Potri.003G125900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.003G125900.3 pacid=42787001 polypeptide=Potri.003G125900.3.p locus=Potri.003G125900 ID=Potri.003G125900.3.v4.1 annot-version=v4.1
ATGATGAGCAACCTTAAGCTAGGGGTAGAAGTGGTTAGTGCGCACAATCTTTTGCCCAAAGATGAGCACGGTTCATCCAGTGCCTTTGTAGAGCTCTGCT
TTGACGGCCAGAGGTTCCGCACAACCATCAAAGAAAAGGATCCCAATCCTGTGTGGAGTGAGTGTTTCTACTTCAACATCCCTGACCCTTCCAACCTACA
CTATCTTACTCTTGATGCCCATGTTTACAATAACATCAGAGCTACCAACTCCAGGTACTTCCTTGGTAAGGTTTGCCTCACTGGGAATTCATTTGTACCC
TACTCGGATGCTGTTGTGTTGCACTACCCTCTGGAAAAGCGTGGAATTTTCTCGCGTGTAAGAGGAGAGCTTGGCCTGAAAGTTTATATCACTGATGATG
CATCCATCAAATCCTCTACCCCACTTCCTGCAGTTGAATCTTTGCCAACAAAGGATCCAGGCTTAACACATGCCGTGGCTCCAATGGTAGACCCTATGAC
AAATACTGTATCTCATAAAAGAGTTGAGAGACACACCTTTCATCATCTTCCTAACCCAAATCACCAGCAACAACAGCATCAGAATCATTCTTCTGCTCCT
TCAATCACCCATCATGTACCAAAGTATGTGGCTGATGAGATGAAAGCTGCAGAAACACAGCCTCCAAAGTTAGTTCGAATGCACTCTGCATCATCATCAC
AACCCGTTGACCATGCCCTTAAAGAGACAAGTCCCTTTCTTGGAGGGGGAAGAGTTGTTGGGGGTCGTGTTATTCGAGGAGACAAGACTGCAAGCACTTA
TGATCTTGTTGAACGGATGTACTTCCTGTATGTAAGGGTTGTTAAGGCTCGTGATCTTCCTGCCATGGATGTTACAGGAAGTCTCGATCCGTTTGTTGAG
GTGAGAGTTGGAAACTACAGAGGAATTACAAAGCACTTTGAGAAAAAGCAAAATCCAGAGTGGAATCAGGTGTTTGCTTTTTCAAGGGAACGGATGCAAG
CTTCTGTTTTAGAAGTTGTTATTAAGGACAAGGATCTTGTTAAAGATGACTTTGTAGGCGTCATAAGGTTTGATATCAACGAGGTTCCTTCGAGAGTCCC
ACCAGATAGTCCGCTAGCTCCGGAGTGGTATCGGCTTGAGGACAAGAAGGGAGAGAAGATAAAGGGCGAGCTAATGCTTGCGGTGTGGATTGGAACCCAA
GCAGATGAGACCTTTCCTGATGCGTGGCATTCTGATGCAGCTACTCCTGTAGATAATACACCGGCTACCTCCACTGTTACACGTTCAAAGGTCTATCATG
CACCAAGGTTGTGGTATGTACGTGTTAATGTTGTCGAGGCACAAGACTTGGTCCCATCAGAGAAGACACGTTTCCCTGAAGTGTATGCTAAGGTACAGAT
GGGAAACCAAGTTTTGAAAACGAAGACATGTCAGGCTCGAACTTTTAGTGCGTTATGGAATGAGGATCTTTTATTCGTTGCTGCTGAACCCTTCGAGGAC
CATCTAGTTCTTTCAGTTGAGGATCGTGTCGGTCCTGGAAAAGATGAGATAATCGGGATGGTCATCATCCCATTGCGCTCTGTGGAGAAGCGTGCTGATG
ATCGGATAATTCATTCTCGCTGGTTTAACCTGGAAAAGCCAGTTGCTGTAGATGTAGATCAGTTCAAGAAAGACAAGTTCTCTAGCCGTATTCATCTTCG
AGCCTGCTTAGATGGAGGATACCATGTTCTTGATGAGTCAACTCATTACAGCAGTGATCTTTGTCCAACGGCAAAACAGCTTTGGAGGCCTCCGATTGGG
ATACTGGAACTAGGCATCTTAAATGCTGTGGGACTCCATCCTCTGAAAACACGAGATGGGAGGGGTACAGCCGATACGTACTGTGTTGCAAAGTATGGTC
ACAAATGGGTCCGGACACGCACCCTTATTGACAACCCGTCTCCAAAATACAATGAGCAGTACACATGGGAAGTTTTTGATCCAGCTACAGTTCTTACAGT
TGGTGTATTTGACAACAGCCAGCTTGGGGGAAAAGGTTCAAATGGAAAGGACCTAAAGATTGGGAAGGTTCGGATTCGCATCTCTACACTTGAAACTGGG
