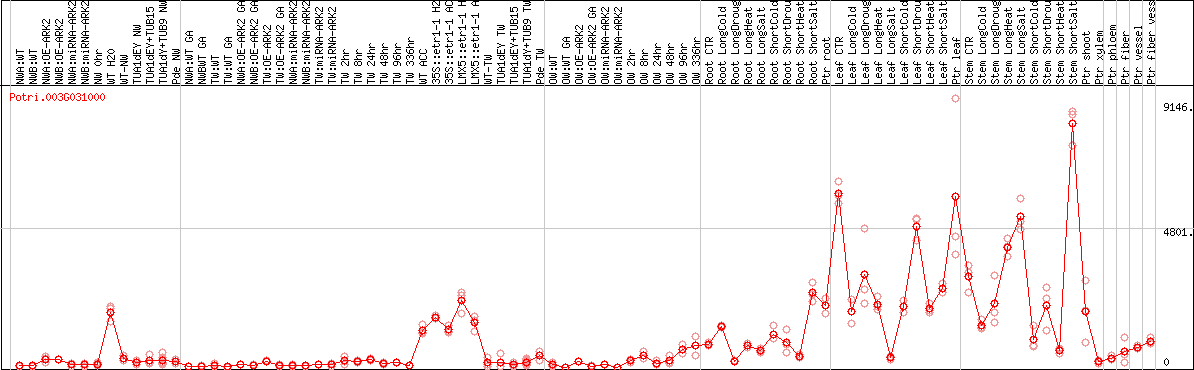
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G031000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
