Potri.003G143000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.003G143000.1 pacid=42785675 polypeptide=Potri.003G143000.1.p locus=Potri.003G143000 ID=Potri.003G143000.1.v4.1 annot-version=v4.1
ATGCAAAAACCAATAGAAGGTCGAGGAGTAGGAGGAGGTGGAGGTGAGCCAGAAGGAATGAAGTCAATGGAGTTGGATCTTGATCTCGATAACTCTTGGC
CTTTAGATCAGATCTCTTTCATGTCCTCAAACCCCATGTCTCCTTTTCTTATTTCTACTTCTACTGAACAACCTTGTTCTCCTCTTTGGGCTTTCTCTGA
TGCCGTTGATGATAGACTTGCTGCTACTGCTTCTGGTCAGGCTTCCCCTGCTTTCGCTGCTGCTGCTGCCCCCCGCTTGTCTGATTATCCTATCTTACTC
ACATGTAATCCAAATTTGATAACCGAAAGCCAAGGGGAAAATGATGATAACAGTAAACTTCCTTCTCCATTTTTGGGATTGATGCCTATTGACAACCCAG
ATGGGTATTGTATGATAAAAGAGAGAATGACACAGGCCCTAAGATATTTCAAAGAATCAACTGAACAACATGTTCTAGCTCAGGTTTGGGCACCTGTGAA
GAATGGGGGTCAACATGTATTGACAACTTCAGGGCAACCCTTTGTTCTGGACCCTCATAGTAATGGACTTCATCAGTATAGGATGGTCTCTCTCATGTAC
ATGTTCTCTGTGGACGGCGAGAGTGACAGAGAGCTTGGGCTTCCTGGCCGTGTTTTCAGACAAAAATCTCCAGAATGGACTCCAAATGTTCAGTATTACT
CCAGCAAGGAGTATTCACGGCTTGATCATGCTCTACGCTACAATGTCCGAGGAACATTGGCTTTGCCGGTCTTTGAACCTTCTGGACAGTCTTGTGTTGG
TGTTCTTGAGCTCATTATGAATTCACAGAAGATCAACTATGCACCTGAGGTTGATAAAGTGTGCAAGGCACTTGAGGCAGTAAATTTAAAAAGTTCAGAG
ATATTGGATCCTCCAAGCATACAGATTTGTAATGAAGGTCGCCAAAATGCGCTGTCTGAAATCTTGGAGATATTGACAATGGTGTGTGAAACTCATAAAT
TGCCCTTGGCTCAGACATGGGTTCCATGCATACATCGTAGTGTCTTAACGTATGGTGGTGGTCTGAAGAAGAGTTGTACCAGCTTTGATGGCAATTGCAA
TGGCCAAGTCTGTATGTCCACTACTGATGTGGCATTCTATGTAGTGGATGCTCGCATGTGGGGTTTTCGGGAGGCCTGTCTTGAGCATCACTTACAAAAG
GGTCAGGGTGTTGCTGGGAGGGCGTTTTTGTCCCAGAACTCATGCTTCTGCCCAGATATCACCCAGTTCTGCAAGACTGAGTATCCTCTAGTACATTATG
CACGCATGTTTGGATTAACCAGCTGTTTTGCAATCTTCTTGCGGAGCAGTTATACTGGAGATGATGATTATATTCTTGAATTTTTTCTGCCCCCTAGCAT
CACAGACAGTCACGAACAGAAGACATTTTTAGGCTCCATATTGGCAACAATGAAGCAGGATTTCCAGAGTCTCAAGGTTGCTTCAGGGATGGACCTTGAA
GAGGAAGGATTTGTAGAGATGATTGAAGCCACTACTAATGGGAGACTTGAATGCATTCAGATACCCCAACCTACAAAATCTCCCCCTGGTGATAATATGT
TGCCAAATGAAGGACACATTGAGCAAATTGATTCAGAAAAGAACAAGTTAATGTTTGACTTGGATGTCATAAAAAATGGAGGCAGTGCTGTTCAAGCTGA
CAGAAGACAAACTCCTCCATCTCATCCAGAGAAGAAAGGGACAAAAAAGCCAACAGAGAGGAAACGCGGAAAGGCAGAGAAAACAATTAGTCTAGAGGTT
CTCCAACAATATTTTGCTGGAAGTCTGAAAGATGCTGCAAAGAGACTTGGTGTTTGCCCTACTACAATGAAGCGCATTTGCAGGCAGCATGGGATCTCCC
GCTGGCCATCTCGCAAGATCAACAAGGTTAATCGTTCCCTGTCTAAGCTAAAGTGGGTAATTGAATCAGTCCAAGGCACTGAAGGAACATTTGATTTAAC
TCCTCTTACTACAAGTCCACTTCATGTTGCTGATGGCACCATCTCTTGGCCTTCCAATTTGAATGGGAGCAATCAACAGACCTCACCAAACTCTAAACCC
