External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G62760 171 / 1e-52
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT5G62360 153 / 7e-47
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT5G62350 137 / 9e-41
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT1G62770 135 / 4e-40
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT4G25260 130 / 4e-38
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT3G47380 126 / 2e-36
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT1G14890 126 / 3e-36
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT2G01610 125 / 7e-36
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT4G25250 123 / 3e-35
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT5G51520 119 / 1e-33
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G119300
322 / 1e-113
AT1G62760 167 / 3e-51
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128300
197 / 4e-64
AT5G62360 192 / 1e-62
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128100
195 / 2e-63
AT5G62360 221 / 1e-73
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128200
194 / 5e-63
AT5G62360 172 / 1e-54
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128700
147 / 9e-45
AT5G62350 221 / 7e-74
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.012G127500
146 / 4e-44
AT5G62350 220 / 2e-73
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.012G127400
140 / 4e-42
AT4G25250 150 / 7e-46
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128400
140 / 7e-42
AT5G62360 134 / 2e-39
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128500
134 / 1e-39
AT4G25250 144 / 2e-43
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028910
187 / 4e-60
AT1G62760 169 / 1e-51
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10004327
178 / 2e-56
AT1G62760 164 / 8e-50
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10038915
151 / 5e-46
AT5G62360 174 / 5e-55
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031138
147 / 2e-44
AT5G62360 172 / 3e-54
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031133
146 / 5e-44
AT5G62350 211 / 8e-70
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10027199
144 / 4e-43
AT5G62360 169 / 3e-53
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031713
140 / 8e-42
AT5G62350 202 / 4e-66
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10032232
134 / 2e-39
AT1G62760 141 / 9e-41
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031711
134 / 2e-39
AT5G62360 145 / 4e-44
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10024595
134 / 4e-39
AT1G62760 141 / 6e-41
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04043
PMEI
Plant invertase/pectin methylesterase inhibitor
Representative CDS sequence
>Potri.003G113600.1 pacid=42786331 polypeptide=Potri.003G113600.1.p locus=Potri.003G113600 ID=Potri.003G113600.1.v4.1 annot-version=v4.1
ATGGAAAGCTCCTTTGTAGGCCTTGCTCTAGCGGCTATTCTCGTTATCCAATTCTCTACCCCAGTAAACTCTTCATGCTCAGCCACTAGACCCTCACCAA
GAAACATAAGATATATCAAAACATCATGTTATGATACCACACTCTATCCTAAATTATGCTACCACACCCTTGCAATCTATGCAAGCACAATCAAAACAAA
CCCTAAACTACTAGCTAATACTGCCCTTCATGTGAGCCTTAAAAGCACCAACTCGACATCAAGGTTGATGAAAAGGGCGTCCAAGACTCCTGGTCTGGAT
CCTAGAGTGCTTGCGGCCATGTTGGATTGTGTAGAGGAGGTTGGTGATGCAGTATATGAGCTTCAAAGGTCTATTGAAGAGATGGATCATGCTGGCGGGT
CAAATTTTTCTATGGTAATGAATGATGTGGTAACGTGGGTTAGTGCATCATTAACAGATGATGATACCTGCATGGATGGATTTGCTGAAGGGGCTGTTAA
TAAAAAAGTGAAGACCACTGTTAAGAGACATCTAGGAAGAATTGCACGTTTGACTAGTAATGCTTTGGCTCTTGTCAATAGATATGCTTCAAGTAAAGCT
AACTTACCTTAA
AA sequence
>Potri.003G113600.1 pacid=42786331 polypeptide=Potri.003G113600.1.p locus=Potri.003G113600 ID=Potri.003G113600.1.v4.1 annot-version=v4.1
MESSFVGLALAAILVIQFSTPVNSSCSATRPSPRNIRYIKTSCYDTTLYPKLCYHTLAIYASTIKTNPKLLANTALHVSLKSTNSTSRLMKRASKTPGLD
PRVLAAMLDCVEEVGDAVYELQRSIEEMDHAGGSNFSMVMNDVVTWVSASLTDDDTCMDGFAEGAVNKKVKTTVKRHLGRIARLTSNALALVNRYASSKA
NLP
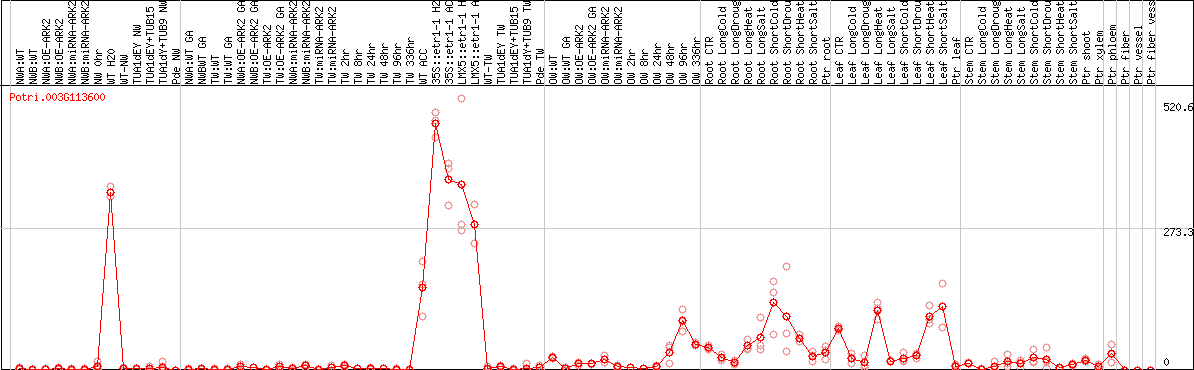
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G113600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.