Pt-SWI1.2 (Potri.003G106800) [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-SWI1.2 | ||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.003G106800.1 pacid=42786612 polypeptide=Potri.003G106800.1.p locus=Potri.003G106800 ID=Potri.003G106800.1.v4.1 annot-version=v4.1
ATGACTACCATGGAGCTGGTTGATGTTGCAGTGATAGACCACCCATCGGAAATAAAAAGGAGGCAGAACTCCGAGGACGCCGATAGAAGGCTTTTTTTGG
GCGGACATTGCCTGCATCACCCAACATTTACCACAGCACCACCATTTGACCCAGCTGAGCATATTAAAGTGAGCTCTTTTTATGAAGTTGATCACTCCAA
GCTGCCTCATAAATCCCCTGATCAACTCAACAAAACCCGGGTTGTGATGGTGAATGAAAAGACCAGGATGAGAGTCTCGCTGAGGTTTCCAAGCATCAAT
TCTCTAAGATGTTACTTCAATGAGATTGAAGCTATTAATTACAAGAAAGACATGAAAACGAAGAAGCAGCAGCTACCAGCATTCGACGAGAAATACATTA
TAGGATCAGAAGTTGCAGGGGAAGCTCTTTATAGGAGAATCTCTTCTCAAGAAATGGCAGACAAGAGTTACTCATGGAGTTTCTGGATGGTTAAACATCC
TTCGGTTTCACCTCGAAAAGTGTCATACCCACCTACAAGTACTCATGTTAATAAATTTGTTGGTGCAAGGAAGGTGTCTCTCATGTCTGAGCTCAACGGG
ACAGGCATGGTTAAGTGGGGTCAGCGCCGGCAGGTCAGGTTCTTGGCTAAACACGTAGAGGATAAACGTGAAATAGTGATTGCATCGAAGGATTTGATTA
AAAGCGAAGAAGAGAAAGACAGTGATGGTAGTGATGATGACACAGACGATGAGGACGAGGAGGAGGTCGATGTTAAGTTAGTAGTAAACAAGTCAAGTGA
AGCTAAAAGGAAATTACGTAAGAGAAAGTGTCAAGGTGGGTCTGGTATTAGCAAATTATCACCAAAAAAGAAAAGGCGTAAAATTGAAAAGAAGAACCAG
ATTGTGGTCTATAGGCAAAAGAAGAACAAACTCATCAAGAATTCTATTGACAGATGGTCTGCGGGGAGGTATAAATTGGCTGAGGAAAACATGTTAAAGG
TAATGAAAGAGCAAAATGCTGTGTTTCGACGCCCAATTTTAAGGCCAGAATTGAGAGCTGAGGCACGGAAGTTGATTGGGGATACTGGGCTGTTAGACCA
CTTGTTGAAGCATATGTCAGGGAAGGTGGCTCCGGGAGGAGAAGAGAGATTCAGAAGGAGGCATAACGCAGATGGAGCAATGGAGTATTGGCTGGAGAAG
GCTGATTTGGTTGATATCAGGAAAGAGGCTGGTGTGCAGGATCCTTATTGGACACCTCCACCTGGGTGGAAACCTGGTGATAATCCTAGTCAGGATCCAG
TTTGTGCTAGAGAGATCAAGGAACTCAGAGAAGAAATTGCTAAAATTAAAGGGGAGATGGAGGCAATGGTGTCTAAAAAACACGGGGAGGAATTAGCAAT
GGTGGCAGCACCGAATTATTCTCCTACAAGTCAGGACATGGAGCATGACAACTTCTTAATTCCACTGAAGGAAATGTACATTGATTTGGTGAATAAGAAG
GTAAAGATGGAGGAACAACTAAAGGAAATTTCAGAATCTTTGTATGGGATGAAGGAAGAAATGGAGAAGCTAAAAACCAGAGTGGAGAAATCAAACAGAG
CAGAATCAACTGAAAAGCCAGCTTTATTAATGGGCTCAACAGAGTCAATCACGCCAGCAGGAACTGGAAGAAAGGGGAAAGGAGTAATGCATCAGGAAAA
