External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G79870 444 / 5e-158
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
AT1G12550 275 / 2e-91
HPR3
hydroxypyruvate reductase 3, D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1)
AT2G45630 255 / 2e-83
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
AT4G34200 105 / 3e-25
EDA9
embryo sac development arrest 9, D-3-phosphoglycerate dehydrogenase (.1)
AT1G17745 101 / 1e-23
PGDH
D-3-phosphoglycerate dehydrogenase (.1.2)
AT3G19480 98 / 2e-22
D-3-phosphoglycerate dehydrogenase (.1)
AT1G68010 87 / 4e-19
ATHPR1, HPR
hydroxypyruvate reductase (.1.2)
AT1G72190 86 / 7e-19
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1)
AT5G28310 77 / 2e-16
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G14780 78 / 5e-16
FDH
formate dehydrogenase (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G183700
412 / 2e-145
AT1G79870 404 / 3e-142
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Potri.014G073400
271 / 3e-89
AT2G45630 405 / 2e-141
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Potri.002G151100
270 / 5e-89
AT2G45630 400 / 3e-140
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Potri.014G073500
265 / 2e-87
AT2G45630 377 / 4e-131
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Potri.002G151200
261 / 6e-86
AT2G45630 382 / 3e-133
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Potri.001G113250
256 / 7e-84
AT1G12550 340 / 9e-117
hydroxypyruvate reductase 3, D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1)
Potri.003G119000
236 / 7e-76
AT2G45630 327 / 2e-111
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Potri.013G046150
173 / 1e-53
AT1G79870 164 / 3e-50
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Potri.014G022800
103 / 1e-24
AT3G19480 860 / 0.0
D-3-phosphoglycerate dehydrogenase (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10035866
507 / 0
AT1G79870 466 / 6e-167
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Lus10025796
499 / 5e-180
AT1G79870 466 / 1e-166
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Lus10025795
423 / 1e-149
AT1G79870 438 / 6e-156
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Lus10035867
421 / 5e-149
AT1G79870 431 / 8e-153
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Lus10037552
408 / 8e-144
AT1G79870 410 / 6e-145
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Lus10036537
261 / 1e-85
AT2G45630 384 / 8e-134
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Lus10041393