CGTGTATATACGCACTCTTATCCCTTGCTGGTTCTTCATCCTACCGGGGTTAAGAAGATGGGGGAATTGCACTTGGCGATTAGATTTACATGCATATCTT
TTGCAAACATGCTTTATCAGTACTCACGACCACTTCTACCCAAAATGCACTATATCAGGCCATTCAATGTGATGCAGCTAGATATGCTCCGCCACCAAGC
TGTCAACATAGTAGCATTACGGTTAGGTCGGGCAGAACCTCCTCTTAGGAAAGAAGTGGTGGAGTACATGTCAGATGTGGACTCACACCTCTGGAGCATG
CGTCGAAGCAAGGCAAATTTTCTTCGCTTGATGACTGTTTTCTCAGGATTATTTACAGCTGGAAAGTGGTTCGAGGATATTTGCATGTGGAAGAATCCTA
TCACTACTGTGCTTGTTCATGTGCTCTATCTTATGCTTGCTTGCTTCCCAGAACTTATTCTGCCCACAGTTTTTCTTTACATGTTTCTGATAGGCATTTG
GAATTACCGATATCGACCAAGGTACCCTCCCCACATGAACACAAAGATTTCACAGGCTGAGGCAGTGCATCCTGATGAGCTAGATGAGGAATTTGATACA
TTCCCAACAAGCAGGAGCCCTGAGCTGGTGGGAATGAGGTATGATAGGTTACGAAGTGTTGCTGGAAGGATTCAAACTGTCATAGGCGATATAGCAACCC
AAGGGGAGCGATTTCAGGCACTGTTAAGCTGGCGTGATCCACGGGCCACTGCCATCTTTGTTATATTCTGCCTTGTAGCTGCACTGGTTTTGTTTGTGAC
GCCATTTCAGGTTATAGCAGCTTTGGCAGGCTTCTACATGATGAGGCATCCAAGATTTCGCTACAGGACACCATCTGTGCCCATTAACTTCTTTCGCAGG
CTTCCTGCTCGGACAGATAGTATGCTGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.003G125900.3 pacid=42787001 polypeptide=Potri.003G125900.3.p locus=Potri.003G125900 ID=Potri.003G125900.3.v4.1 annot-version=v4.1
MMSNLKLGVEVVSAHNLLPKDEHGSSSAFVELCFDGQRFRTTIKEKDPNPVWSECFYFNIPDPSNLHYLTLDAHVYNNIRATNSRYFLGKVCLTGNSFVP
YSDAVVLHYPLEKRGIFSRVRGELGLKVYITDDASIKSSTPLPAVESLPTKDPGLTHAVAPMVDPMTNTVSHKRVERHTFHHLPNPNHQQQQHQNHSSAP
SITHHVPKYVADEMKAAETQPPKLVRMHSASSSQPVDHALKETSPFLGGGRVVGGRVIRGDKTASTYDLVERMYFLYVRVVKARDLPAMDVTGSLDPFVE
VRVGNYRGITKHFEKKQNPEWNQVFAFSRERMQASVLEVVIKDKDLVKDDFVGVIRFDINEVPSRVPPDSPLAPEWYRLEDKKGEKIKGELMLAVWIGTQ
ADETFPDAWHSDAATPVDNTPATSTVTRSKVYHAPRLWYVRVNVVEAQDLVPSEKTRFPEVYAKVQMGNQVLKTKTCQARTFSALWNEDLLFVAAEPFED
HLVLSVEDRVGPGKDEIIGMVIIPLRSVEKRADDRIIHSRWFNLEKPVAVDVDQFKKDKFSSRIHLRACLDGGYHVLDESTHYSSDLCPTAKQLWRPPIG
ILELGILNAVGLHPLKTRDGRGTADTYCVAKYGHKWVRTRTLIDNPSPKYNEQYTWEVFDPATVLTVGVFDNSQLGGKGSNGKDLKIGKVRIRISTLETG
RVYTHSYPLLVLHPTGVKKMGELHLAIRFTCISFANMLYQYSRPLLPKMHYIRPFNVMQLDMLRHQAVNIVALRLGRAEPPLRKEVVEYMSDVDSHLWSM
RRSKANFLRLMTVFSGLFTAGKWFEDICMWKNPITTVLVHVLYLMLACFPELILPTVFLYMFLIGIWNYRYRPRYPPHMNTKISQAEAVHPDELDEEFDT
FPTSRSPELVGMRYDRLRSVAGRIQTVIGDIATQGERFQALLSWRDPRATAIFVIFCLVAALVLFVTPFQVIAALAGFYMMRHPRFRYRTPSVPINFFRR
LPARTDSML
|
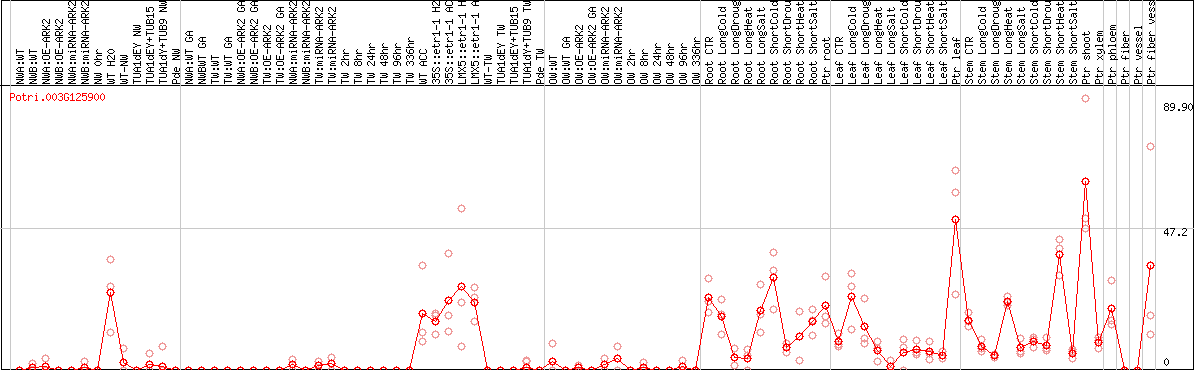
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G125900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