CCTGAATATCATGGCAACAGGAATGGATCACCTACCTGTAGAAAACCAGGAAGTGATGGGCAGGCTGGTTTTGAAGATCAATTGCTAGGATATAGAATAT
TGAGTCAAGAGAAACTCACTGTGCAAAACAGGTTTTCACCAGAGTTAGGTCGAGGCTCAAACAGATCCAAAAAAAGGAGTGGCTCAAGGGATGGGAGTGC
AGGGACTCCTACTTCTCATGACTCATGCCAAGGTAGCCCAGAAAATGAAAGTGCACCTGTAAAGGACCCATCTGTTTCTCCTGTCCATGAGCGGTGTATT
AAAGCTGGGGGATCACCTGGACTGGCTCTTCAACAAACCAAGGAACAAAATCTGTCATCTGCATACTCAATACCTGATGCTCTTGTTGCAACAGAAGCCC
ATGAACCGTTTGGTGGAATGCTAATAGAGGATGCGGGGAGCTCGAAAGATTTGAGAAACCTTTGTCCTGCTGTGGCTGAAGCTATTGTGGATGAACGAGT
TCCAGAATCTAGTTGGACAGATCCTCCATGTTTCAATATGCTTCCCACACAAATGTTTGCTGCTCCTTTGCATGCAATTCCGCAGGCAACACCTAGGCAA
GAAATGAAGAGTGTAACCATAAAGGCAACATACAGGGAAGACGTAATAAGGTTTCGAATTTCTCTGAGTTCTGGGATTGTGGAGTTGAAAGAGGAAGTGG
CCAAGAGGCTGAAGCTGGAGGTGGGTACTTTTGACATCAAGTATCTGGATGATGATCAAGAGTGGGTTCTGATAGCATGTGATGCTGACTTGCTGGAGTG
CATGGATGTTTCAAGATCATCAAGCAGTAATATAATCAGGCTCTCAGTTCATGATGCAAATGCCAATCTTGGGAGCTCCTGTGAGAGCACTGGGGAGTTA
TGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.003G143000.1 pacid=42785675 polypeptide=Potri.003G143000.1.p locus=Potri.003G143000 ID=Potri.003G143000.1.v4.1 annot-version=v4.1
MQKPIEGRGVGGGGGEPEGMKSMELDLDLDNSWPLDQISFMSSNPMSPFLISTSTEQPCSPLWAFSDAVDDRLAATASGQASPAFAAAAAPRLSDYPILL
TCNPNLITESQGENDDNSKLPSPFLGLMPIDNPDGYCMIKERMTQALRYFKESTEQHVLAQVWAPVKNGGQHVLTTSGQPFVLDPHSNGLHQYRMVSLMY
MFSVDGESDRELGLPGRVFRQKSPEWTPNVQYYSSKEYSRLDHALRYNVRGTLALPVFEPSGQSCVGVLELIMNSQKINYAPEVDKVCKALEAVNLKSSE
ILDPPSIQICNEGRQNALSEILEILTMVCETHKLPLAQTWVPCIHRSVLTYGGGLKKSCTSFDGNCNGQVCMSTTDVAFYVVDARMWGFREACLEHHLQK
GQGVAGRAFLSQNSCFCPDITQFCKTEYPLVHYARMFGLTSCFAIFLRSSYTGDDDYILEFFLPPSITDSHEQKTFLGSILATMKQDFQSLKVASGMDLE
EEGFVEMIEATTNGRLECIQIPQPTKSPPGDNMLPNEGHIEQIDSEKNKLMFDLDVIKNGGSAVQADRRQTPPSHPEKKGTKKPTERKRGKAEKTISLEV
LQQYFAGSLKDAAKRLGVCPTTMKRICRQHGISRWPSRKINKVNRSLSKLKWVIESVQGTEGTFDLTPLTTSPLHVADGTISWPSNLNGSNQQTSPNSKP
PEYHGNRNGSPTCRKPGSDGQAGFEDQLLGYRILSQEKLTVQNRFSPELGRGSNRSKKRSGSRDGSAGTPTSHDSCQGSPENESAPVKDPSVSPVHERCI
KAGGSPGLALQQTKEQNLSSAYSIPDALVATEAHEPFGGMLIEDAGSSKDLRNLCPAVAEAIVDERVPESSWTDPPCFNMLPTQMFAAPLHAIPQATPRQ
EMKSVTIKATYREDVIRFRISLSSGIVELKEEVAKRLKLEVGTFDIKYLDDDQEWVLIACDADLLECMDVSRSSSSNIIRLSVHDANANLGSSCESTGEL
|
DESeq2's median of ratios [POPLAR]
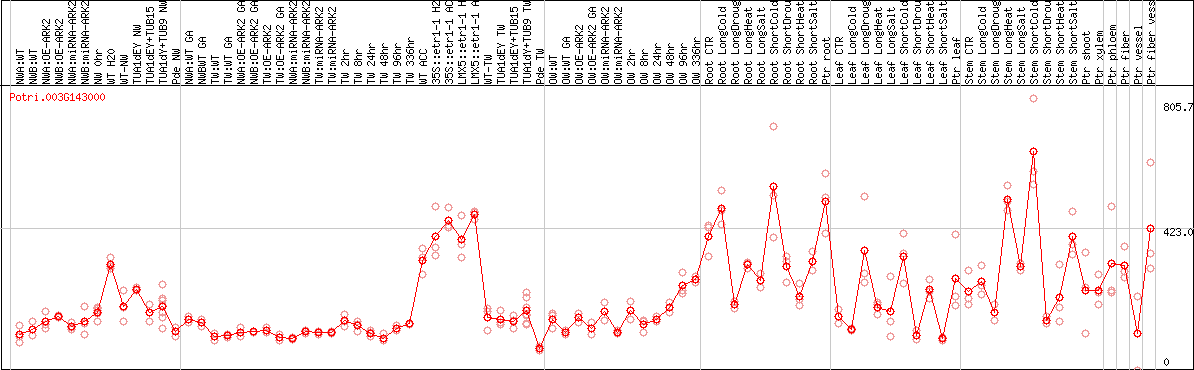
Coexpressed genes
Potri.003G143000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
