External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G15660 201 / 6e-67
ATGRX4
A. THALIANA GLUTAREDOXIN 4, glutaredoxin 4 (.1.2)
AT3G54900 110 / 3e-31
ATGRXCP, CXIP1
GLUTAREDOXIN, CAX interacting protein 1 (.1)
AT4G04950 109 / 2e-28
AtGRXS17
Arabidopsis thaliana monothiol glutaredoxin 17, thioredoxin family protein (.1)
AT2G38270 96 / 3e-24
ATGRX2, CXIP2
GLUTAREDOXIN, CAX-interacting protein 2 (.1)
AT5G20500 53 / 4e-09
Glutaredoxin family protein (.1)
AT5G63030 46 / 8e-07
GRXC1
glutaredoxin C1, Thioredoxin superfamily protein (.1)
AT2G20270 45 / 4e-06
Thioredoxin superfamily protein (.1.2)
AT5G18600 41 / 4e-05
Thioredoxin superfamily protein (.1)
AT5G40370 41 / 6e-05
GRXC2
glutaredoxin C2, Glutaredoxin family protein (.1.2)
AT5G58530 41 / 0.0001
Glutaredoxin family protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008687
185 / 2e-57
AT3G15660 197 / 7e-62
A. THALIANA GLUTAREDOXIN 4, glutaredoxin 4 (.1.2)
Lus10018487
100 / 2e-26
AT4G04950 280 / 5e-93
Arabidopsis thaliana monothiol glutaredoxin 17, thioredoxin family protein (.1)
Lus10011187
98 / 6e-26
AT4G04950 322 / 8e-110
Arabidopsis thaliana monothiol glutaredoxin 17, thioredoxin family protein (.1)
Lus10041501
99 / 1e-25
AT3G54900 159 / 4e-49
GLUTAREDOXIN, CAX interacting protein 1 (.1)
Lus10012592
98 / 1e-25
AT3G54900 156 / 2e-48
GLUTAREDOXIN, CAX interacting protein 1 (.1)
Lus10026132
101 / 3e-25
AT3G14460 594 / 0.0
LRR and NB-ARC domains-containing disease resistance protein (.1)
Lus10002847
90 / 1e-21
AT2G38270 384 / 4e-135
GLUTAREDOXIN, CAX-interacting protein 2 (.1)
Lus10011188
48 / 5e-07
AT4G04950 297 / 1e-99
Arabidopsis thaliana monothiol glutaredoxin 17, thioredoxin family protein (.1)
Lus10017148
47 / 6e-07
AT5G20500 182 / 2e-60
Glutaredoxin family protein (.1)
Lus10022844
47 / 8e-07
AT4G28730 177 / 3e-57
glutaredoxin C5, Glutaredoxin family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0172
Thioredoxin
PF00462
Glutaredoxin
Glutaredoxin
Representative CDS sequence
>Potri.003G060600.2 pacid=42787462 polypeptide=Potri.003G060600.2.p locus=Potri.003G060600 ID=Potri.003G060600.2.v4.1 annot-version=v4.1
ATGGCAAGGTTACTGTCGAATACAATCTTGAAGGGTATTTCGCGTGCATCACAGTCTCCTAGAATTGTGCCTGCATCTTTTAACCACGTCAAGCTCAGAT
TCTCTACCACCATTCCCAATGATCCCGACAGTCATGCAGATTTTCAACCAAATAATAAGGCTGTGAATGAGAGTGGGAGTTGTAGCGGAATTAATATTAA
GGAACTTGTTGACAAGGATGTCAAGGAACACCCAATAGTAATTTACATGAAGGGATATCCTGACCTTCCTCAATGTGGATTTAGCGCTCTGGCGGTTAGA
GTATTGAAACAATACAATGTTCCGATAACTGCTAGAAATATATTGGAGTATCCTGACCTAAGAACTGGAGTAAAAGCATACAGCAATTGGCCTACATTTC
CCCAAATTTTTATCAAGGGAGAGTTTATTGGTGGCTCAGATATAATTATGAATATGCACCAGACTGGCGAACTGAAAGAAAAGCTCCAAGATATTTCAGG
AAAAGAGGAGTCTGAATAA
AA sequence
>Potri.003G060600.2 pacid=42787462 polypeptide=Potri.003G060600.2.p locus=Potri.003G060600 ID=Potri.003G060600.2.v4.1 annot-version=v4.1
MARLLSNTILKGISRASQSPRIVPASFNHVKLRFSTTIPNDPDSHADFQPNNKAVNESGSCSGINIKELVDKDVKEHPIVIYMKGYPDLPQCGFSALAVR
VLKQYNVPITARNILEYPDLRTGVKAYSNWPTFPQIFIKGEFIGGSDIIMNMHQTGELKEKLQDISGKEESE
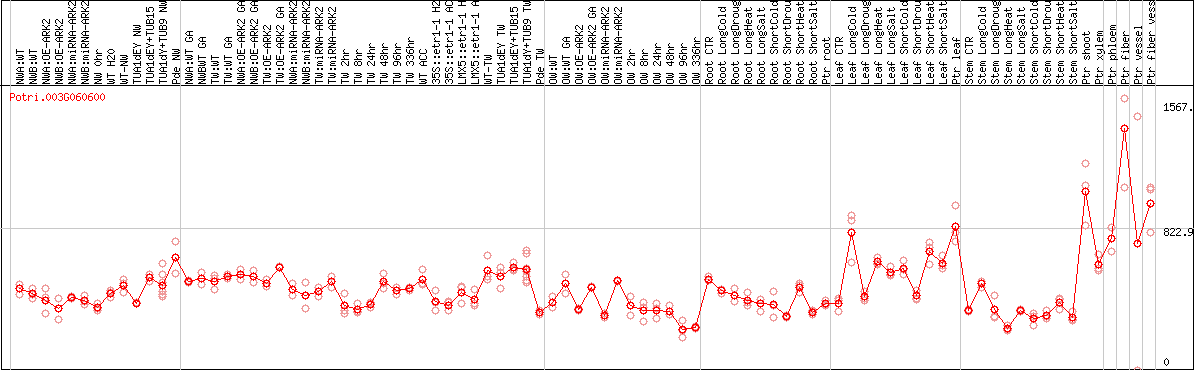
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G060600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.