AGAAGCAACGGTTTTAGGGGAATCAGCACAAGAACAATGCAAGTCATCATCAGGAGGCATCATAGCACCAAGAACAGAATCACCAGCACCAACGGAGGAC
AGGGCAGCAAAGATAGAGAGGCTGAAAAGCGGGTTTAGAATATGCAAGCCCCAGGGAAGTTTCCTGTGGCCGGATATGACTACCTTAACCCCTCACCCTC
AGGTTGTGGTCCTACTAGAAGACCTCATTGCGGTACAAACACCTCCCTCAGTGTCCTCCACTACACCAAAACAATCTCACTTCCTCTTTGCTCCTCCATC
TCAAACCCATACACCCCACCGTACTTTCCCTGTGAAGCCATTAGCTGAGAGAAGGCCTGTCACCATTCCCCAATCCACAGCTGCCACGACTCCAACCAGC
TGTCCTCCCCTTGATCAAATGACTCACTCCCAGTATGAGAATAGCAGCATTTCCACTTCTACTACCATCACCACCACTACCAAAACCCCTCTCATCAACC
TTAATGAGCCACTGAATACCAATCAAACTGATGATTATGGATTGTTTTATGGGTCTCAGTCTCATGCTGAAGCCTCTCCTCACCCTGTCACTTACCAAAG
AAGACATCATCAAAATGTGACCACCAGTATTGCCATGCCAAGTTTGGGACCCACAAAGAAAGGGATGATGAGCCAATGGGAGGAAGGTGATCGGAGAACA
GGAATGATAAGGTACTGTGAGCAGTGTGAGCAGCAACAGGGATGCTCCTCTGCCTCTTCCATTGCATCTTCTTCCTTGCCAATGGGAAAGGGGACTTGGT
TGGCTCTGGCTACTTCTAAGGCTTCCGTGGAGCACAAATCTAAAAGGGGTTAA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.003G106800.1 pacid=42786612 polypeptide=Potri.003G106800.1.p locus=Potri.003G106800 ID=Potri.003G106800.1.v4.1 annot-version=v4.1
MTTMELVDVAVIDHPSEIKRRQNSEDADRRLFLGGHCLHHPTFTTAPPFDPAEHIKVSSFYEVDHSKLPHKSPDQLNKTRVVMVNEKTRMRVSLRFPSIN
SLRCYFNEIEAINYKKDMKTKKQQLPAFDEKYIIGSEVAGEALYRRISSQEMADKSYSWSFWMVKHPSVSPRKVSYPPTSTHVNKFVGARKVSLMSELNG
TGMVKWGQRRQVRFLAKHVEDKREIVIASKDLIKSEEEKDSDGSDDDTDDEDEEEVDVKLVVNKSSEAKRKLRKRKCQGGSGISKLSPKKKRRKIEKKNQ
IVVYRQKKNKLIKNSIDRWSAGRYKLAEENMLKVMKEQNAVFRRPILRPELRAEARKLIGDTGLLDHLLKHMSGKVAPGGEERFRRRHNADGAMEYWLEK
ADLVDIRKEAGVQDPYWTPPPGWKPGDNPSQDPVCAREIKELREEIAKIKGEMEAMVSKKHGEELAMVAAPNYSPTSQDMEHDNFLIPLKEMYIDLVNKK
VKMEEQLKEISESLYGMKEEMEKLKTRVEKSNRAESTEKPALLMGSTESITPAGTGRKGKGVMHQEKEATVLGESAQEQCKSSSGGIIAPRTESPAPTED
RAAKIERLKSGFRICKPQGSFLWPDMTTLTPHPQVVVLLEDLIAVQTPPSVSSTTPKQSHFLFAPPSQTHTPHRTFPVKPLAERRPVTIPQSTAATTPTS
CPPLDQMTHSQYENSSISTSTTITTTTKTPLINLNEPLNTNQTDDYGLFYGSQSHAEASPHPVTYQRRHHQNVTTSIAMPSLGPTKKGMMSQWEEGDRRT
GMIRYCEQCEQQQGCSSASSIASSSLPMGKGTWLALATSKASVEHKSKRG
|
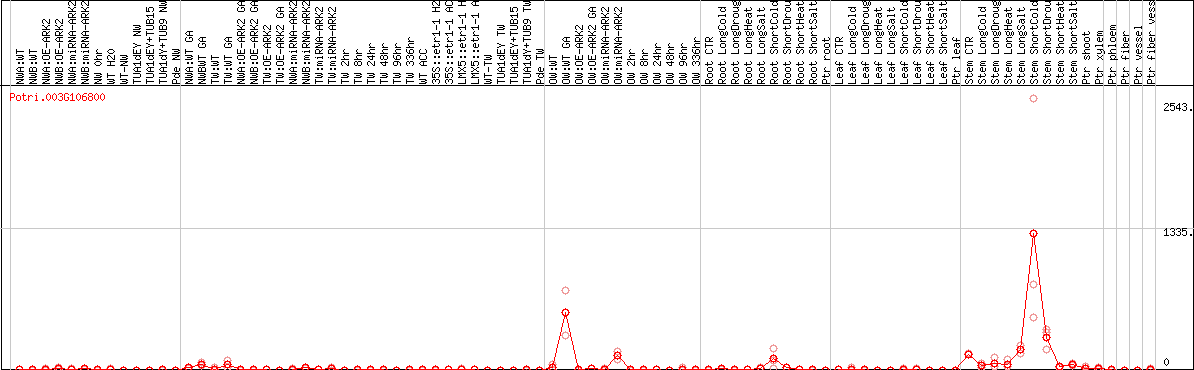
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G106800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
