External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G48655 77 / 6e-17
RING/U-box superfamily protein (.1.2.3)
AT3G07200 76 / 7e-17
RING/U-box superfamily protein (.1.2)
AT2G42030 49 / 2e-06
RING/U-box superfamily protein (.1)
AT4G03510 47 / 5e-06
ATRMA1, RMA1
RING membrane-anchor 1 (.1.2)
AT1G18660 47 / 7e-06
zinc finger (C3HC4-type RING finger) family protein (.1), zinc finger (C3HC4-type RING finger) family protein (.2), zinc finger (C3HC4-type RING finger) family protein (.3), zinc finger (C3HC4-type RING finger) family protein (.4)
AT4G21070 47 / 9e-06
ATBRCA1
ARABIDOPSIS THALIANA BREAST CANCER SUSCEPTIBILITY1, breast cancer susceptibility1 (.1)
AT3G58030 46 / 9e-06
RING/U-box superfamily protein (.1.2.3.4)
AT2G32950 45 / 2e-05
FUS1, EMB168, DET340, ATCOP1, COP1
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
AT3G16090 45 / 2e-05
AtHrd1A
homolog of yeast Hrd1, RING/U-box superfamily protein (.1)
AT1G65040 45 / 3e-05
AtHrd1B
homolog of yeast Hrd1, RING/U-box superfamily protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G082200
285 / 2e-97
AT3G07200 86 / 2e-20
RING/U-box superfamily protein (.1.2)
Potri.002G245500
75 / 3e-16
AT5G48655 154 / 2e-47
RING/U-box superfamily protein (.1.2.3)
Potri.014G191400
74 / 5e-16
AT5G48655 158 / 8e-49
RING/U-box superfamily protein (.1.2.3)
Potri.014G159300
48 / 3e-06
AT2G32950 1091 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.002G141300
48 / 3e-06
AT2G44950 877 / 0.0
REDUCED DORMANCY 4, histone mono-ubiquitination 1 (.1)
Potri.014G055800
47 / 8e-06
AT2G44950 917 / 0.0
REDUCED DORMANCY 4, histone mono-ubiquitination 1 (.1)
Potri.001G358100
47 / 1e-05
AT4G21070 649 / 0.0
ARABIDOPSIS THALIANA BREAST CANCER SUSCEPTIBILITY1, breast cancer susceptibility1 (.1)
Potri.004G002700
45 / 2e-05
AT2G32950 832 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.012G068100
45 / 2e-05
AT1G18660 679 / 0.0
zinc finger (C3HC4-type RING finger) family protein (.1), zinc finger (C3HC4-type RING finger) family protein (.2), zinc finger (C3HC4-type RING finger) family protein (.3), zinc finger (C3HC4-type RING finger) family protein (.4)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022774
79 / 2e-17
AT5G48655 153 / 7e-47
RING/U-box superfamily protein (.1.2.3)
Lus10022569
72 / 2e-15
AT3G07200 81 / 3e-19