255 / 1e-83
AT2G45630 392 / 7e-137
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
Lus10006708
244 / 3e-79
AT1G12550 358 / 4e-124
hydroxypyruvate reductase 3, D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1)
Lus10014133
229 / 2e-73
AT1G12550 340 / 1e-116
hydroxypyruvate reductase 3, D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1)
Lus10011442
125 / 3e-35
AT1G79870 117 / 1e-32
D-isomer specific 2-hydroxyacid dehydrogenase family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0325
Form_Glyc_dh
PF00389
2-Hacid_dh
D-isomer specific 2-hydroxyacid dehydrogenase, catalytic domain
CL0063
NADP_Rossmann
PF02826
2-Hacid_dh_C
D-isomer specific 2-hydroxyacid dehydrogenase, NAD binding domain
Representative CDS sequence
>Potri.003G052700.1 pacid=42786844 polypeptide=Potri.003G052700.1.p locus=Potri.003G052700 ID=Potri.003G052700.1.v4.1 annot-version=v4.1
ATGGAGTCCATAGGTGTCCTGATGACCTGCCCACCATTTGATCCTTACCTCGTCGAACAACTTGAGAAACGCTTCACTCTTTTCAAATTCCATTCCATTC
CTGACAAAGCCCACTTCCTCAATTCCAACAAAGCCTCCATCCGCGCGGTCGTTGGCAACGCCAGTGCAGGCGCAGACGCACAGCTCATCCACCAGTTGCC
CAACCTCGAAATCGTGTCCAGTTTCAGTGTTGGAATCGACAAGATTGACCTGGCCAAGTGTAGAGAGAGAGGTATCAGGGTCACTAATACCCCTGATGTT
TTGACCGATGATGTTGCTGATTTGGCTATTGGGTTGATGTTGGCTGTCTTGAGGAGGCTTTGTCCGAGTGATCGCTATGTTAGGAGTGGTCAGTGGAAAA
GGGGTGATTACAAGTTGACCACTAAGTTTACTGGGAAGTCAGTTGGCATTATTGGGTTGGGCAGGATTGGCCTGGCGATTGCCAAGAGAGCTGAAGCATT
TAGTTGCCCAATTAGTTACCACACTAGAGCAGAGAAATCAGATGTAAAATACAAGTACTATCCTAGTGTTGTTGAGTTGGCTGCCAACTGTCAAATTCTC
GTTGTTGCATGTGCATTAACTGAAGAAACTCGCCACATTATCAATCGTGAAGTCATCAATGCATTGGGTCCAAAGGGTGTCCTTATCAATATCGGTAGAG
GTCCTCATGTTGATGAACCTGAGCTGGTTTCTGCACTAGTTGAAGGCCGGTTAGGCGGTGCTGGGCTCGACGTGTTTCAAGATGAACCTAATGTACCCGA
GGAACTCTTTGGGCTTGAAAATGTAGTCCTCTTGCCTCATGTAGGGAGTGGCACCATGGAATCCAGGAAAGAAATGGCTGACCTTGTTGTTGGCAATTTG
GAGGCGCACTTCTTAAACAAACCATTGTTAACTCCCGTGCTTTGA
AA sequence
>Potri.003G052700.1 pacid=42786844 polypeptide=Potri.003G052700.1.p locus=Potri.003G052700 ID=Potri.003G052700.1.v4.1 annot-version=v4.1
MESIGVLMTCPPFDPYLVEQLEKRFTLFKFHSIPDKAHFLNSNKASIRAVVGNASAGADAQLIHQLPNLEIVSSFSVGIDKIDLAKCRERGIRVTNTPDV
LTDDVADLAIGLMLAVLRRLCPSDRYVRSGQWKRGDYKLTTKFTGKSVGIIGLGRIGLAIAKRAEAFSCPISYHTRAEKSDVKYKYYPSVVELAANCQIL
VVACALTEETRHIINREVINALGPKGVLINIGRGPHVDEPELVSALVEGRLGGAGLDVFQDEPNVPEELFGLENVVLLPHVGSGTMESRKEMADLVVGNL
EAHFLNKPLLTPVL
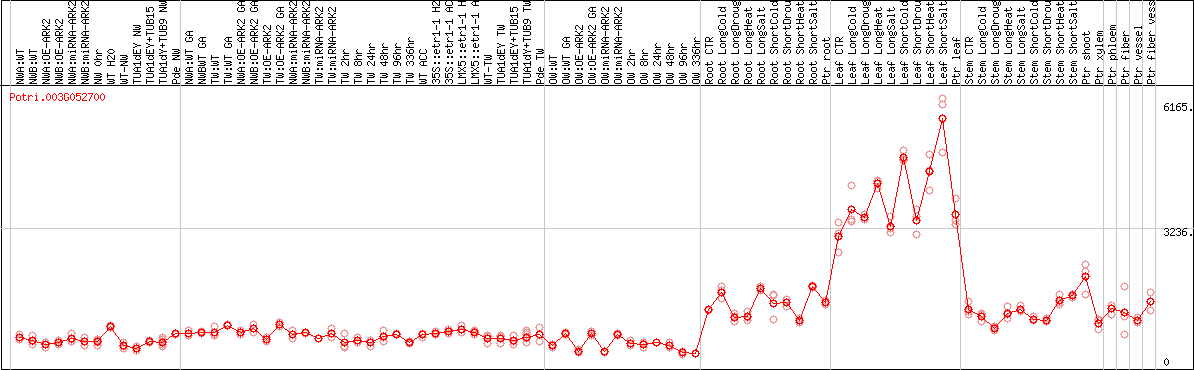
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G052700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.