External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G17260 477 / 3e-171
Lactate/malate dehydrogenase family protein (.1)
AT3G15020 59 / 4e-10
mMDH2
mitochondrial malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1.2)
AT1G53240 59 / 5e-10
mMDH1
mitochondrial malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G122400
517 / 0
AT4G17260 545 / 0.0
Lactate/malate dehydrogenase family protein (.1)
Potri.008G135920
337 / 1e-117
AT4G17260 372 / 1e-130
Lactate/malate dehydrogenase family protein (.1)
Potri.004G054200
61 / 2e-10
AT1G53240 520 / 0.0
mitochondrial malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
Potri.011G096300
55 / 1e-08
AT3G15020 541 / 0.0
mitochondrial malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1.2)
Potri.017G152000
52 / 9e-08
AT1G53240 277 / 8e-93
mitochondrial malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004350
476 / 5e-171
AT4G17260 550 / 0.0
Lactate/malate dehydrogenase family protein (.1)
Lus10002982
199 / 7e-65
AT4G17260 198 / 6e-64
Lactate/malate dehydrogenase family protein (.1)
Lus10006030
121 / 1e-31
AT4G17260 124 / 9e-32
Lactate/malate dehydrogenase family protein (.1)
Lus10002983
83 / 2e-19
AT4G17260 159 / 3e-48
Lactate/malate dehydrogenase family protein (.1)
Lus10028931
63 / 3e-12
AT4G17260 88 / 1e-21
Lactate/malate dehydrogenase family protein (.1)
Lus10013680
62 / 7e-11
AT1G53240 520 / 0.0
mitochondrial malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
Lus10017939
62 / 9e-11
AT1G53240 578 / 0.0
mitochondrial malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
Lus10038323
59 / 6e-10
AT3G15020 578 / 0.0
mitochondrial malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1.2)
Lus10021666
57 / 3e-09
AT3G47520 542 / 0.0
plastidic NAD-dependent malate dehydrogenase, malate dehydrogenase (.1)
Lus10000275
57 / 5e-09
AT3G47520 543 / 0.0
plastidic NAD-dependent malate dehydrogenase, malate dehydrogenase (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00056
Ldh_1_N
lactate/malate dehydrogenase, NAD binding domain
CL0341
LDH_C
PF02866
Ldh_1_C
lactate/malate dehydrogenase, alpha/beta C-terminal domain
Representative CDS sequence
>Potri.003G111201.1 pacid=42785859 polypeptide=Potri.003G111201.1.p locus=Potri.003G111201 ID=Potri.003G111201.1.v4.1 annot-version=v4.1
ATGCTTGATCTCCAACACGCTGCAGCTTTTTTACCCCGCACCAAAATTATTGCGTCCACTGATTACCTCGTCACTGTTGGATCTGATCTTTGTATTGTTA
CTGCTGGGGCTCGACAGATTGCGGGGGAGTCTAGGTTGAATTTGTTGCAGAGGAATGTTGCTTTGTTTCGGGGAATTATACCTCCGCTTGCTAAGTATTC
TCCCGGCACGATCTTGATGATTGTGTCGAATCCTGTTGATGTGTTGACTTATGTGGCGTGGAAGTTGTCGGGGTTTCCTTCGAATCGAGTTGTTGGATCT
GGTACGAATCTGGATAGTTCCAGGTTTCGGTTCTTGATTGCTGATCACCTTGATGTCAATGCCCAGGATGTGCAGGCCTCCATTATTGGCGAGCACGGGG
ATAGTTCTGTGGCATTATGGTCCAGCATAAGTGTTGGGGGAGTGCCCGTATTGAGCTTCTTAGAGAAGCAGCAAATTCCATATGAGAAGGAGACACTAGA
GGGGATTCACAAAGCAGTTGTAGACAGTGCGTATGAAGTGATTAGTCTCAAGGGTTACACATCTTGGGCGATCGGGTACTCAGCTGCTAACTTGGCTCGG
TCCATACTCAGAGACCAGAGGAAAATCCACCCCGTTTCAGTTCTTGCAAAGGGGTTTTATGGCATTGATGATGGCGATGTGTTCTTGAGCTTACCTGCAC
AGTTAGGTAGGGGAGGGGTTTTAGGGGTGACCAATGTGCATTTGACGGATGAGGAGGCACAAAGGCTTAGGAAGTCTGCTCAGACCATCTTGAAGGTGCA
AAGCCAGTTGGGACTTTGA
AA sequence
>Potri.003G111201.1 pacid=42785859 polypeptide=Potri.003G111201.1.p locus=Potri.003G111201 ID=Potri.003G111201.1.v4.1 annot-version=v4.1
MLDLQHAAAFLPRTKIIASTDYLVTVGSDLCIVTAGARQIAGESRLNLLQRNVALFRGIIPPLAKYSPGTILMIVSNPVDVLTYVAWKLSGFPSNRVVGS
GTNLDSSRFRFLIADHLDVNAQDVQASIIGEHGDSSVALWSSISVGGVPVLSFLEKQQIPYEKETLEGIHKAVVDSAYEVISLKGYTSWAIGYSAANLAR
SILRDQRKIHPVSVLAKGFYGIDDGDVFLSLPAQLGRGGVLGVTNVHLTDEEAQRLRKSAQTILKVQSQLGL
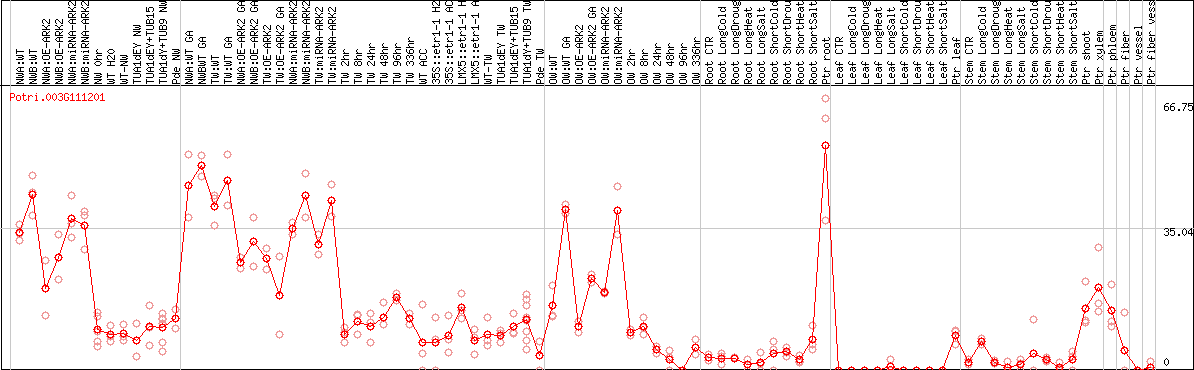
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G111201 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.