Potri.003G107200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.003G107200.1 pacid=42786512 polypeptide=Potri.003G107200.1.p locus=Potri.003G107200 ID=Potri.003G107200.1.v4.1 annot-version=v4.1
ATGGATCAAACCCCAAACCCTCATTTTCAAACCCCACAAGACCAATCCAATCCACCACCACCACCACCACCACCACCACCGGAAACCCTAGATTTCATCC
CCGTTTGCCCTAACTCGCCCCCTCCTCCTCCAGCTGAAGAACCTCAATTTCCCGATCCCCAAAACCCACACAAAACCCTAATCTCCAATCCTCCCCAAAT
TGCCCCTGAAAATGGCCACAATCCCCAAACTACCACACCAAAACCCGAGATCCCCAAGTCCTTACTCTCCGAAAATGGCGTCGCTAACACAAACAGTGGT
GACAGAGACTGCTCTGGTGGAGAGGAGGAAACCACCAGCCGACGCCGCCGTCGCAGTCGTTGGGACCCACCGGCGGATGCTGGTGCCGATGGTAGTAACA
ATAATGATTCGGGTAGCGGCACGAGGAAGCGAAAATCGAGGTGGGCTGACGACGAGCCAAAGCCCGTGATTCAGTTACCTGATTTTATGAAAGATTTTAC
TGGAGGTATTGAATTTGATCCGGAAATTCAAGCTTTGAATGCTAGGCTATTGGAGATTAGCAGGATGTTGCAGTCTGGTTTGCCTCTTGATGATAGACCT
GAAGGAGCTAGATCTCCTTCACCTGAACCAATTTATGATAATATGGGAATTAGGATTAACACTAGAGAGTATAGGGCAAGAGAAAGATTGAACAAAGAAA
GACAAGAGATTATTTCGCAGATTATTAAGAGAAACCCGGCCTTTAAACCGCCAGCAGATTATAGGCCCCCCAAGCTTCAAAAGAAGCTTTATATACCTAT
GAAAGAGTATCCAGGCTATAATTTTATTGGTTTGATTATTGGGCCAAGAGGGAATACCCAGAAGAGAATGGAGAGGGAGACAGGTGGTAAAATTGTTATT
AGAGGTAAGGGGTCAGTGAAGGAAGGGAGATTGCAGCAGAAGAGGGATCTGAAACCAGATCCATCAGAGAATGAGGATTTGCATGTTTTGGTCGAGGCGG
AGACACAGGAGGCATTGGATGCTGCTGCAGGGATGGTTGAGAAGTTGTTGCAACCTGTGGATGAAGTCTTGAATGAGCATAAGAGGCAGCAGCTTAGGGA
ACTTGCTGCTTTGAATGGAACAATTAGGGATGAGGAGTATTGTAGGCTATGTGGTGAGCCAGGCCATAGGCAGTATGCATGCCCTTCGCGGACCTCTACG
TTTAAGAGTGATGTGCTTTGTAAGATTTGTGGCGATGGTGGACATCCAACCATTGATTGTCCCATGAAGGGAACAGCTGGGAAGAAAATGGATGATGAGT
ATCAGAACTTTTTGGCTGAGTTAGGTGGGACTATGCCTGAATCTGCAACTAAGCAAACTGCTACCTTAGCTTTGGAGTCAAGTGGTTCTGGAAACAATCC
TCCCTGGGCTGGTAGTAATACCGGGGGTCTTGGTAGTGCGAATCAAGCAGGGCTGGGAGCAAATGGGCTCAAACCTAAAGAATATGATGATACGAATTTG
TATATTGGTTACTTGCCACCCAATCTTGATGATGATGGTTTGATTGGTTTGTTTTCTTCTTTTGGTGAAATCGTGATGGCCAAGGTGATTAAGGATCGGA