RING/U-box superfamily protein (.1.2)
Lus10018271
51 / 4e-08
AT3G07200 51 / 2e-08
RING/U-box superfamily protein (.1.2)
Lus10040641
48 / 8e-07
AT3G07200 46 / 1e-06
RING/U-box superfamily protein (.1.2)
Lus10036151
48 / 4e-06
AT4G21070 381 / 6e-118
ARABIDOPSIS THALIANA BREAST CANCER SUSCEPTIBILITY1, breast cancer susceptibility1 (.1)
Lus10029990
46 / 2e-05
AT2G32950 1088 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10002584
45 / 3e-05
AT1G61620 459 / 2e-164
phosphoinositide binding (.1)
Lus10018609
45 / 3e-05
AT5G44280 375 / 3e-124
ARABIDOPSIS THALIANA RING 1A, RING 1A (.1.2)
Lus10018409
45 / 3e-05
AT1G61620 459 / 3e-164
phosphoinositide binding (.1)
Lus10040359
45 / 3e-05
AT3G16090 652 / 0.0
homolog of yeast Hrd1, RING/U-box superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0229
RING
PF13923
zf-C3HC4_2
Zinc finger, C3HC4 type (RING finger)
Representative CDS sequence
>Potri.003G148300.1 pacid=42786391 polypeptide=Potri.003G148300.1.p locus=Potri.003G148300 ID=Potri.003G148300.1.v4.1 annot-version=v4.1
ATGGATTTACTGGAGAATTTTGACAGCAACATGGTTTTTATGGAGAAAGCAGAAAACACTATGGATTTACTGGGGAGTGTGGATCTTCTGGTTGAAAATT
TGTGGGAAGCCGAGAGAAGTTACTATCAGCACTTTGATCTCAACTTTCCAGTTCCTGTTGAGAACAATATACCTCCCTGTGGCCCGTGGTTTCCTGCCCA
ACCACAGATTTTTCAAGCTGAAGCATATGGATTGACACGACAACGTGGAACTGCTCGAGATGTTGAGGTCATTGATGATGATATTGCTATAATCTCCCCC
AACACATTTGATCAAGCCAGGAAGAAAGCTAGGAGAAATCCTGATGCAGAAGTAAGGGAAAATATAAATATGGCCAATCAGGTTCATGCTGCTGGGAGTG
TGCCCCAACTCCCTGGATCATCCCAGGCTGTGCCACCATCCCAGTTCCCTGGATCGTCCCAGACTGTGCCACCGCCCCAACTTTCTGGATTGTCCCTGAC
GGTGCCACCGCCTCATTTTTCAGGATTGTCCCAAACTGCACCGCCGCCACCAGCACCAATGTTTTGTTGCCCGATTTGTATGGACGAGATGAAAGAAGCA
ACATCGACAAAATGTGGTCATGTTTTCTGCAAGAGTTGCATTGAAAAAGCACTGGCTGTCCAGAAGAAGTGCCCTACTTGCAGGATGAAATGTATAGCGA
AATCCATCTTCAGGATCTTTCTTCCTGCATTTCTTTAA
AA sequence
>Potri.003G148300.1 pacid=42786391 polypeptide=Potri.003G148300.1.p locus=Potri.003G148300 ID=Potri.003G148300.1.v4.1 annot-version=v4.1
MDLLENFDSNMVFMEKAENTMDLLGSVDLLVENLWEAERSYYQHFDLNFPVPVENNIPPCGPWFPAQPQIFQAEAYGLTRQRGTARDVEVIDDDIAIISP
NTFDQARKKARRNPDAEVRENINMANQVHAAGSVPQLPGSSQAVPPSQFPGSSQTVPPPQLSGLSLTVPPPHFSGLSQTAPPPPAPMFCCPICMDEMKEA
TSTKCGHVFCKSCIEKALAVQKKCPTCRMKCIAKSIFRIFLPAFL
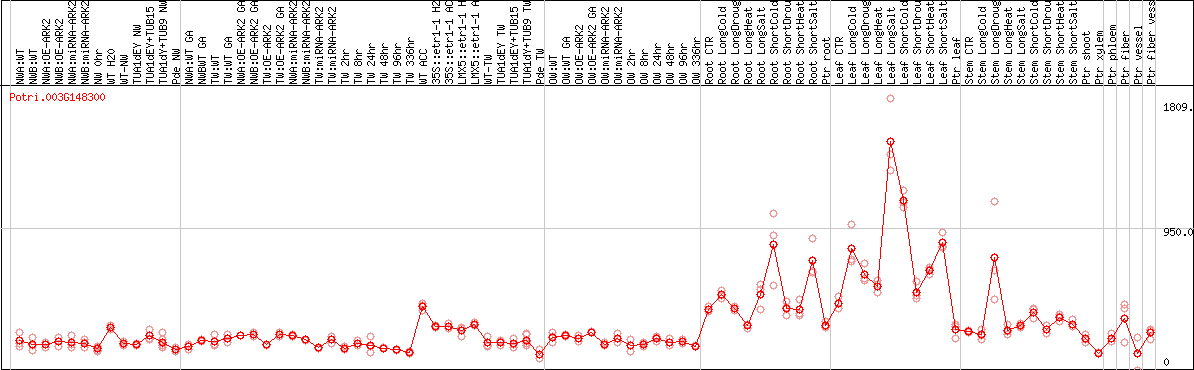
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G148300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.