TTACTGGATTGAGTAAAGGTTATGGTTTTGTGAAGTATTGTGATGTTCAAATGGCTAATAATGCAATCGCAAGCATGAATGGTTATCGCATTGATGGTCG
AACCATTGCTGTGAGAGTTGCTGGTAAGCCACCTCAGCCTACTGTCCCGCCAGGTCCTCCAACATCAACCATGCCCGCATACCCTATCCCAACTCAACCA
CTTGGTGGTGCCTATCCCTCGCAGCAGTTTACAGCAGGTGGTCCCCTTCCAAATGGTCCACCTACCAGCTATGTCGGGGCGCACGCCAGCTATAGAGGAA
CTCCGGTTCCATGGGGACCACCAGTTCCCTCTCCTTATGGTCCTTATGCTCCTCCTCCTCCTCCTCCTCCTCCTGGGTCAACCATGTACCCTCCCATTCC
TGGACAGCCTATACCTCCTTATGGTGTGCAATATCCTCTACCAGTGCAGCCAGTTCCCTCGGGCACACTGACTCAGACCGTGGCATGTAGTGAGGCTCAA
CAAAGCTACCCACCAGGAGTGCCATCTGAAAACAGCTTATCTGCTCCACTTGCAGCCTCAAATGTGTATGGGCACTCGATTGGTTATTCATCTTATTACA
GTGCAGTTCCTCCTCCTCCTCCCCCCCCAGCAACAGACCATTCACAGGGAATGGGTAATGTGCCTTGGGCCTCAAACTCGACCATGCCACCTCCACATTC
TTCTTCTGCAGAGAAAGCAAGATATGGTGCAGATGCAGAGTATGCGAAGTTTATGGCAGAGATGAAATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.003G107200.1 pacid=42786512 polypeptide=Potri.003G107200.1.p locus=Potri.003G107200 ID=Potri.003G107200.1.v4.1 annot-version=v4.1
MDQTPNPHFQTPQDQSNPPPPPPPPPPETLDFIPVCPNSPPPPPAEEPQFPDPQNPHKTLISNPPQIAPENGHNPQTTTPKPEIPKSLLSENGVANTNSG
DRDCSGGEEETTSRRRRRSRWDPPADAGADGSNNNDSGSGTRKRKSRWADDEPKPVIQLPDFMKDFTGGIEFDPEIQALNARLLEISRMLQSGLPLDDRP
EGARSPSPEPIYDNMGIRINTREYRARERLNKERQEIISQIIKRNPAFKPPADYRPPKLQKKLYIPMKEYPGYNFIGLIIGPRGNTQKRMERETGGKIVI
RGKGSVKEGRLQQKRDLKPDPSENEDLHVLVEAETQEALDAAAGMVEKLLQPVDEVLNEHKRQQLRELAALNGTIRDEEYCRLCGEPGHRQYACPSRTST
FKSDVLCKICGDGGHPTIDCPMKGTAGKKMDDEYQNFLAELGGTMPESATKQTATLALESSGSGNNPPWAGSNTGGLGSANQAGLGANGLKPKEYDDTNL
YIGYLPPNLDDDGLIGLFSSFGEIVMAKVIKDRITGLSKGYGFVKYCDVQMANNAIASMNGYRIDGRTIAVRVAGKPPQPTVPPGPPTSTMPAYPIPTQP
LGGAYPSQQFTAGGPLPNGPPTSYVGAHASYRGTPVPWGPPVPSPYGPYAPPPPPPPPGSTMYPPIPGQPIPPYGVQYPLPVQPVPSGTLTQTVACSEAQ
QSYPPGVPSENSLSAPLAASNVYGHSIGYSSYYSAVPPPPPPPATDHSQGMGNVPWASNSTMPPPHSSSAEKARYGADAEYAKFMAEMK
|
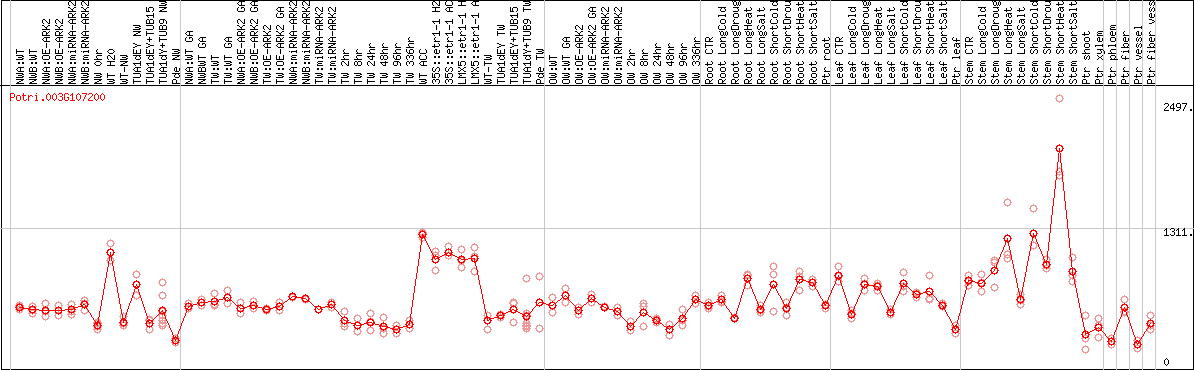
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G107200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
